The document outlines methods of manifold learning for dimension reduction in large datasets, emphasizing the motivation for compressing and processing massive data generated in various fields. It introduces the mathematical formalization of the problem, examples from different data types like molecular dynamics and handwritten digits, and categorizes methods into convex and non-convex optimizations. Key techniques discussed include Principal Components Analysis (PCA), kernel-based PCA, and Laplacian eigenmaps, which aim to preserve the manifold structure of the data while reducing its dimensionality.




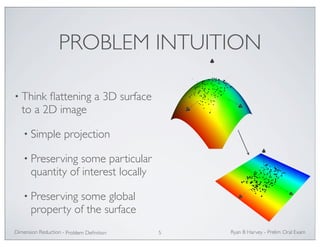
![THE PROBLEM (FORMALIZED)
• Inputs:
• Outputs:
X = [x1, . . . ,xn], xk ∈ RD
Y = [y1, . . . , yn], yk ∈ Rd, d D
M⊂ Rd
• Assumption: data live on some manifold embedded
in RD , and inputs X are samples taken in RD
of the underlying
manifold .
M
• Problem statement: Find a reduced representation Y of
X
which best preserves the manifold structure of the data, as
defined by some metric of interest.
Ryan Dimension Reduction - B Harvey - Prelim Oral Exam
Problem Definition 6](https://image.slidesharecdn.com/ob7etaomrryvt5gi6uxu-signature-0bdd673cabf44901f5f756298743b37f4849bc69f9003ecb916d5b3896bca0e6-poli-141113155943-conversion-gate02/85/Methods-of-Manifold-Learning-for-Dimension-Reduction-of-Large-Data-Sets-6-320.jpg)
![An example from Molecular Dynamics, I
EXAMPLES Text OF documents
DATA SETS
1000 Science News articles, from 8 different categories. We
compute about 10000 coordinates, i-th coordinate of document
d represents frequency in document d of the i-th word in a fixed
dictionary.
Data base of about 60, 000 28 × 28 gray-scale pictures of
handwritten digits, Text
collected by USPS. Point cloud in R282 .
Goal: automatic recognition.
Mauro Maggioni Geometry of data sets in high dimensions and learning Hyperspectral
The dynamics of a small protein in a bath of water molecules is
approximated by a Langevin system of stochastic equations
x˙ = −∇U(x) + w˙ .
Handwritten Digits
10
model in the form of Equation 3, we can synthesize new shapes through the walking
cycle. In these examples only 10 samples were used to embed the manifold for half a
cycle on a unit circle in 2D and to learn the model. Silhouettes at intermediate body
configurations were synthesized (at the middle point between each two centers) using
the learned model. The learned model can successfully interpolate shapes at intermedi-ate
The set of states of the protein is a noisy set of points in R36.
configurations (never seen in the learning) using only two-dimensional embedding.
The figure shows results for three different peoples.
Mauro Maggioni Analysis of High-dimensional Data Sets and Graphs
Learn Mapping from
Embedding to 3D
Learn Nonlinear Mapping
Manifold
Embedding
Visual
input
3D pose
from Embedding
to visual input
Learn Nonlinear
Manifold Embedding
(a) Learning components
Manifold
Embedding
(view based)
Visual
input
3D pose
Image
Closed Form
solution for
inverse mapping
Collections
Error Criteria
Manifold Selection
View Determination
3D pose
interpolation
(b) pose estimation.
(c) Synthesis.
Video
Fig. 4. (a,b) Block diagram for the learning framework and 3D pose estimation. (c) Shape synthe-sis
for three different people. First, third and fifth rows: samples used in learning. Second, fourth,
sixth rows: interpolated shapes at intermediate configurations (never seen in the learning)
Dimension Reduction - Given a visual input (silhouette), and the learned model, we can recover the intrinsic
Ryan B Harvey - Prelim Oral Exam
body configuration, recover the view point, and reconstruct the input and detect any
spatial or temporal outliers. In other words, we can simultaneously solve for the pose,
view point, and reconstruct the input. A block diagram for recovering 3D pose and
view point given learned manifold models are shown in Figure 4. The framework [20]
Molecular
Dynamics
Set of 10, 000 picture (28 by 28 pixels) of 10 handwritten digits. Color represents the label (digit) of each point.
Problem Definition 7](https://image.slidesharecdn.com/ob7etaomrryvt5gi6uxu-signature-0bdd673cabf44901f5f756298743b37f4849bc69f9003ecb916d5b3896bca0e6-poli-141113155943-conversion-gate02/85/Methods-of-Manifold-Learning-for-Dimension-Reduction-of-Large-Data-Sets-7-320.jpg)



















![CONSTRUCTING THE
EIGENVALUE PROBLEM
• Extending the same argument to , with
and , we need to minimize the objective
function:
giving the minimization
F
FTDF = I
• This problem can also be solved by solving the generalized
eigenvalue problem for the minimum eigenvalues.
Ryan Dimension Reduction - B Harvey - Prelim Oral Exam
Laplacian Eigenmaps 29
f ∈ Rd F ∈ Rn × Rd
f (i) = [f (i)
1 , . . . , f (i)
d ]T
i
j ||f (i) − f (j)||2Wij = tr(FTLF)
argmin tr(FTLF)
Lf = λDf](https://image.slidesharecdn.com/ob7etaomrryvt5gi6uxu-signature-0bdd673cabf44901f5f756298743b37f4849bc69f9003ecb916d5b3896bca0e6-poli-141113155943-conversion-gate02/85/Methods-of-Manifold-Learning-for-Dimension-Reduction-of-Large-Data-Sets-29-320.jpg)








![CONNECTION TO THE
GRAPH LAPLACIAN
12
•We will show in three steps that for a function (under
appropriate assumptions) Ef ≈ L2f
:
(1) Fix a point and show that:
xi
[(I −W)f]i ≈ −12
f ∈M
j Wij(xi − xij )TH(xi − xij )
where H is the Hessian of f at xi
.
(2) Show that the expectation .
(3) Put steps (1) and (2) together to achieve the final result.
Ryan Dimension Reduction - B Harvey - Prelim Oral Exam
Locally Linear Embedding 39
E[vTHv] = rLf](https://image.slidesharecdn.com/ob7etaomrryvt5gi6uxu-signature-0bdd673cabf44901f5f756298743b37f4849bc69f9003ecb916d5b3896bca0e6-poli-141113155943-conversion-gate02/85/Methods-of-Manifold-Learning-for-Dimension-Reduction-of-Large-Data-Sets-39-320.jpg)
![• Show:
CONNECTION TO THE
GRAPH LAPLACIAN (1)
[(I −W)f]i ≈ −12
j Wij(xi − xij )TH(xi − xij )
• Consider a coordinate system in the tangent plane centered at
vj = xij − xi
o = xi o
and let . This is a vector originating at .
αj = Wij xi
• Let . Since is in the affine span of its neighbors
(and by construction of W
), we have
where .
Ryan Dimension Reduction - B Harvey - Prelim Oral Exam
Locally Linear Embedding 40
j αj = 1
o = xi =
j αjvj](https://image.slidesharecdn.com/ob7etaomrryvt5gi6uxu-signature-0bdd673cabf44901f5f756298743b37f4849bc69f9003ecb916d5b3896bca0e6-poli-141113155943-conversion-gate02/85/Methods-of-Manifold-Learning-for-Dimension-Reduction-of-Large-Data-Sets-40-320.jpg)

![CONNECTION TO THE
GRAPH LAPLACIAN (1)
[(I −W)f]i = f(o) −
•We have , and using Taylor’s
approximation for , we can write
j ∇f + 12
j Hvj)
• Since and , the first three terms
disappear, and
Ryan Dimension Reduction - B Harvey - Prelim Oral Exam
Locally Linear Embedding 42
j αjf(vj)
f(vj)
[(I −W)f]i = f(o) −
j αjf(vj)
≈ f(o) −
j αjf(o) −
j αjvT
j αj(vT
j αj = 1
j αjvj = o
[(I −W)f]i = f(o) −
j αjf(vj) ≈ −12
j αjvT
j Hvj](https://image.slidesharecdn.com/ob7etaomrryvt5gi6uxu-signature-0bdd673cabf44901f5f756298743b37f4849bc69f9003ecb916d5b3896bca0e6-poli-141113155943-conversion-gate02/85/Methods-of-Manifold-Learning-for-Dimension-Reduction-of-Large-Data-Sets-42-320.jpg)

![CONNECTION TO THE
GRAPH LAPLACIAN (2)
• Then using the Spectral theorem, we can write
E[vTHv] = E
• Since is independent of , we can replace
to get
i
Ryan Dimension Reduction - B Harvey - Prelim Oral Exam
Locally Linear Embedding 44
i λiv, ei2
E
v, ei2
E
v, ei2
= r
E[vTHv] = r (
i λi) = rtr(H) = rLf](https://image.slidesharecdn.com/ob7etaomrryvt5gi6uxu-signature-0bdd673cabf44901f5f756298743b37f4849bc69f9003ecb916d5b3896bca0e6-poli-141113155943-conversion-gate02/85/Methods-of-Manifold-Learning-for-Dimension-Reduction-of-Large-Data-Sets-44-320.jpg)
![CONNECTION TO THE
GRAPH LAPLACIAN (3)
• Now, putting these together, we have
[(I −W)f]i ≈ −12
• LLE minimizes which reduces to
finding eigenfunctions of , which can now
be interpreted as finding eigenfunctions of the iterated
Laplacian . Eigenfunctions of coincide with those of .
Ryan Dimension Reduction - B Harvey - Prelim Oral Exam
Locally Linear Embedding 45
j WijvT
j Hvj
E[vTHv] = rLf
(I −W)T (I −W)f ≈ 12
L2f
fT (I −W)T (I −W)f
(I −W)T (I −W)
L2 L L 2](https://image.slidesharecdn.com/ob7etaomrryvt5gi6uxu-signature-0bdd673cabf44901f5f756298743b37f4849bc69f9003ecb916d5b3896bca0e6-poli-141113155943-conversion-gate02/85/Methods-of-Manifold-Learning-for-Dimension-Reduction-of-Large-Data-Sets-45-320.jpg)