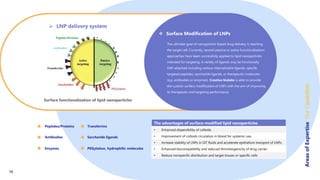
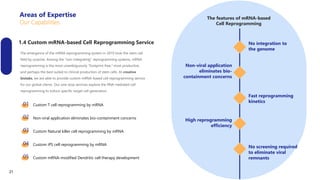
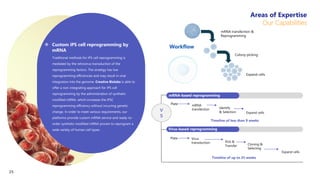
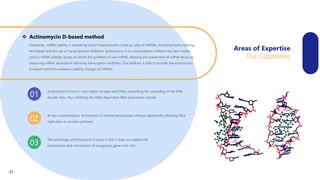
Creative Biolabs offers a comprehensive range of high-quality mRNA services, including custom synthesis, modification, delivery systems, and cell reprogramming to support genomics research and drug development. Their team of over 200 scientists utilizes state-of-the-art platforms to provide tailored solutions and ensure reliable mRNA products through stringent quality control measures. The company's innovative approaches include lipid nanoparticle delivery systems and various mRNA modification strategies to enhance stability and translation efficiency.