Cloning vectors
- 2. INTRODUCTION • Cloning vector is an DNA molecule in which foreign DNA can be injected and which is further capable of replicating within in the host cell to produce multiple clone of recombinant DNA • Ex: plasmid, phage virus • Characters : 1.replicates autonomously 2.origin of replication 3.Selectable markers 4.multiple cloning site /poly linker 5.modern vector =cloning+expression
- 3. Commonly used cloning vectors
- 4. Recognition sequences of some REs Enzyme Recognition site Type of cut end EcoRI G↓A-A-T-T-C 5’ P extension BamHI G↓G-A-T-C-C 5’ P extension PstI C-T-G-C-A↓G 3’ P extension Sau3A1 ↓G-A-T-C 5’ P extension PvuII C-A-G↓C-T-G Blunt end HpaI G-T-T↓A-A-C Blunt end HaeIII G-G↓C-C Blunt end NotI G↓C-G-G-C-C-G-C 5’ P extension
- 5. What determines the choice of vector ? • insert size • vector size • restriction sites • copy number • cloning efficiency • ability to screen for inserts
- 6. Why Clone DNA? • Nucleotide sequence determination • Protein/enzyme/RNA function can be investigated • Mutations can be identified, e.g. gene defects related to specific diseases • Organisms can be ‘engineered’ for specific purposes, e.g. insulin production, insect resistance, etc.
- 7. Types of vectors • Different types of cloning vectors are used for different types of cloning experiments. • The vector is chosen according to the size and type of DNA to be cloned 1.plasmid 2.bacteriophage 3.cosmid 4.Human artificial chromosome 5. BACs 6. YACs
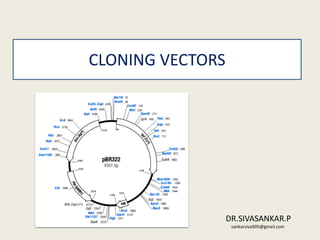
- 8. plasmid • Extrachromosamal double stranded circular self replicating DNA molecules • Not essential for survival of bacteria • 10-100 per cell • Size 1-500kb • Linear plasmid –streptomyces , borella burgdoferi • commercially available ones, eg: pGEM3, pBlueScript • Not well sited for creating genomic libraries • Transformation in efficient • Requires competant cells Types plasmid • pBR322 most commonly used • 4361 bp • Carries genes for ampicillin & tetracyline • Recognition site for EcoR I,hindIII,BamH I , Sal1,Pstl 1
- 9. Contd… • Other plasmid cloning vector • pUC19-2686bp with ampicillin resistant gene ,visual marker • Derivatives of pBR322-, pBR328 ,pBR329
- 10. Vector system Host cell Insert capacity (kb) Plasmid E. coli 0.1-10 Bacteriophage l E. coli 10-20 Cosmid E. coli 35-45 Bacteriophage P1 E. coli 80-100 BAC (bacterial artificial chromosome) E. coli 50-300 P1 bacteriophage-derived AC E. coli 100-300 YAC Yeast 100-2,000 Human AC Cultured human cells >2,000 Cloning vectors and their insert capacities
- 11. Bacteriaophage • Bacterial virus • Can take up larger DNA segments than plasmid Types 1.bacteriophage 2.phage M13 bacteriophage l : • Virus of E coli • DNA 50kbp • Cos end • Transformation is efficient 78-100% • Insert upto 25 kb • Out of 49 kb , 30kb used for lytic cycle • Many insertion site • Insertion vector Ex:GT10,GT11, Zap • Replacement vector ex: l EMBL4
- 12. Phage M13 vector • ss DNA of E coli • Complementary DNA synthesized in the cell • Useful in sequencing of DNA
- 13. Cosmids • Circular ds DNA • Plasmid +bacteriophage l • cos site added to the plasmid • Functions either one • Has both of the origin of replication • Insert upto 40kb • Introduced in E coli • No of copies good • Somewhat unstable ,difficult to maintain • Advantage : large fragment
- 14. Human artificial chromosome • Synthetic vector DNA , character of human chromosome • Self replicating microchromosome • Advantage :can incorporate too long human genes • Insert size >2000kb
- 15. Yeast Artificial Chromosomes • Synthetic DNA • YAC cloning vehicles are plasmids • Telomere-origin of replication • Have ori for bacteria also • Accepts large fragment of human DNA • Final chimeric DNA is a linear DNA molecule with telomeric ends • Large capacity vectors 200-2000Kbp • Yeast & bacteria as host • Cloning efficiency is low
- 16. Bacterial artificial chromosome • F-plasmid of bacteria combines with chromosome & replicates • Insert size 300K bp • Larger genomes can be studied ,can be cultured in bacteria requiring mammalian cells
- 17. THANK YOU
