Next Generation Sequencing for Joubert Poster
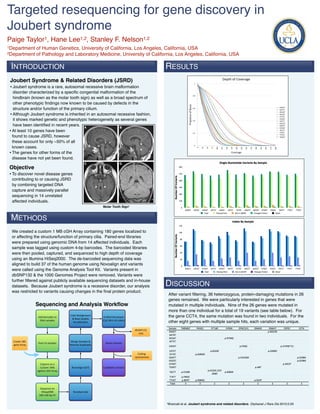
- 1. Targeted resequencing for gene discovery in Joubert syndrome Paige Taylor1, Hane Lee1,2, Stanley F. Nelson1,2 1Department of Human Genetics, University of California, Los Angeles, California, USA 2Department of Pathology and Laboratory Medicine, University of California, Los Angeles, California, USA INTRODUCTION RESULTS Joubert Syndrome & Related Disorders (JSRD) Results • Joubert syndrome is a rare, autosomal recessive brain malformation disorder characterized by a specific congenital malformation of the hindbrain (known as the molar tooth sign) as well as a broad spectrum of other phenotypic findings now known to be caused by defects in the structure and/or function of the primary cilium. • Although Joubert syndrome is inherited in an autosomal recessive fashion, it shows marked genetic and phenotypic heterogeneity as several genes have been identified in recent years. • At least 10 genes have been found to cause JSRD, however these account for only ~50% of all known cases. • The genes for other forms of the disease have not yet been found. Objective • To discover novel disease genes contributing to or causing JSRD by combining targeted DNA capture and massively parallel sequencing in 14 unrelated affected individuals. Molar Tooth Sign1 METHODS We created a custom 1 MB cGH Array containing 180 genes localized to or affecting the structure/function of primary cilia. Paired-end libraries were prepared using genomic DNA from 14 affected individuals. Each sample was tagged using custom 4-bp barcodes. The barcoded libraries were then pooled, captured, and sequenced to high depth of coverage using an Illumina HiSeq2000. The de-barcoded sequencing data was aligned to build 37 of the human genome using Novoalign and variants were called using the Genome Analysis Tool Kit. Variants present in dbSNP132 & the 1000 Genomes Project were removed. Variants were further filtered against publicly available sequencing datasets and in-house datasets. Because Joubert syndrome is a recessive disorder, our analysis DISCUSSION was restricted to variants causing changes in the final protein product. After variant filtering, 36 heterozygous, protein-damaging mutations in 26 genes remained. We were particularly interested in genes that were Sequencing and Analysis Workflow mutated in multiple individuals. Nine of the 26 genes were mutated in more than one individual for a total of 19 variants (see table below). For Indel#Realignment# the gene CCT4, the same mutation was found in two individuals. For the Add#barcodes#to# Unified#Genotyper &#Base#Quality# DNA#samples (Call#SNVs#&#Indels) RecalibraMon other eight genes with multiple sample hits, each variation was unique. dbSNP132,# 1KG Create#180R Merge#Samples#&# Pool#14#samples Novel#Variants gene#Array Remove#Duplicates CodingR synonymous Capture#on#a# Custom#1Mb# Novoalign#(b37) Candidate#Variants Agilent#cGH#Array Sequence#on# HiSeq2000# Novobarcode 100+100#bp#PE 1Brancati et al. Joubert syndrome and related disorders. Orphanet J Rare Dis 2010;5:20