This book club presentation summarized key concepts from Chapter 3 of Andreas Wagner's book "The Origins of Evolutionary Innovations". Specifically, it discussed how regulatory innovations like changes to transcription factor binding sites can evolve. It provided examples of how regulatory networks for galactose metabolism have diverged between yeast and Candida albicans. The presentation noted that genotype networks of regulatory circuits can be large but organisms can tolerate many regulatory changes without phenotype changes, making innovations possible.















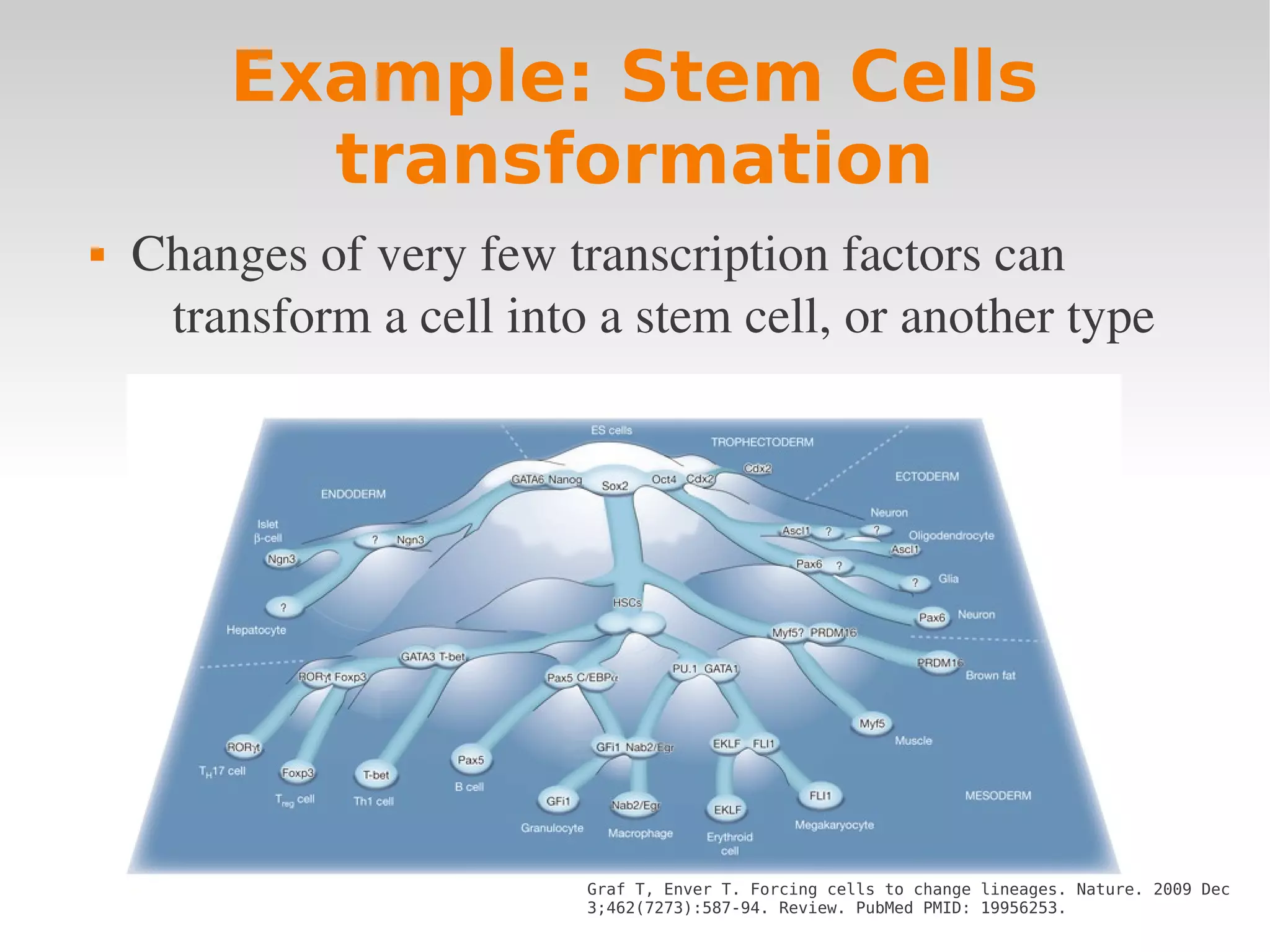

![Some phenotypes have
more genotypes than others
[1]: A. Wagner, The Origins of Evolutionary Innovations. Figure 3.2
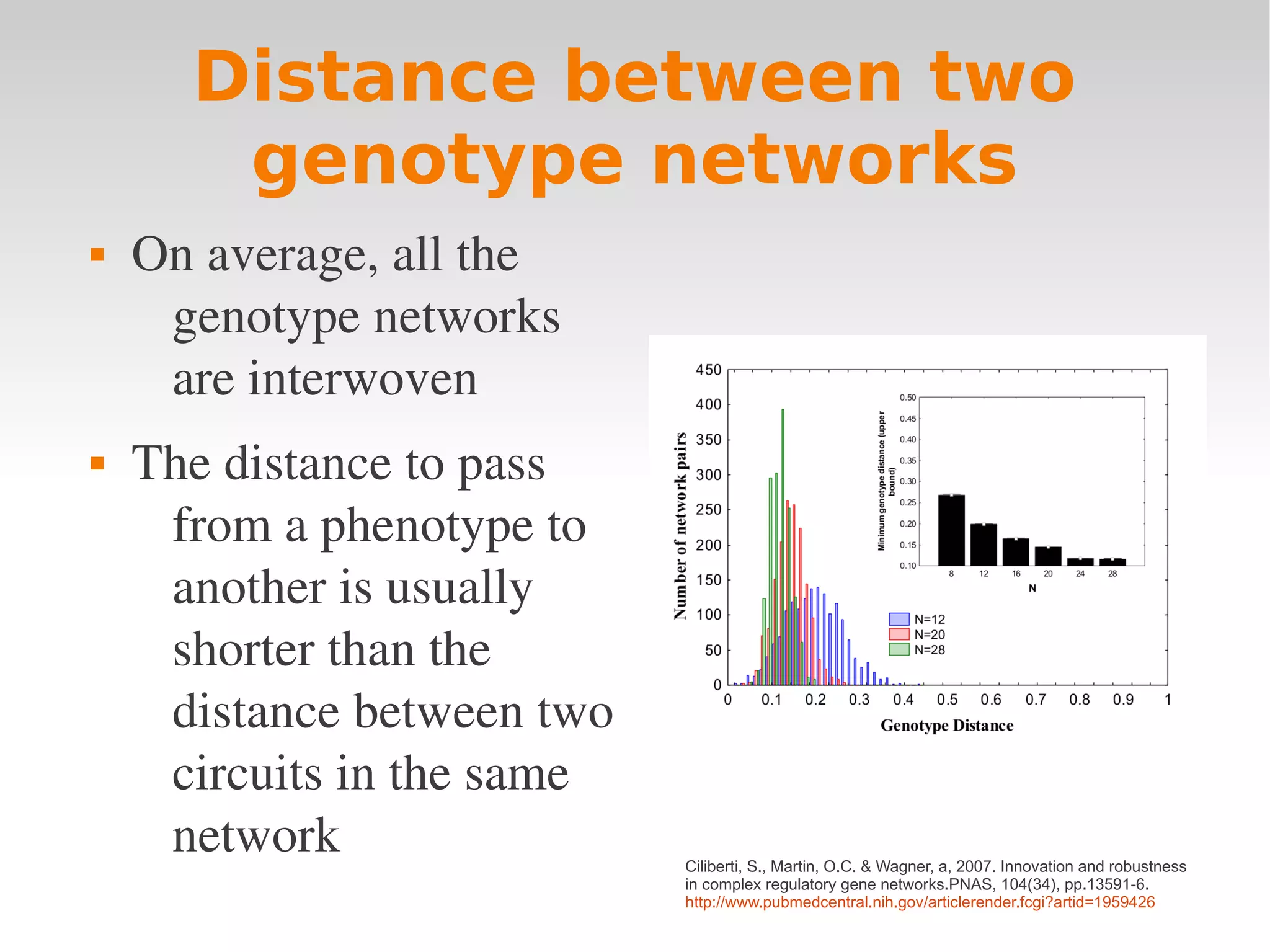
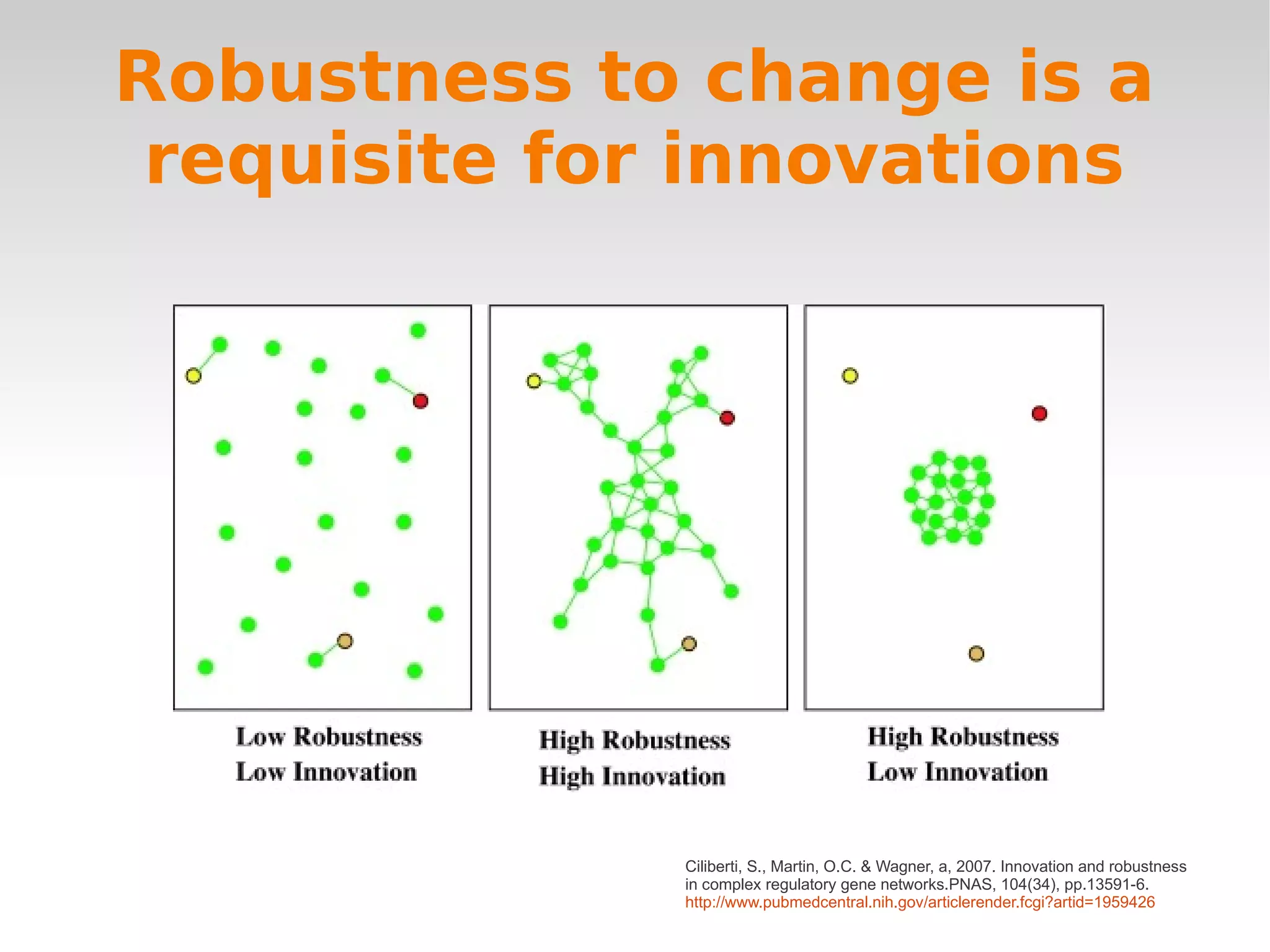
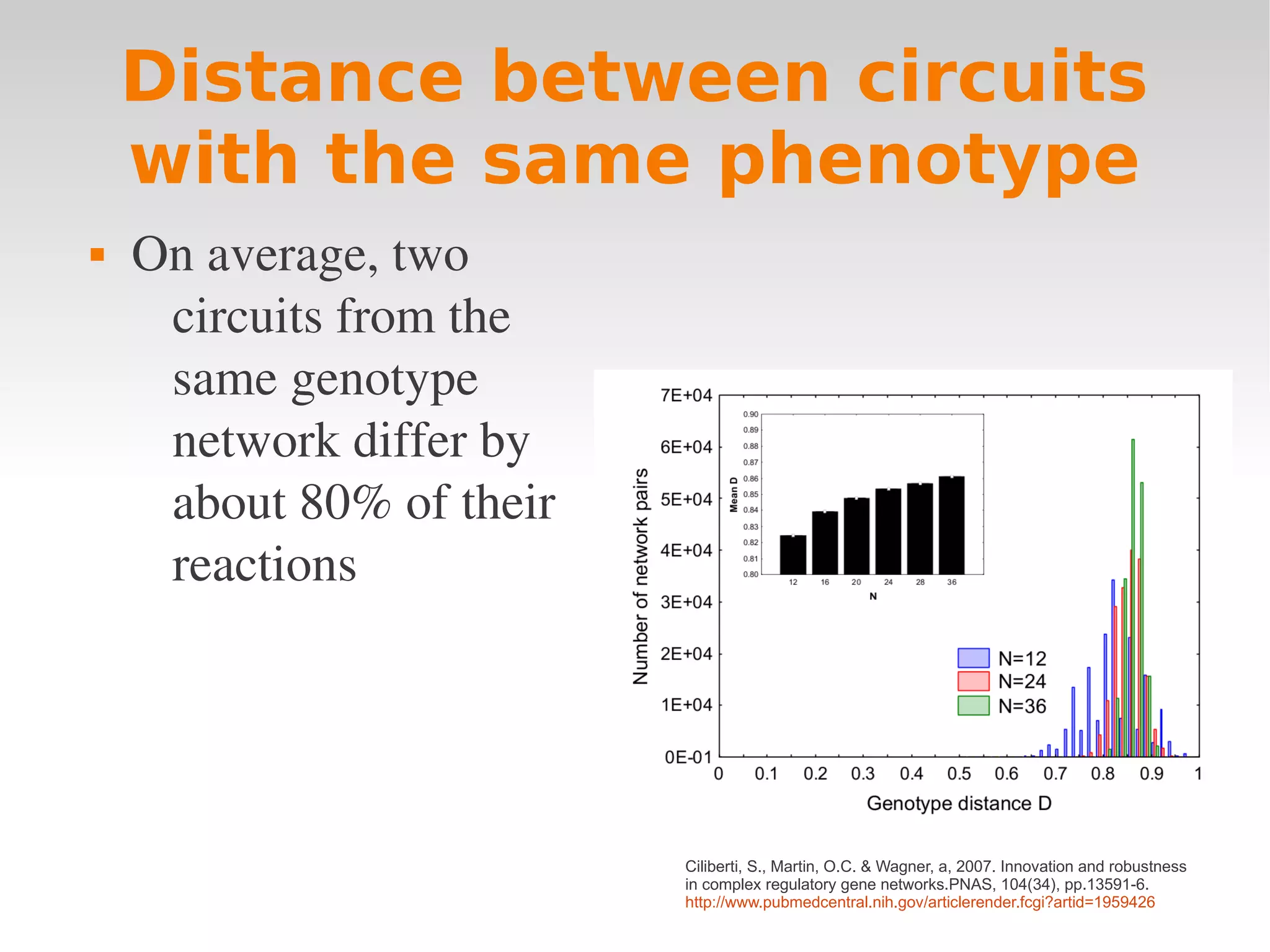
Ciliberti, S., Martin, O.C. & Wagner, a, 2007. Innovation and robustness
in complex regulatory gene networks.PNAS, 104(34), pp.13591-6.
http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=1959426](https://image.slidesharecdn.com/wagnerchapter3-120210061827-phpapp01/75/Wagner-chapter-3-19-2048.jpg)
![Phenotypes of neighbors
Neighbors of two
genotypes in the
same network are
usually different
[1]: A. Wagner, The Origins of Evolutionary Innovations. Figure 3.6
[1]: A. Wagner, The Origins of Evolutionary
Innovations. Figure 2.6](https://image.slidesharecdn.com/wagnerchapter3-120210061827-phpapp01/75/Wagner-chapter-3-21-2048.jpg)