S1 Mapping
- 1. S1 MAPPING Presented by Wardah Shah Roll no. 07 Submitted to Dr. Abrar Qureshi Dept. Of Biotechnology
- 2. INTRODUCTION • Gene expression is the process by which information encoded in a gene is used in the synthesis of a functional gene product, that is, protein in case of mRNA transcript translation. • To understand how a gene is expressed, the RNA transcript must be studied, in particularly, 1. The removal of introns, 2. The exact locations of start and end points of transcription, 3. The signals that determine the start and end of transcription.
- 3. • S1 nuclease is a restriction enzyme that cleaves only single stranded nucleic acids, ssDNA or ssRNA, leading to formation of 5' phosphomononucleotide product or 5' phosphooligonucleotide product. • It has been isolated from Aspargillus oryzae. S1 ENDONUCLEASE
- 4. Properties of S1 Nuclease: • At high ionic strength, low pH (4-4.5) and in the presence of Zn ions (cofactor), S1 nuclease digests ssDNA very efficiently. • It is relatively stable against denaturing agents like urea, SDS and formaldehyde. • It removes single stranded regions from dsDNA. (conversion of cohesive ends to blunt ends) • Cleaves hairpin loops generated during synthesis of DNA. • Used I nuclease mapping techniques and nuclease protection assay
- 5. S1 Mapping: • Technique developed by Berk and Sharp by the study of adenovirus mRNAs • S1 Mapping is a laboratory method used for locating the start and end points of transcripts and for mapping introns. • This technique is used for quantifying the amount of mRNA transcripts, it can therefore identify the level of transcription of the gene in the cell at a given time.
- 6. DNA-mRNA Hybridization: • When hybrid is formed between complementary DNA strand and its mRNA transcript, then the boundaries between double and single stranded regions marks the start and end points of the mRNA. • Introns, which are present in DNA, but not in the mRNA, will loop out as additional single stranded regions. • On treatment with S1 Nuclease, all single stranded regions are cleaved out, leaving behind the hybrid regions. • Upon treatment with alkali, the RNA strands are degraded and the DNA fragments are recovered and their sizes measured on gel electrophoresis. • LIMITATION: Order of DNA sequence cannot be determined.
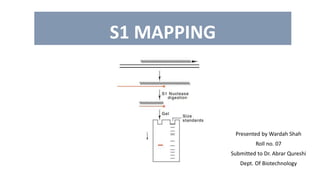
- 7. LOCATING START SITE: • Modifications to hybridization technique allow the precise start and end points of the transcript and of any introns it contains to be mapped onto the DNA sequence • The figure shows an example of S1 mapping for locating the start site of a transcript. • Here, a 400 bp fragment is created by restriction digestion by Sau3A. This fragment contains the start site of the gene. • The fragment is inserted into M13 cloning vector. The mRNA transcript is made to anneal with the singe stranded DNA in the vector. • The transcript anneals only to the complementary sequence. Rest of the single stranded vector and single stranded mRNA is cleaves by action of S1 endonuclease. • The double stranded hybrid is treated with alkali to degrade the RNA and recover the DNA which is run on gel electrophoresis. • The size of this fragment corresponds to the distance between the transcription start point and right hand Sau3A site. • The same strategy is used to locate the end site.
- 8. SUMMARY • In S1 mapping, a labeled DNA probe is used to detect 5’- or 3’-end of a transcript. • Hybridization of the probe to the transcript protects a portion of the probe from digestion by S1 nuclease, specific for single-stranded polynucleotides. • Length of the section of probe protected by the transcript locates the end of the transcript relative to the known location of an end of the probe. • Amount of probe protected is proportional to concentration of transcript, so S1 mapping can be quantitative.
- 9. THANK YOU