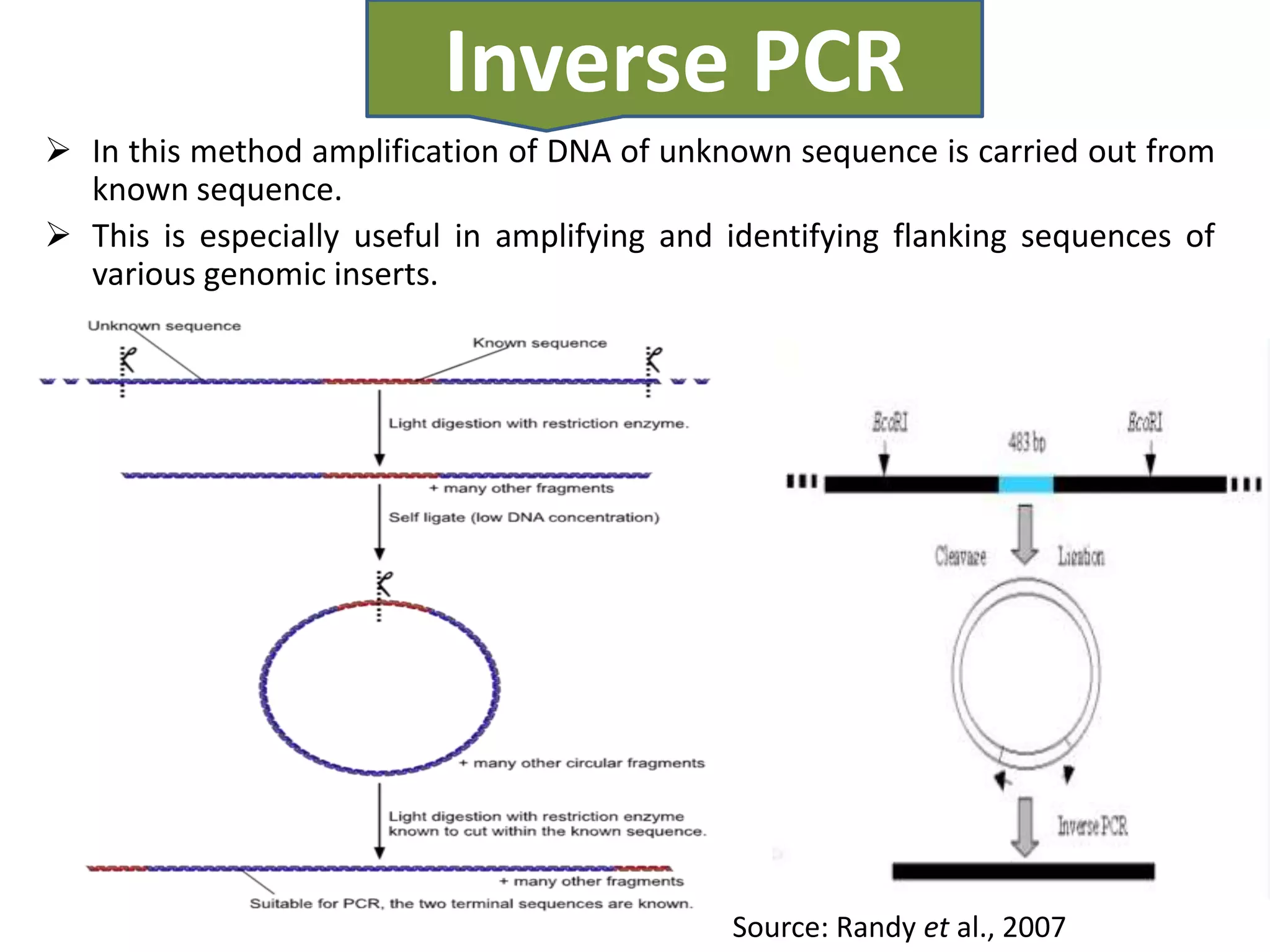
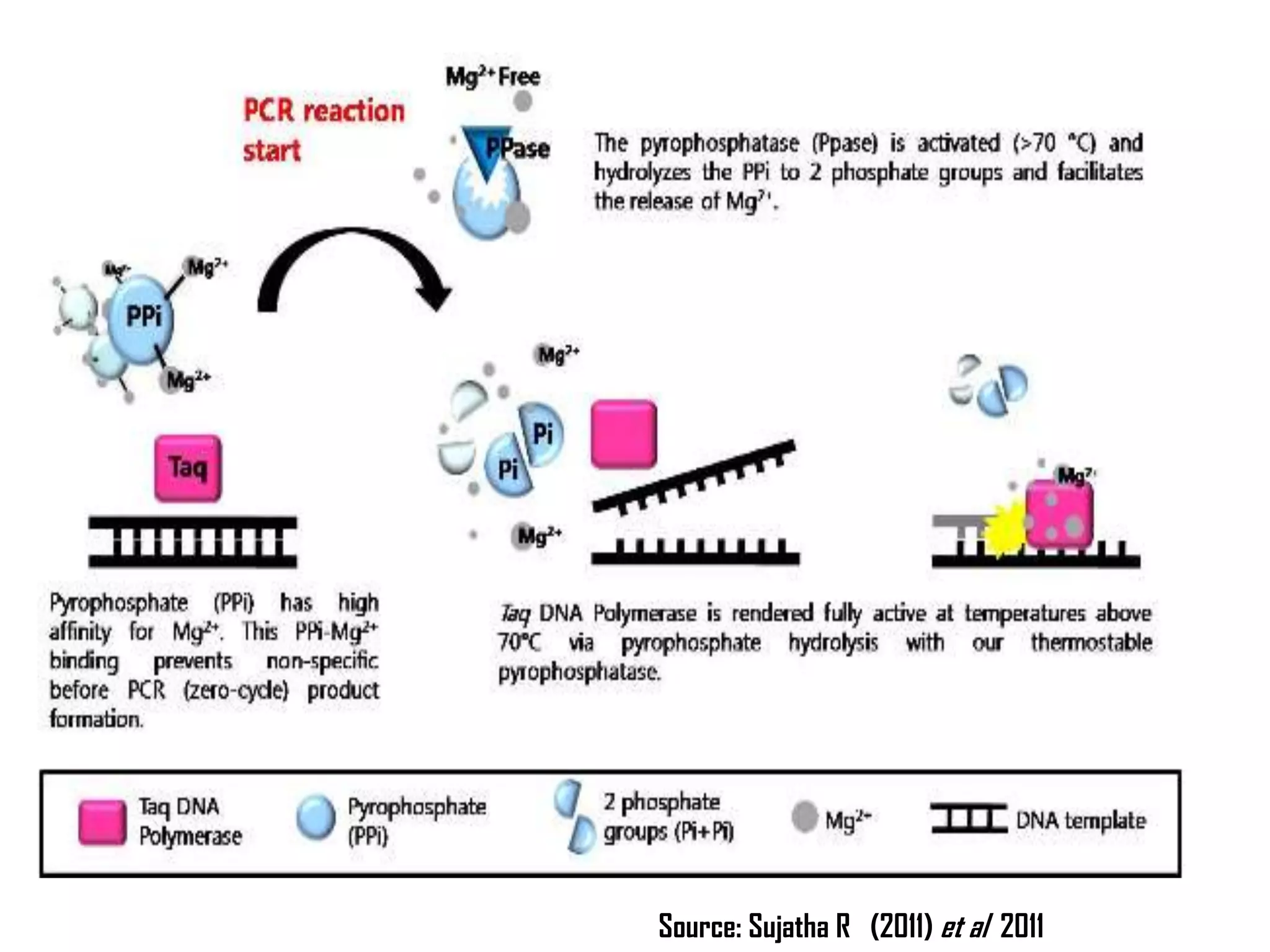
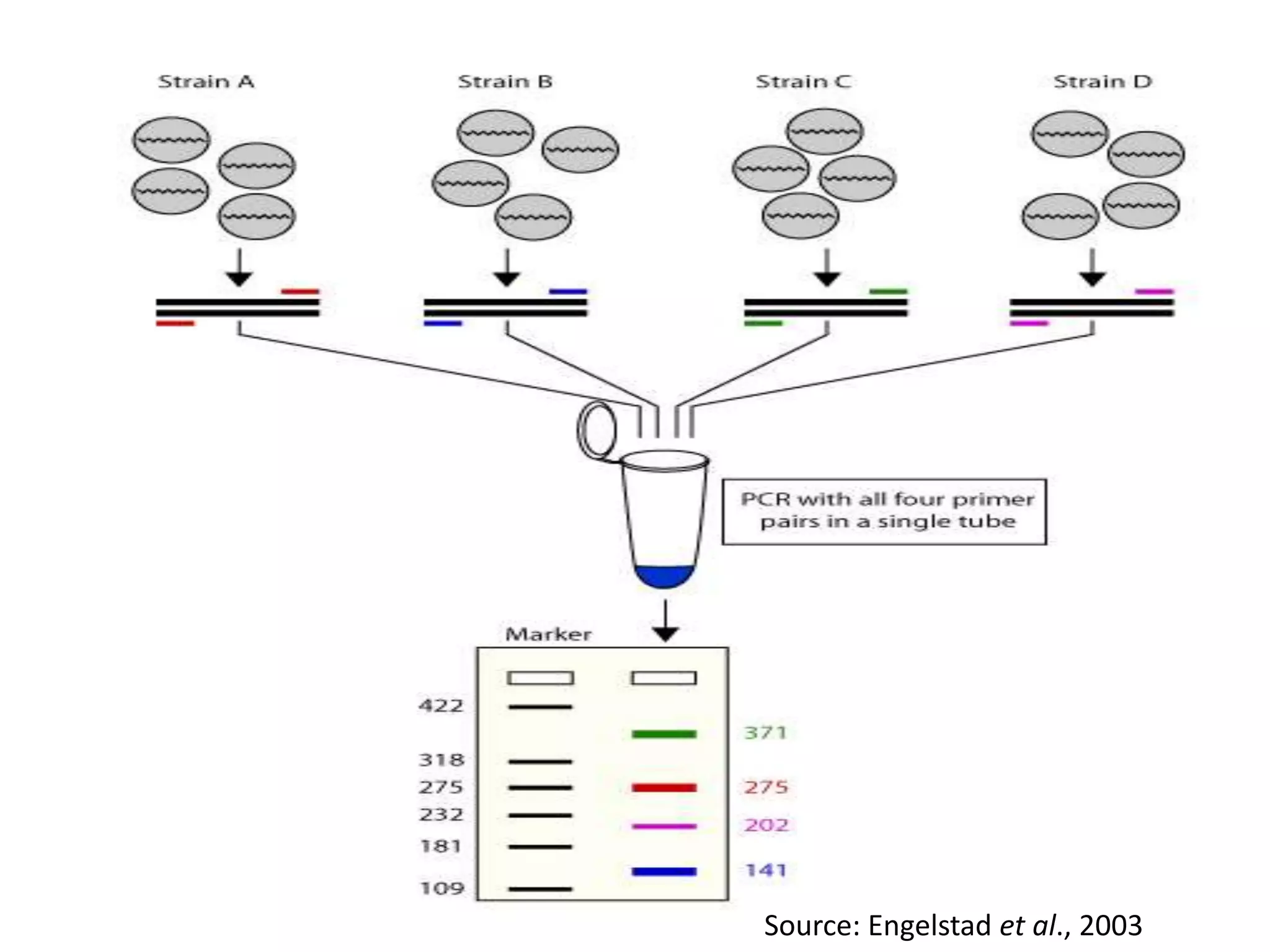
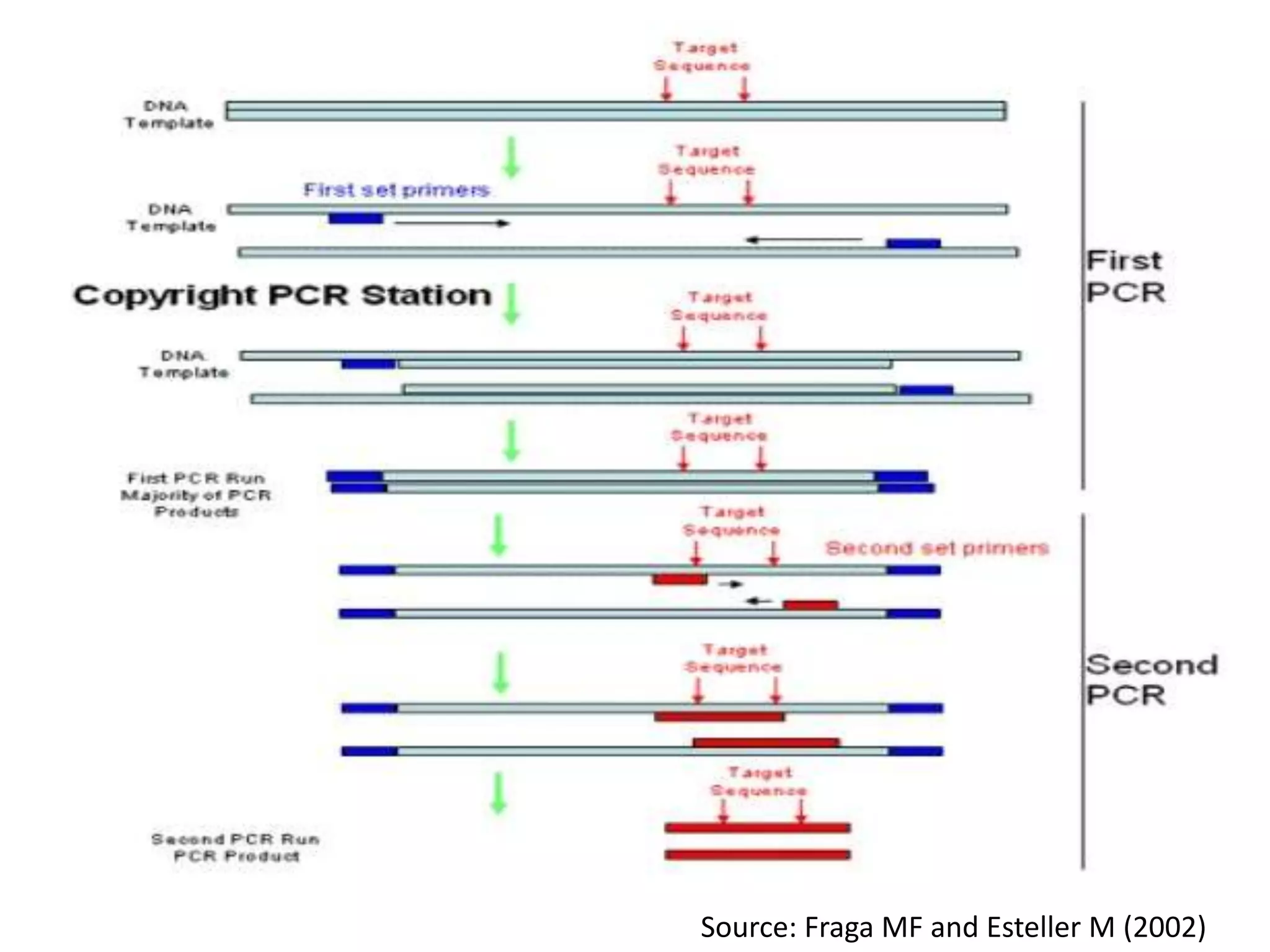
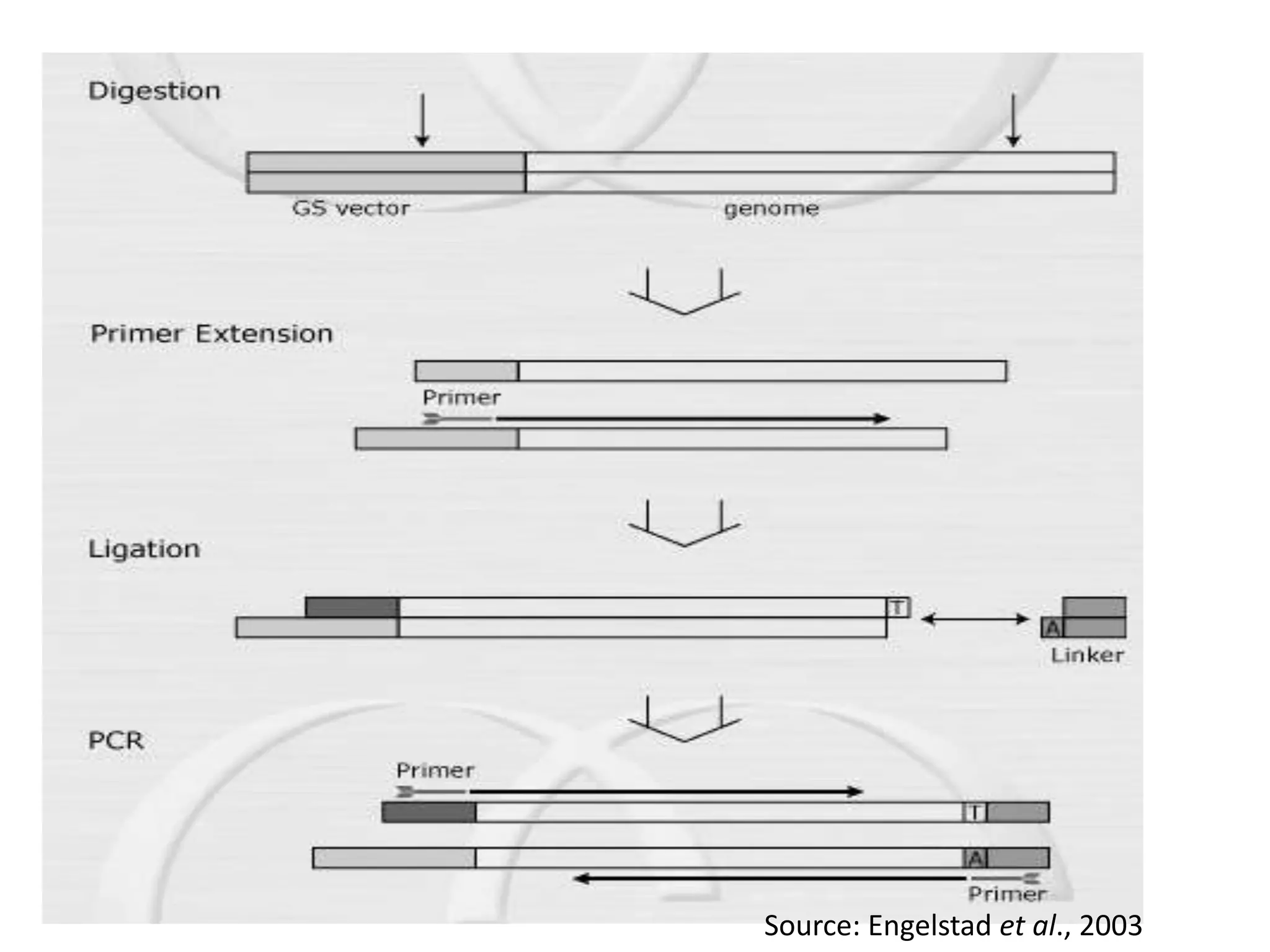
There are many types of polymerase chain reaction (PCR) that can be classified in different ways. Some of the major types include hot-start PCR, which prevents primer dimer formation; multiplex PCR, which amplifies multiple targets simultaneously; nested PCR, which uses two rounds of PCR for greater specificity; and ligation-mediated PCR, which uses linkers ligated to DNA fragments before amplification. PCR has various applications and techniques that optimize the process for different needs.