Embed presentation
Download to read offline




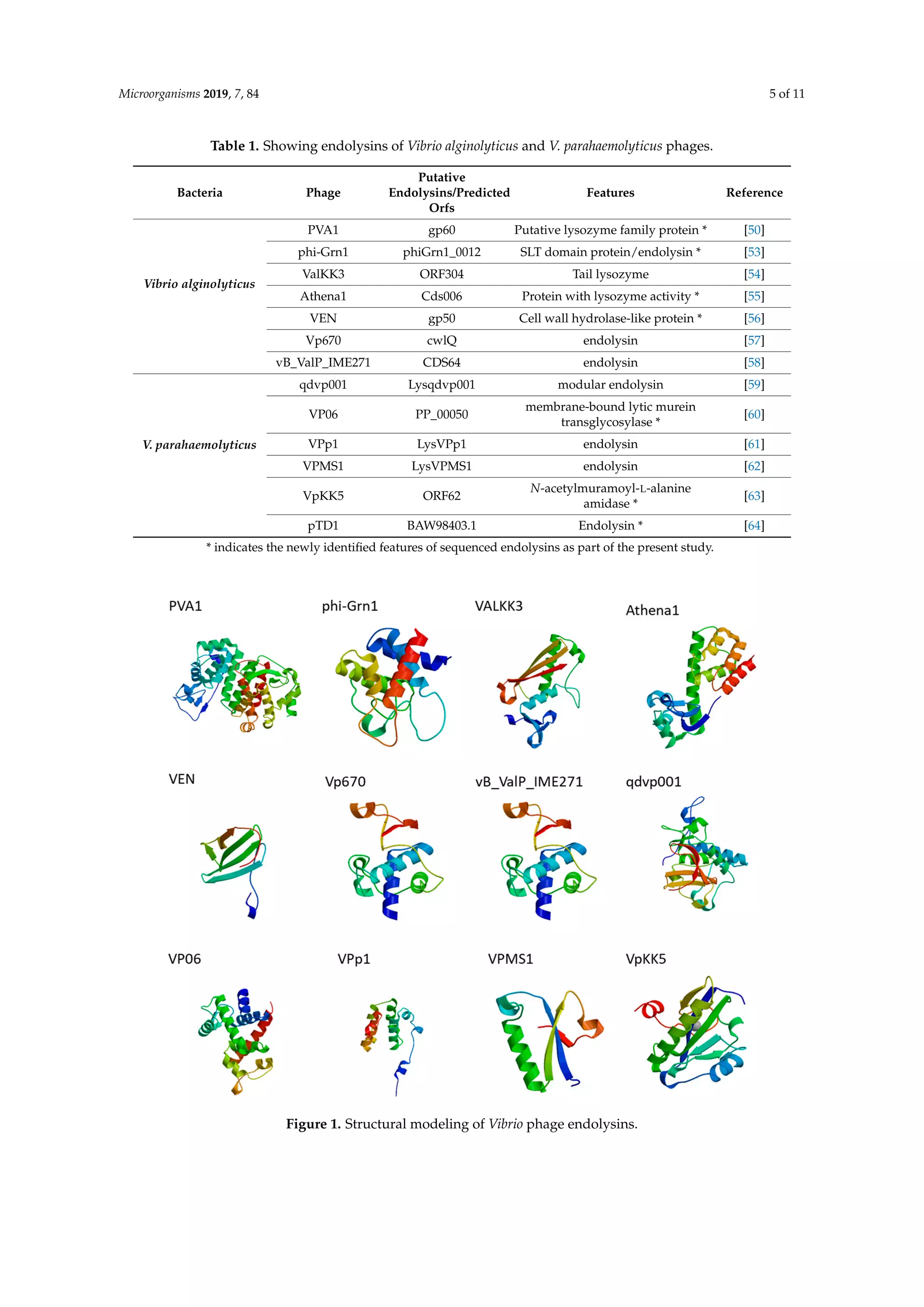







This document reviews the potential of bacteriophage-derived endolysins as antimicrobial agents against the multidrug-resistant pathogens Vibrio alginolyticus and Vibrio parahaemolyticus. It highlights the issues of antibiotic resistance in these bacteria, which pose significant health risks due to the reliance on antibiotics in aquaculture. The study emphasizes the promising application of endolysins in combating these pathogens, underlining their mechanisms, characteristics, and the urgent need for alternative treatments in aquaculture settings.










