Session 3: High throughput analysis of chemical and genetic factors that influence phosphine toxicity and resistance
•Download as PPTX, PDF•
0 likes•336 views
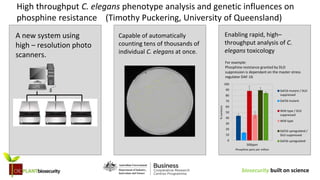
A new high-resolution photo scanning system enables rapid, high-throughput analysis of C. elegans toxicology. For example, an experiment showed that phosphine resistance granted by DLD suppression is dependent on the master stress regulator DAF-16. The scanning system is capable of automatically counting tens of thousands of individual C. elegans at once, allowing for high throughput analysis of C. elegans phenotypes and genetic influences on phosphine resistance.
Report
Share
Report
Share
Recommended
Satellite RNA

in this chapter covers the symptoms modulation and diseases severity increases and decreases. and role of SiRNA in diseases severity reduction. and also covers the types of SRNA..
Detection techniques of Tomato_Brown_Rugose_Fructose_Virus

It is based on result of detection techniques used in Department of virology at Universitat Politechnica de Valencia, Spain as a part of second semester internship.
Transcriptomics: A Tool for Plant Disease Management

Transcriptomics is one of the omics we can use to manage the plant diseases by modifications in the transcription process
Pathogen confusion as a strategy for controlling diseases caused by Xylella...

Pathogen confusion as a strategy for
controlling diseases caused by
Xylella fastidosa
Detection of pathogenic microorganisms

We describe here the detection system for viable pathogenic microorganisms in foods.
Recommended
Satellite RNA

in this chapter covers the symptoms modulation and diseases severity increases and decreases. and role of SiRNA in diseases severity reduction. and also covers the types of SRNA..
Detection techniques of Tomato_Brown_Rugose_Fructose_Virus

It is based on result of detection techniques used in Department of virology at Universitat Politechnica de Valencia, Spain as a part of second semester internship.
Transcriptomics: A Tool for Plant Disease Management

Transcriptomics is one of the omics we can use to manage the plant diseases by modifications in the transcription process
Pathogen confusion as a strategy for controlling diseases caused by Xylella...

Pathogen confusion as a strategy for
controlling diseases caused by
Xylella fastidosa
Detection of pathogenic microorganisms

We describe here the detection system for viable pathogenic microorganisms in foods.
DNA-based methods for bioaerosol analysis

Information for producing phylogenetic/taxonomic libraries of airborne bacteria and fungi. Includes fundamental background information, approaches for sequencing and data analysis, two case studies, and a review of sampling methods
Genetic Engineering of Papaya for Ring Spot Virus

Details about need and methods for Genetic Engineering of Papaya for Ring Spot Virus.
Measuring RNA and DNA biomarkers in liquid biopsies

The interest in liquid biopsies is booming, for obvious reasons: liquid biopsies allow serial follow-up of patients and result in higher patient comfort. Biogazelle offers various customized workflows to fully deploy the biomarker potential from your liquid biopsies.
Assessment of microbial population diversity in polymicrobial research sample...

Assessment of microbial population diversity in polymicrobial research sample...Thermo Fisher Scientific
Analysis of 16S sequences in microbial populations using NGS gives a rapid overview of the community diversity, and is usually performed by sequencing one or two hypervariable regions (V-regions), out of the nine present in the 16S rRNA gene. In this study we compared the community structure of fecal, oral and water microbiomes by analyzing sequences from a single variable region, or from the seven V-regions (V2, V3, V4, V6-7, V8 and V9) included into Ion 16S™ Metagenomics Kit (multi-V analysis)Rapid 16S Next Generation Sequencing for Bacterial Identification in Polymicr...

Rapid 16S Next Generation Sequencing for Bacterial Identification in Polymicr...Thermo Fisher Scientific
In order to identify prokaryotic species in a sample, it is often necessary to culture the sample for hours or days to increase the abundance of bacteria to assayable levels. This often precludes the rapid identification of infectious species.
Furthermore, some species are not easily culturable. We
have developed a facile research method for identifying
bacterial species by 16S ribosomal RNA sequencing on the
Ion Torrent platform. The Ion 16S™ Metagenomics Kit is
designed to PCR amplify the hypervariable regions of the 16S
gene of bacteria. We used this kit to construct libraries from
15 retrospective samples of synovial fluid with various
bacterial species either spiked in or present at collection.
Libraries were sequenced on the Ion PGM™ system and the
data analysis performed using the Ion Reporter™ workflow
which provides an automated analysis solution. Bacteria
present in the samples were correctly identified in samples
containing a single spiked-in species, mixed-species samples,
and in infected samples. Thus, the Ion Torrent™ platform
provides a mechanism for rapidly identifying bacteria that are
present in research samples without culturing.Microbial sequencing

Until recently, the properties and compositions of the microbiota in the planet are still largely a black box. Next generation sequencing (NGS) has proven to be an invaluable tool for investigating diverse environmental and host-associated microbial communities, helping to generate enormous new data sets that can be mined for information on the composition and functional properties of vastly great numbers of microbial communities.
Recombinant Antibody Overview II - Creative Biolabs

It is created by Creative Biolabs who is specialized in providing recombinant antibodies and engineered antibodies. The part two of recombinant antibody overview, we will explain introduction of recombinant antibody ant recombinant antibody expression.
Virus resistance plant, production

introduction
What is virus
What is virus resistance plant
History
Gene use for develop virus resistance plant
Coat protein gene
cDNA of satellite RNA
Defective viral genome
Antisense RNA approach and
Ribozyme – mediated protection
conclusion
References
16S classifier

The diversity of microbial species in a metagenomic study is commonly assessed using 16S rRNA gene sequencing. With the rapid developments in genome sequencing technologies, the focus has shifted towards the sequencing of hypervariable regions of 16S rRNA gene instead of full length gene sequencing. Therefore, 16S Classifier is developed using a machine learning method, Random Forest, for faster and accurate taxonomic classification of short hypervariable regions of 16S rRNA sequence. It displayed precision values of up to 0.91 on training datasets and the precision values of up to 0.98 on the test dataset. On real metagenomic datasets, it showed up to 99.7% accuracy at the phylum level and up to 99.0% accuracy at the genus level. 16S Classifier is available freely at http://metagenomics.iiserb.ac.in/16Sclassifier and http://metabiosys.iiserb.ac.in/16Sclassifier.
Recombinanant dna technology

This presentation consist of Basic Overview of recombinent DNA Technology/ Genetic Engineering
PCR Webinar: COVID-19 (2020)

Polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it to a large enough amount to study in detail. PCR was invented in 1984 by the American biochemist Kary Mullis at Cetus Corporation. It is fundamental to much of genetic testing including analysis of ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory and clinical laboratory research for a broad variety of applications including biomedical research and criminal forensics.
neoplex pcr test kit [genematrix advantages of neo_plex covid-19 Rapid Test K...

This Presentation Briefly Explains All About Neoplex Pcr Test Kit, for Corona Virus Detection. Pcr Covid-19 Rapid test Kits Have Advantages Over Other Manufacturers.
Molecular Basis for Genetic Resistance of Fusarium virguliforme, the Causal A...

This poster was presented at the Undergraduate Research Symposium at the University of Illinois at Urbana-Champaign. It summarizes a semester-long research project I participated in through the Department of Natural Resources and Environmental Sciences.
Genome informed diagnostics

Research investigating the use a genome-informed approach to develop diagnostic tools, for the detection of exotic phytopathogenic bacteria that pose a threat to Australian agriculture.
Virus and viroid testing of solanaceous and cucurbit seed shipments to Australia

Virus and viroid testing of solanaceous and cucurbit seed shipments to AustraliaPlant Biosecurity Cooperative Research Centre
The aim of this research project is to establish Australian developed seed testing protocols as an international standard for the detection of viroids and cucumber green mottle mosaic virus (CGMMV) in seed, and to reduce the risks of contaminated traded seed.More Related Content
What's hot
DNA-based methods for bioaerosol analysis

Information for producing phylogenetic/taxonomic libraries of airborne bacteria and fungi. Includes fundamental background information, approaches for sequencing and data analysis, two case studies, and a review of sampling methods
Genetic Engineering of Papaya for Ring Spot Virus

Details about need and methods for Genetic Engineering of Papaya for Ring Spot Virus.
Measuring RNA and DNA biomarkers in liquid biopsies

The interest in liquid biopsies is booming, for obvious reasons: liquid biopsies allow serial follow-up of patients and result in higher patient comfort. Biogazelle offers various customized workflows to fully deploy the biomarker potential from your liquid biopsies.
Assessment of microbial population diversity in polymicrobial research sample...

Assessment of microbial population diversity in polymicrobial research sample...Thermo Fisher Scientific
Analysis of 16S sequences in microbial populations using NGS gives a rapid overview of the community diversity, and is usually performed by sequencing one or two hypervariable regions (V-regions), out of the nine present in the 16S rRNA gene. In this study we compared the community structure of fecal, oral and water microbiomes by analyzing sequences from a single variable region, or from the seven V-regions (V2, V3, V4, V6-7, V8 and V9) included into Ion 16S™ Metagenomics Kit (multi-V analysis)Rapid 16S Next Generation Sequencing for Bacterial Identification in Polymicr...

Rapid 16S Next Generation Sequencing for Bacterial Identification in Polymicr...Thermo Fisher Scientific
In order to identify prokaryotic species in a sample, it is often necessary to culture the sample for hours or days to increase the abundance of bacteria to assayable levels. This often precludes the rapid identification of infectious species.
Furthermore, some species are not easily culturable. We
have developed a facile research method for identifying
bacterial species by 16S ribosomal RNA sequencing on the
Ion Torrent platform. The Ion 16S™ Metagenomics Kit is
designed to PCR amplify the hypervariable regions of the 16S
gene of bacteria. We used this kit to construct libraries from
15 retrospective samples of synovial fluid with various
bacterial species either spiked in or present at collection.
Libraries were sequenced on the Ion PGM™ system and the
data analysis performed using the Ion Reporter™ workflow
which provides an automated analysis solution. Bacteria
present in the samples were correctly identified in samples
containing a single spiked-in species, mixed-species samples,
and in infected samples. Thus, the Ion Torrent™ platform
provides a mechanism for rapidly identifying bacteria that are
present in research samples without culturing.Microbial sequencing

Until recently, the properties and compositions of the microbiota in the planet are still largely a black box. Next generation sequencing (NGS) has proven to be an invaluable tool for investigating diverse environmental and host-associated microbial communities, helping to generate enormous new data sets that can be mined for information on the composition and functional properties of vastly great numbers of microbial communities.
Recombinant Antibody Overview II - Creative Biolabs

It is created by Creative Biolabs who is specialized in providing recombinant antibodies and engineered antibodies. The part two of recombinant antibody overview, we will explain introduction of recombinant antibody ant recombinant antibody expression.
Virus resistance plant, production

introduction
What is virus
What is virus resistance plant
History
Gene use for develop virus resistance plant
Coat protein gene
cDNA of satellite RNA
Defective viral genome
Antisense RNA approach and
Ribozyme – mediated protection
conclusion
References
16S classifier

The diversity of microbial species in a metagenomic study is commonly assessed using 16S rRNA gene sequencing. With the rapid developments in genome sequencing technologies, the focus has shifted towards the sequencing of hypervariable regions of 16S rRNA gene instead of full length gene sequencing. Therefore, 16S Classifier is developed using a machine learning method, Random Forest, for faster and accurate taxonomic classification of short hypervariable regions of 16S rRNA sequence. It displayed precision values of up to 0.91 on training datasets and the precision values of up to 0.98 on the test dataset. On real metagenomic datasets, it showed up to 99.7% accuracy at the phylum level and up to 99.0% accuracy at the genus level. 16S Classifier is available freely at http://metagenomics.iiserb.ac.in/16Sclassifier and http://metabiosys.iiserb.ac.in/16Sclassifier.
Recombinanant dna technology

This presentation consist of Basic Overview of recombinent DNA Technology/ Genetic Engineering
PCR Webinar: COVID-19 (2020)

Polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it to a large enough amount to study in detail. PCR was invented in 1984 by the American biochemist Kary Mullis at Cetus Corporation. It is fundamental to much of genetic testing including analysis of ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory and clinical laboratory research for a broad variety of applications including biomedical research and criminal forensics.
neoplex pcr test kit [genematrix advantages of neo_plex covid-19 Rapid Test K...

This Presentation Briefly Explains All About Neoplex Pcr Test Kit, for Corona Virus Detection. Pcr Covid-19 Rapid test Kits Have Advantages Over Other Manufacturers.
Molecular Basis for Genetic Resistance of Fusarium virguliforme, the Causal A...

This poster was presented at the Undergraduate Research Symposium at the University of Illinois at Urbana-Champaign. It summarizes a semester-long research project I participated in through the Department of Natural Resources and Environmental Sciences.
What's hot (20)
Measuring RNA and DNA biomarkers in liquid biopsies

Measuring RNA and DNA biomarkers in liquid biopsies
Assessment of microbial population diversity in polymicrobial research sample...

Assessment of microbial population diversity in polymicrobial research sample...
Rapid 16S Next Generation Sequencing for Bacterial Identification in Polymicr...

Rapid 16S Next Generation Sequencing for Bacterial Identification in Polymicr...
Recombinant Antibody Overview II - Creative Biolabs

Recombinant Antibody Overview II - Creative Biolabs
neoplex pcr test kit [genematrix advantages of neo_plex covid-19 Rapid Test K...

neoplex pcr test kit [genematrix advantages of neo_plex covid-19 Rapid Test K...
Molecular Basis for Genetic Resistance of Fusarium virguliforme, the Causal A...

Molecular Basis for Genetic Resistance of Fusarium virguliforme, the Causal A...
More from Plant Biosecurity Cooperative Research Centre
Genome informed diagnostics

Research investigating the use a genome-informed approach to develop diagnostic tools, for the detection of exotic phytopathogenic bacteria that pose a threat to Australian agriculture.
Virus and viroid testing of solanaceous and cucurbit seed shipments to Australia

Virus and viroid testing of solanaceous and cucurbit seed shipments to AustraliaPlant Biosecurity Cooperative Research Centre
The aim of this research project is to establish Australian developed seed testing protocols as an international standard for the detection of viroids and cucumber green mottle mosaic virus (CGMMV) in seed, and to reduce the risks of contaminated traded seed.Psyllid microflora: Implications for liberibacter disease surveillance and pe...

Psyllid microflora: Implications for liberibacter disease surveillance and pe...Plant Biosecurity Cooperative Research Centre
This research is improving the understanding of the microflora of Australian psyllids, for better biosecurity preparedness and response management.Challenges with implementing a new diagnostic platform in Post Entry Quarantine

Challenges with implementing a new diagnostic platform in Post Entry QuarantinePlant Biosecurity Cooperative Research Centre
The diagnosis of viral pathogens is a crucial component of plant biosecurity surveillance and preventing the introduction of exotic plant viruses and viroids at the border. Existing quarantine procedures can be time-consuming and require detailed knowledge of potential infecting viral pathogens. Currently, imported plants can spend as long as two years in quarantine, with associated costs.
To simplify the post-entry quarantine process researchers have developed a plant diagnostic toolkit for plant viruses and viroids. The toolkit takes advantage of the natural antiviral system of plants, using small RNA next generation sequencing (sRNA-seq) technology to detect nearly all known viruses and viroids in a single test. The new test, and associated toolkit, will reduce the time imported plant material spends in Australia’s quarantine system while improving accuracy of detection in a single sRNA-seq experiment.Creating an enabling environment for fruit fly area-wide management

Creating an enabling environment for fruit fly area-wide managementPlant Biosecurity Cooperative Research Centre
This research has developed recommendations for stakeholders involved in area-wide management of fruit fly, including social and institutional requirements.Building resilience in Indigenous communities through engagement

Building resilience in Indigenous communities through engagementPlant Biosecurity Cooperative Research Centre
This project aims to build the ability of indigenous communities (Maori and Aboriginal), regulatory authorities and industries to better manage the impact of biosecurity threats. Models have been developed for Indigenous engagement.Building collaboration to enable social innovation in biosecurity systems

Building collaboration to enable social innovation in biosecurity systemsPlant Biosecurity Cooperative Research Centre
This social biosecurity project, aims to improve plant biosecurity management by developing the capacity of regional and remote communities to engage in biosecurity surveillance activities.Design and Evaluation of Targeted Biosecurity Surveillance Systems

Design and Evaluation of Targeted Biosecurity Surveillance SystemsPlant Biosecurity Cooperative Research Centre
Surveillance systems are an essential component of biosecurity. Design of biosecurity surveillance systems may include designs of grids of static traps, plans for field sampling, or deployment of potentially "game-changing" mobile trap technology. The aim of these systems is to achieve defined detection objectives, (e.g. early detection, supporting area-freedom status) at minimum cost. This project will develop and apply statistically-based surveillance systems that account for organism biology, trap behaviour and landscape characteristics.Rethinking biosecurity inspections using statistical modelling and simulation...

Rethinking biosecurity inspections using statistical modelling and simulation...Plant Biosecurity Cooperative Research Centre
Ships arriving in Australia may have visited multiple ports along the way. These complex pathways present opportunities for pest species, such as the Asian Gypsy Moth, to arrive into Australia from indirect routes. Understanding those pathways that link Australia directly or indirectly to countries in which a pest or disease occurs is necessary to identify arriving ships with the highest likelihood of carrying hitchhiker species. This project proposes to address three important questions:
1. What general shipping pathways pose the greatest risk?
2. How to make decisions regarding what ships to search?
3. How much inspection to conduct?Plant pest impacts: a common set of metrics

This research project is collecting data on past pest invasions in both Australia and New Zealand, in order to identify common patterns in plant biosecurity pests.
Natural dispersal as a biosecurity risk -are we prepared?

Natural dispersal as a biosecurity risk -are we prepared?Plant Biosecurity Cooperative Research Centre
Long distance natural (wind-assisted) dispersal of exotic plant pests and pathogens into Australia, is a very real and underestimated, biosecurity risk.Developing tools for in-field surveillance of pathogens

Developing tools for in-field surveillance of pathogensPlant Biosecurity Cooperative Research Centre
This research will investigate technologies to enable the development of spore traps capable of in-field detection, and identification, of specific biosecurity threats.New tools for field grains surveillance

New tools for field grains surveillance and diagnostics of high priority exotic plant pests.
Tomato potato psyllid and Liberibacter ecology

Research project about tomato potato psyllid and Liberibacter ecology.
Smart-trap design and deployment strategies

Smart-trap design and deployment strategies, for Hessian fly.
Session 1: A risk-return prioritisation tool for global trade inspections

Session 1: A risk-return prioritisation tool for global trade inspectionsPlant Biosecurity Cooperative Research Centre
The spread of invasive species continues to provide significant challenges to those government biosecurity agencies charged with protecting a country’s borders. In an increasingly connected world, these invasive species are potentially able to spread further and more rapidly. Human mediated pathways such as ships and airlines are the most obvious ways in which invasive species can be spread. Direct routes from one port to another are currently monitored, but indirect pathways,
in which a ship picks up an invasive species and then travels to a number of different locations before arriving at the final destination, present more challenging scenarios. For the Australian Government Department of Agriculture, one particular concern is for ships arriving into Australia carrying viable eggs of the Asian gypsy moth (Lymantria dispar). We are developing a real time tool that will analyse the pathways for incoming ships and determine the likelihood the ship could be carrying viable eggs.Session 10: Better biosecurity risk management through collaborative planning...

Session 10: Better biosecurity risk management through collaborative planning...Plant Biosecurity Cooperative Research Centre
Planning and decision making to manage plant biosecurity risks is inherently complex, often contentious, involves unknowns and uncertainty, and needs to be adaptable to rapidly changing situations. The aim of this project is to develop a collaborative planning and shared decision making
framework that will result in better and faster decisions to respond more quickly to plant biosecurity risks, resulting in reduced impacts and costs, and more equitable and favourable outcomes for stakeholders and affected parties.Session 10: Building resilience in indigenous communities through engagement

Session 10: Building resilience in indigenous communities through engagementPlant Biosecurity Cooperative Research Centre
Biosecurity issues impact on key crops and environmental values across NZ and Australia. A key outcome for the project team will be the ability of indigenous communities, and relevant regulatory authorities and industries, to better manage the social, environmental and economic impacts of biosecurity threats, and to participate in biosecurity strategies through improved bicultural engagement models that build empowerment and ownership in indigenous communities and their response to those threats. The teams have developed an engagement model adapted to the indigenous peoples and their communities of each country.Session 10: Creating an enabling environment for industry-driven fruit fly ar...

Session 10: Creating an enabling environment for industry-driven fruit fly ar...Plant Biosecurity Cooperative Research Centre
This presentation reports on research that explored how to strengthen support to local industries pursuing fruit fly area-wide management (AWM).Session 10: Participating in fruit fly control: Why being motivated is not go...

Session 10: Participating in fruit fly control: Why being motivated is not go...Plant Biosecurity Cooperative Research Centre
The results of a baseline study on motivation and incentives involved in the decisions to control fruit fly highlight the variability of motivations within demographic groups.More from Plant Biosecurity Cooperative Research Centre (20)
Virus and viroid testing of solanaceous and cucurbit seed shipments to Australia

Virus and viroid testing of solanaceous and cucurbit seed shipments to Australia
Psyllid microflora: Implications for liberibacter disease surveillance and pe...

Psyllid microflora: Implications for liberibacter disease surveillance and pe...
Challenges with implementing a new diagnostic platform in Post Entry Quarantine

Challenges with implementing a new diagnostic platform in Post Entry Quarantine
Creating an enabling environment for fruit fly area-wide management

Creating an enabling environment for fruit fly area-wide management
Building resilience in Indigenous communities through engagement

Building resilience in Indigenous communities through engagement
Building collaboration to enable social innovation in biosecurity systems

Building collaboration to enable social innovation in biosecurity systems
Design and Evaluation of Targeted Biosecurity Surveillance Systems

Design and Evaluation of Targeted Biosecurity Surveillance Systems
Rethinking biosecurity inspections using statistical modelling and simulation...

Rethinking biosecurity inspections using statistical modelling and simulation...
Natural dispersal as a biosecurity risk -are we prepared?

Natural dispersal as a biosecurity risk -are we prepared?
Developing tools for in-field surveillance of pathogens

Developing tools for in-field surveillance of pathogens
Session 1: A risk-return prioritisation tool for global trade inspections

Session 1: A risk-return prioritisation tool for global trade inspections
Session 10: Better biosecurity risk management through collaborative planning...

Session 10: Better biosecurity risk management through collaborative planning...
Session 10: Building resilience in indigenous communities through engagement

Session 10: Building resilience in indigenous communities through engagement
Session 10: Creating an enabling environment for industry-driven fruit fly ar...

Session 10: Creating an enabling environment for industry-driven fruit fly ar...
Session 10: Participating in fruit fly control: Why being motivated is not go...

Session 10: Participating in fruit fly control: Why being motivated is not go...
Recently uploaded
Orion Air Quality Monitoring Systems - CWS

Professional air quality monitoring systems provide immediate, on-site data for analysis, compliance, and decision-making.
Monitor common gases, weather parameters, particulates.
Multi-source connectivity as the driver of solar wind variability in the heli...

The ambient solar wind that flls the heliosphere originates from multiple
sources in the solar corona and is highly structured. It is often described
as high-speed, relatively homogeneous, plasma streams from coronal
holes and slow-speed, highly variable, streams whose source regions are
under debate. A key goal of ESA/NASA’s Solar Orbiter mission is to identify
solar wind sources and understand what drives the complexity seen in the
heliosphere. By combining magnetic feld modelling and spectroscopic
techniques with high-resolution observations and measurements, we show
that the solar wind variability detected in situ by Solar Orbiter in March
2022 is driven by spatio-temporal changes in the magnetic connectivity to
multiple sources in the solar atmosphere. The magnetic feld footpoints
connected to the spacecraft moved from the boundaries of a coronal hole
to one active region (12961) and then across to another region (12957). This
is refected in the in situ measurements, which show the transition from fast
to highly Alfvénic then to slow solar wind that is disrupted by the arrival of
a coronal mass ejection. Our results describe solar wind variability at 0.5 au
but are applicable to near-Earth observatories.
THE IMPORTANCE OF MARTIAN ATMOSPHERE SAMPLE RETURN.

The return of a sample of near-surface atmosphere from Mars would facilitate answers to several first-order science questions surrounding the formation and evolution of the planet. One of the important aspects of terrestrial planet formation in general is the role that primary atmospheres played in influencing the chemistry and structure of the planets and their antecedents. Studies of the martian atmosphere can be used to investigate the role of a primary atmosphere in its history. Atmosphere samples would also inform our understanding of the near-surface chemistry of the planet, and ultimately the prospects for life. High-precision isotopic analyses of constituent gases are needed to address these questions, requiring that the analyses are made on returned samples rather than in situ.
Astronomy Update- Curiosity’s exploration of Mars _ Local Briefs _ leadertele...

Article written for leader telegram
Earliest Galaxies in the JADES Origins Field: Luminosity Function and Cosmic ...

We characterize the earliest galaxy population in the JADES Origins Field (JOF), the deepest
imaging field observed with JWST. We make use of the ancillary Hubble optical images (5 filters
spanning 0.4−0.9µm) and novel JWST images with 14 filters spanning 0.8−5µm, including 7 mediumband filters, and reaching total exposure times of up to 46 hours per filter. We combine all our data
at > 2.3µm to construct an ultradeep image, reaching as deep as ≈ 31.4 AB mag in the stack and
30.3-31.0 AB mag (5σ, r = 0.1” circular aperture) in individual filters. We measure photometric
redshifts and use robust selection criteria to identify a sample of eight galaxy candidates at redshifts
z = 11.5 − 15. These objects show compact half-light radii of R1/2 ∼ 50 − 200pc, stellar masses of
M⋆ ∼ 107−108M⊙, and star-formation rates of SFR ∼ 0.1−1 M⊙ yr−1
. Our search finds no candidates
at 15 < z < 20, placing upper limits at these redshifts. We develop a forward modeling approach to
infer the properties of the evolving luminosity function without binning in redshift or luminosity that
marginalizes over the photometric redshift uncertainty of our candidate galaxies and incorporates the
impact of non-detections. We find a z = 12 luminosity function in good agreement with prior results,
and that the luminosity function normalization and UV luminosity density decline by a factor of ∼ 2.5
from z = 12 to z = 14. We discuss the possible implications of our results in the context of theoretical
models for evolution of the dark matter halo mass function.
Structures and textures of metamorphic rocks

It is useful for the Under Graduating students for easy understanding and it's useful for the exam preparations.
filosofia boliviana introducción jsjdjd.pptx

La filosofía boliviana y la búsqueda por construir pensamientos propios
insect taxonomy importance systematics and classification

documents provide information about insect classification and taxonomy of insect
extra-chromosomal-inheritance[1].pptx.pdfpdf![extra-chromosomal-inheritance[1].pptx.pdfpdf](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
![extra-chromosomal-inheritance[1].pptx.pdfpdf](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
Slide 1: Title Slide
Extrachromosomal Inheritance
Slide 2: Introduction to Extrachromosomal Inheritance
Definition: Extrachromosomal inheritance refers to the transmission of genetic material that is not found within the nucleus.
Key Components: Involves genes located in mitochondria, chloroplasts, and plasmids.
Slide 3: Mitochondrial Inheritance
Mitochondria: Organelles responsible for energy production.
Mitochondrial DNA (mtDNA): Circular DNA molecule found in mitochondria.
Inheritance Pattern: Maternally inherited, meaning it is passed from mothers to all their offspring.
Diseases: Examples include Leber’s hereditary optic neuropathy (LHON) and mitochondrial myopathy.
Slide 4: Chloroplast Inheritance
Chloroplasts: Organelles responsible for photosynthesis in plants.
Chloroplast DNA (cpDNA): Circular DNA molecule found in chloroplasts.
Inheritance Pattern: Often maternally inherited in most plants, but can vary in some species.
Examples: Variegation in plants, where leaf color patterns are determined by chloroplast DNA.
Slide 5: Plasmid Inheritance
Plasmids: Small, circular DNA molecules found in bacteria and some eukaryotes.
Features: Can carry antibiotic resistance genes and can be transferred between cells through processes like conjugation.
Significance: Important in biotechnology for gene cloning and genetic engineering.
Slide 6: Mechanisms of Extrachromosomal Inheritance
Non-Mendelian Patterns: Do not follow Mendel’s laws of inheritance.
Cytoplasmic Segregation: During cell division, organelles like mitochondria and chloroplasts are randomly distributed to daughter cells.
Heteroplasmy: Presence of more than one type of organellar genome within a cell, leading to variation in expression.
Slide 7: Examples of Extrachromosomal Inheritance
Four O’clock Plant (Mirabilis jalapa): Shows variegated leaves due to different cpDNA in leaf cells.
Petite Mutants in Yeast: Result from mutations in mitochondrial DNA affecting respiration.
Slide 8: Importance of Extrachromosomal Inheritance
Evolution: Provides insight into the evolution of eukaryotic cells.
Medicine: Understanding mitochondrial inheritance helps in diagnosing and treating mitochondrial diseases.
Agriculture: Chloroplast inheritance can be used in plant breeding and genetic modification.
Slide 9: Recent Research and Advances
Gene Editing: Techniques like CRISPR-Cas9 are being used to edit mitochondrial and chloroplast DNA.
Therapies: Development of mitochondrial replacement therapy (MRT) for preventing mitochondrial diseases.
Slide 10: Conclusion
Summary: Extrachromosomal inheritance involves the transmission of genetic material outside the nucleus and plays a crucial role in genetics, medicine, and biotechnology.
Future Directions: Continued research and technological advancements hold promise for new treatments and applications.
Slide 11: Questions and Discussion
Invite Audience: Open the floor for any questions or further discussion on the topic.
FAIR & AI Ready KGs for Explainable Predictions

The increased availability of biomedical data, particularly in the public domain, offers the opportunity to better understand human health and to develop effective therapeutics for a wide range of unmet medical needs. However, data scientists remain stymied by the fact that data remain hard to find and to productively reuse because data and their metadata i) are wholly inaccessible, ii) are in non-standard or incompatible representations, iii) do not conform to community standards, and iv) have unclear or highly restricted terms and conditions that preclude legitimate reuse. These limitations require a rethink on data can be made machine and AI-ready - the key motivation behind the FAIR Guiding Principles. Concurrently, while recent efforts have explored the use of deep learning to fuse disparate data into predictive models for a wide range of biomedical applications, these models often fail even when the correct answer is already known, and fail to explain individual predictions in terms that data scientists can appreciate. These limitations suggest that new methods to produce practical artificial intelligence are still needed.
In this talk, I will discuss our work in (1) building an integrative knowledge infrastructure to prepare FAIR and "AI-ready" data and services along with (2) neurosymbolic AI methods to improve the quality of predictions and to generate plausible explanations. Attention is given to standards, platforms, and methods to wrangle knowledge into simple, but effective semantic and latent representations, and to make these available into standards-compliant and discoverable interfaces that can be used in model building, validation, and explanation. Our work, and those of others in the field, creates a baseline for building trustworthy and easy to deploy AI models in biomedicine.
Bio
Dr. Michel Dumontier is the Distinguished Professor of Data Science at Maastricht University, founder and executive director of the Institute of Data Science, and co-founder of the FAIR (Findable, Accessible, Interoperable and Reusable) data principles. His research explores socio-technological approaches for responsible discovery science, which includes collaborative multi-modal knowledge graphs, privacy-preserving distributed data mining, and AI methods for drug discovery and personalized medicine. His work is supported through the Dutch National Research Agenda, the Netherlands Organisation for Scientific Research, Horizon Europe, the European Open Science Cloud, the US National Institutes of Health, and a Marie-Curie Innovative Training Network. He is the editor-in-chief for the journal Data Science and is internationally recognized for his contributions in bioinformatics, biomedical informatics, and semantic technologies including ontologies and linked data.
Comparative structure of adrenal gland in vertebrates

Adrenal gland comparative structures in vertebrates
Citrus Greening Disease and its Management

Citrus Greening was one of the major causes of decline in the citrus production. So, effective management cultural practices should be incorporated
Recently uploaded (20)
Multi-source connectivity as the driver of solar wind variability in the heli...

Multi-source connectivity as the driver of solar wind variability in the heli...
ESR_factors_affect-clinic significance-Pathysiology.pptx

ESR_factors_affect-clinic significance-Pathysiology.pptx
THE IMPORTANCE OF MARTIAN ATMOSPHERE SAMPLE RETURN.

THE IMPORTANCE OF MARTIAN ATMOSPHERE SAMPLE RETURN.
Astronomy Update- Curiosity’s exploration of Mars _ Local Briefs _ leadertele...

Astronomy Update- Curiosity’s exploration of Mars _ Local Briefs _ leadertele...
Earliest Galaxies in the JADES Origins Field: Luminosity Function and Cosmic ...

Earliest Galaxies in the JADES Origins Field: Luminosity Function and Cosmic ...
Circulatory system_ Laplace law. Ohms law.reynaults law,baro-chemo-receptors-...

Circulatory system_ Laplace law. Ohms law.reynaults law,baro-chemo-receptors-...
erythropoiesis-I_mechanism& clinical significance.pptx

erythropoiesis-I_mechanism& clinical significance.pptx
insect taxonomy importance systematics and classification

insect taxonomy importance systematics and classification
Comparative structure of adrenal gland in vertebrates

Comparative structure of adrenal gland in vertebrates
Session 3: High throughput analysis of chemical and genetic factors that influence phosphine toxicity and resistance
- 1. biosecurity built on science A new system using high – resolution photo scanners. Enabling rapid, high– throughput analysis of C. elegans toxicology For example: Phosphine resistance granted by DLD suppression is dependant on the master stress regulator DAF-16 500ppm 0 10 20 30 40 50 60 70 80 90 100 Phosphine parts per million %survivors Daf16 mutant / DLD suppressed Daf16 mutant Wild type / DLD suppressed Wild type Daf16 upregulated / DLD suppressed Daf16 upregulated Capable of automatically counting tens of thousands of individual C. elegans at once. High throughput C. elegans phenotype analysis and genetic influences on phosphine resistance (Timothy Puckering, University of Queensland)
