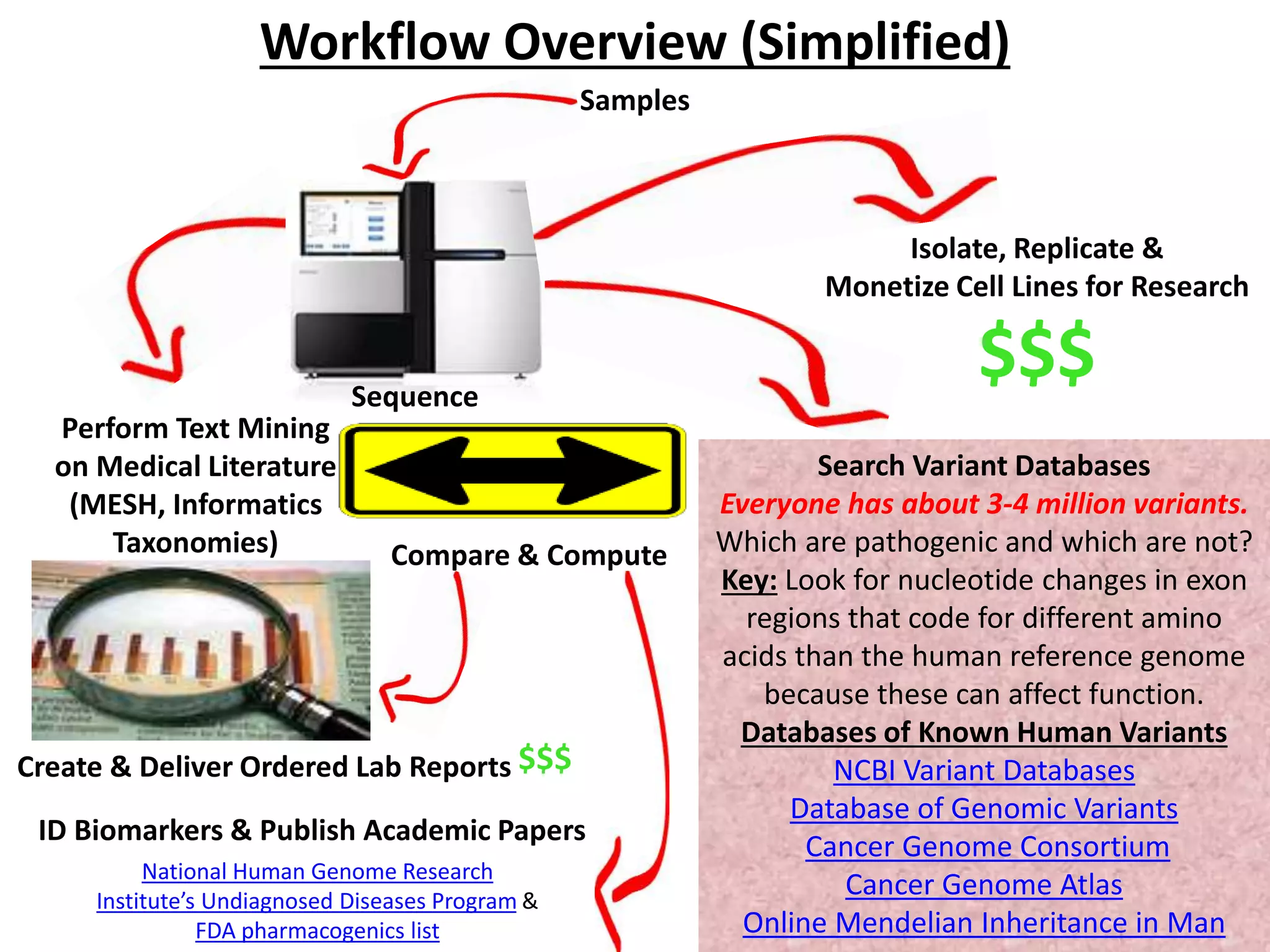
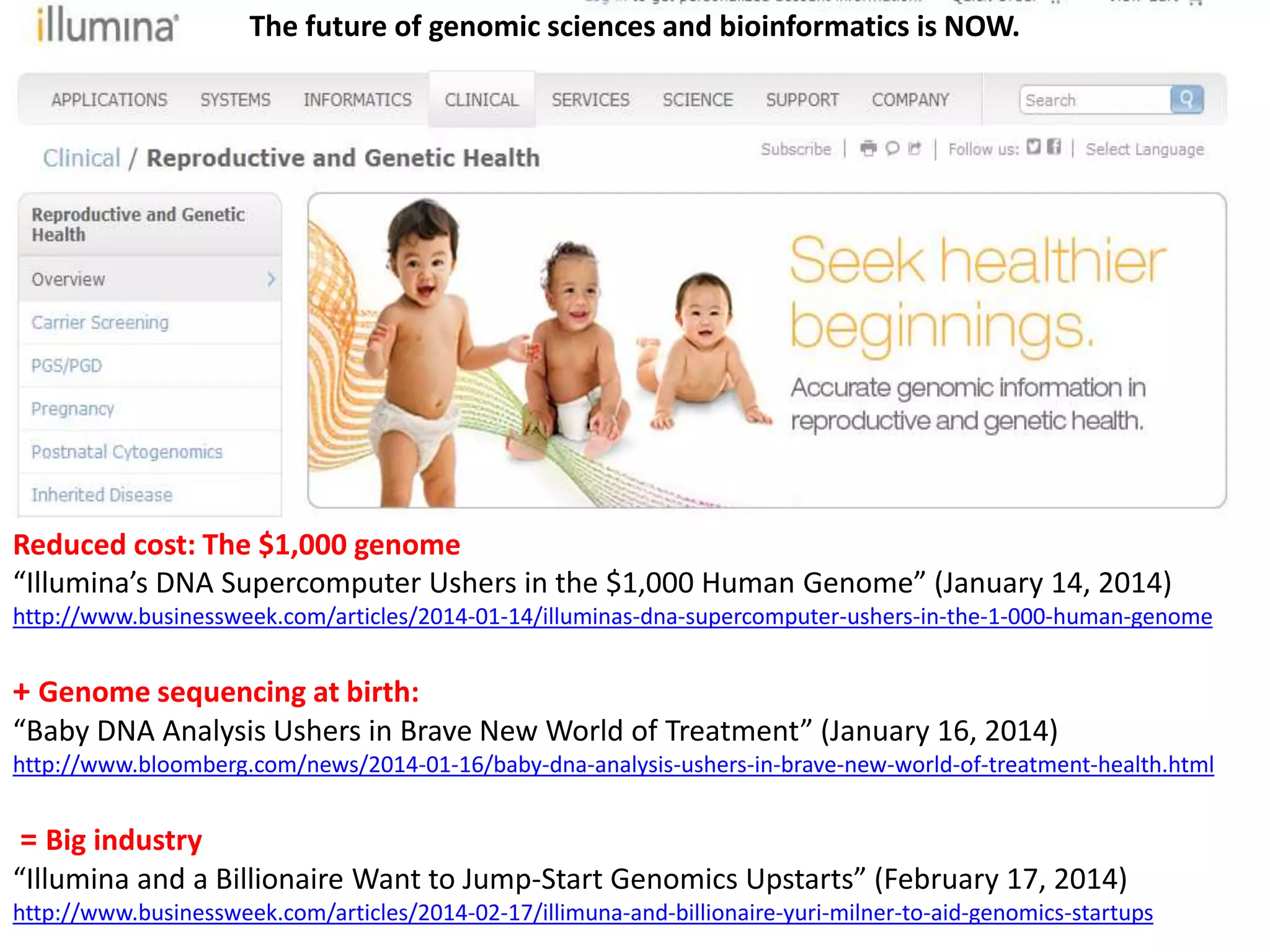
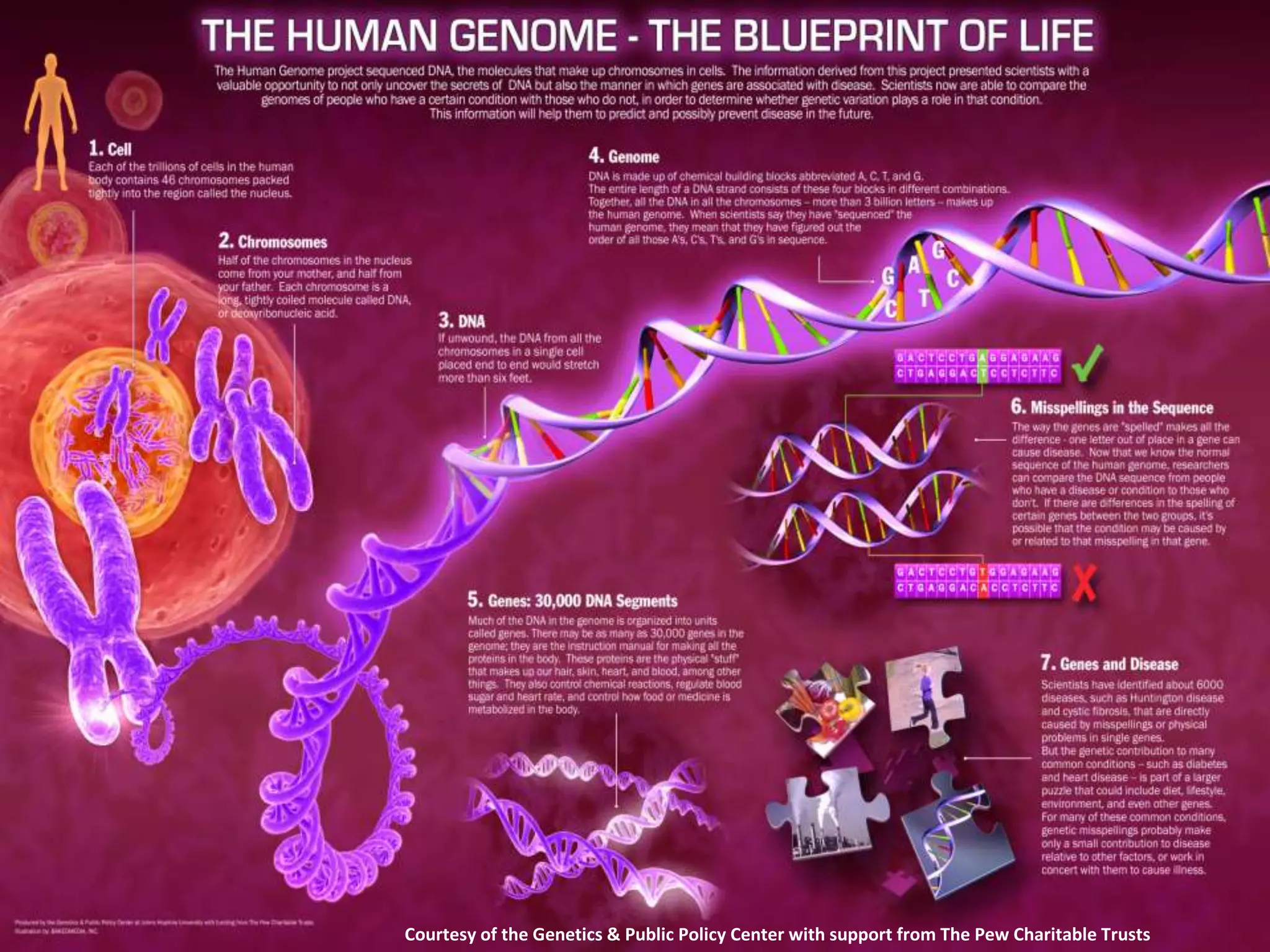
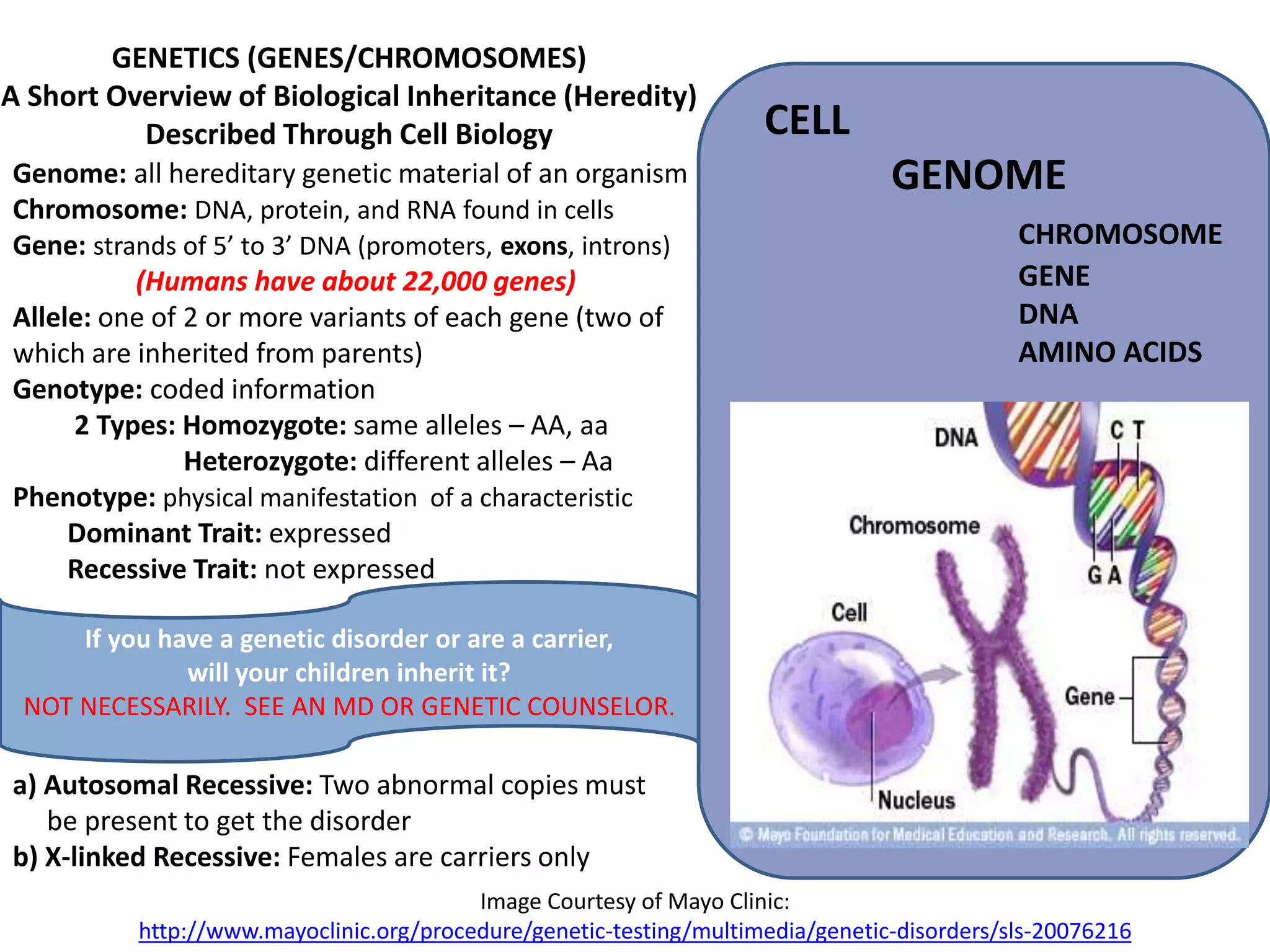
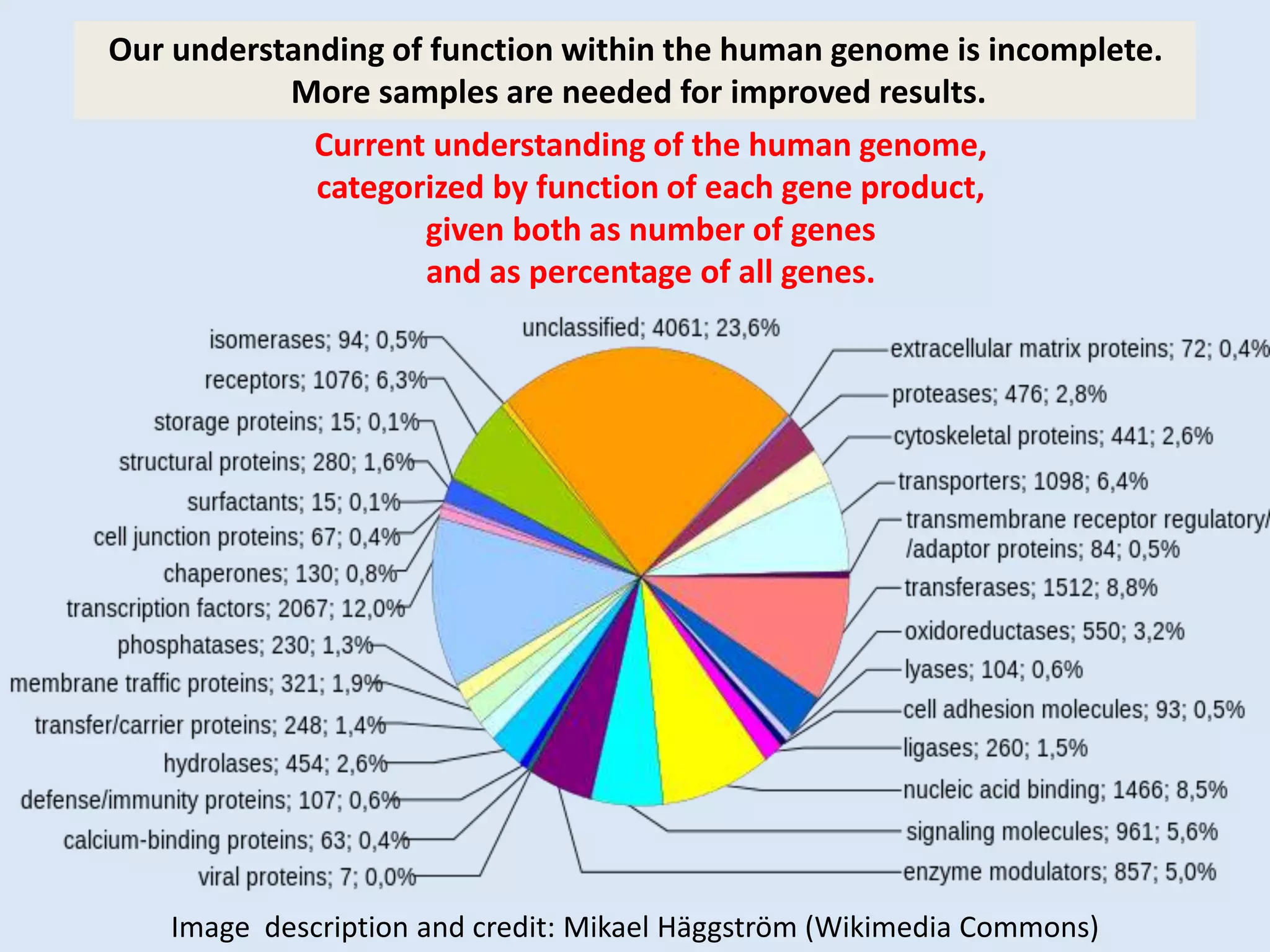
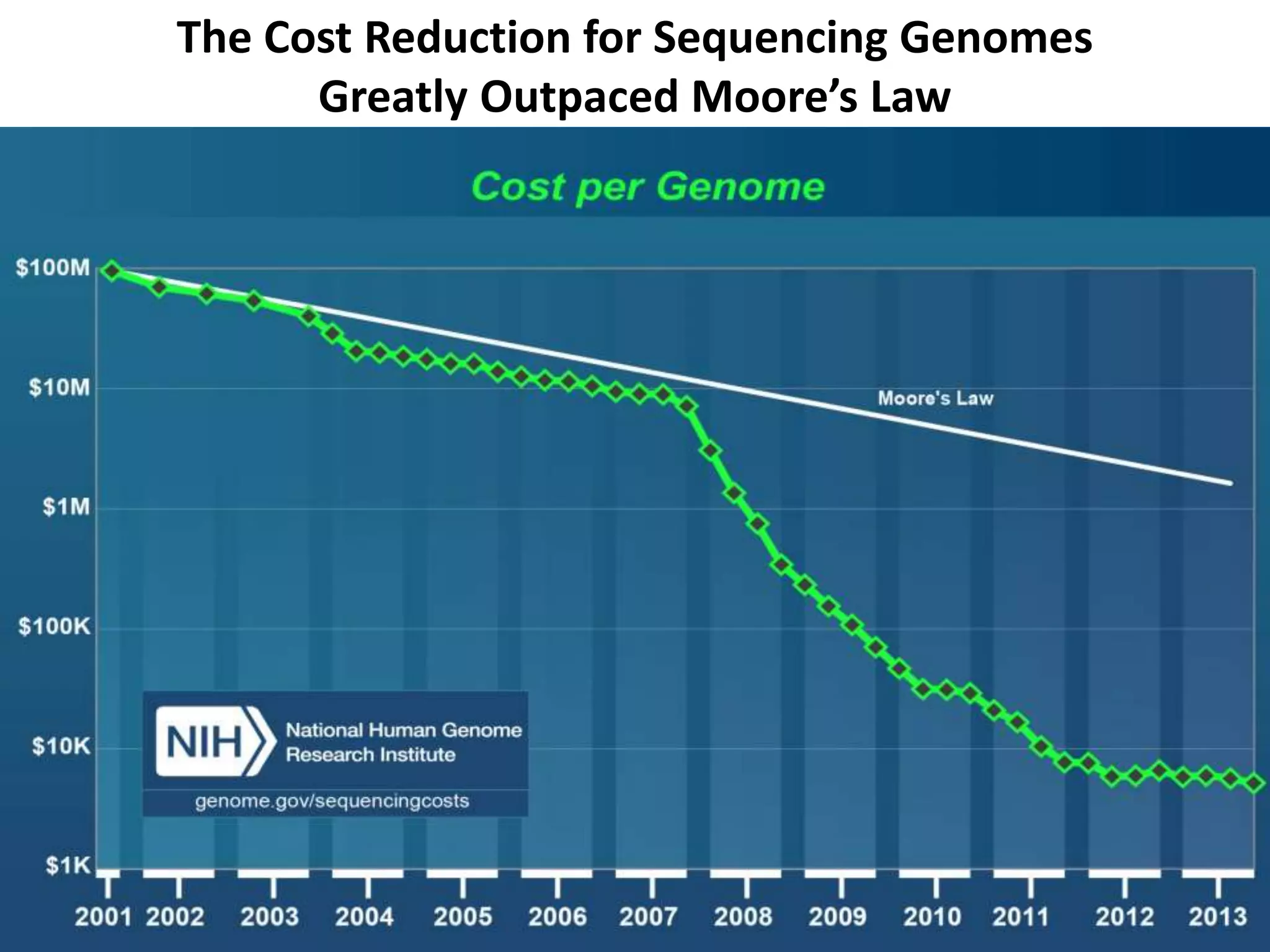
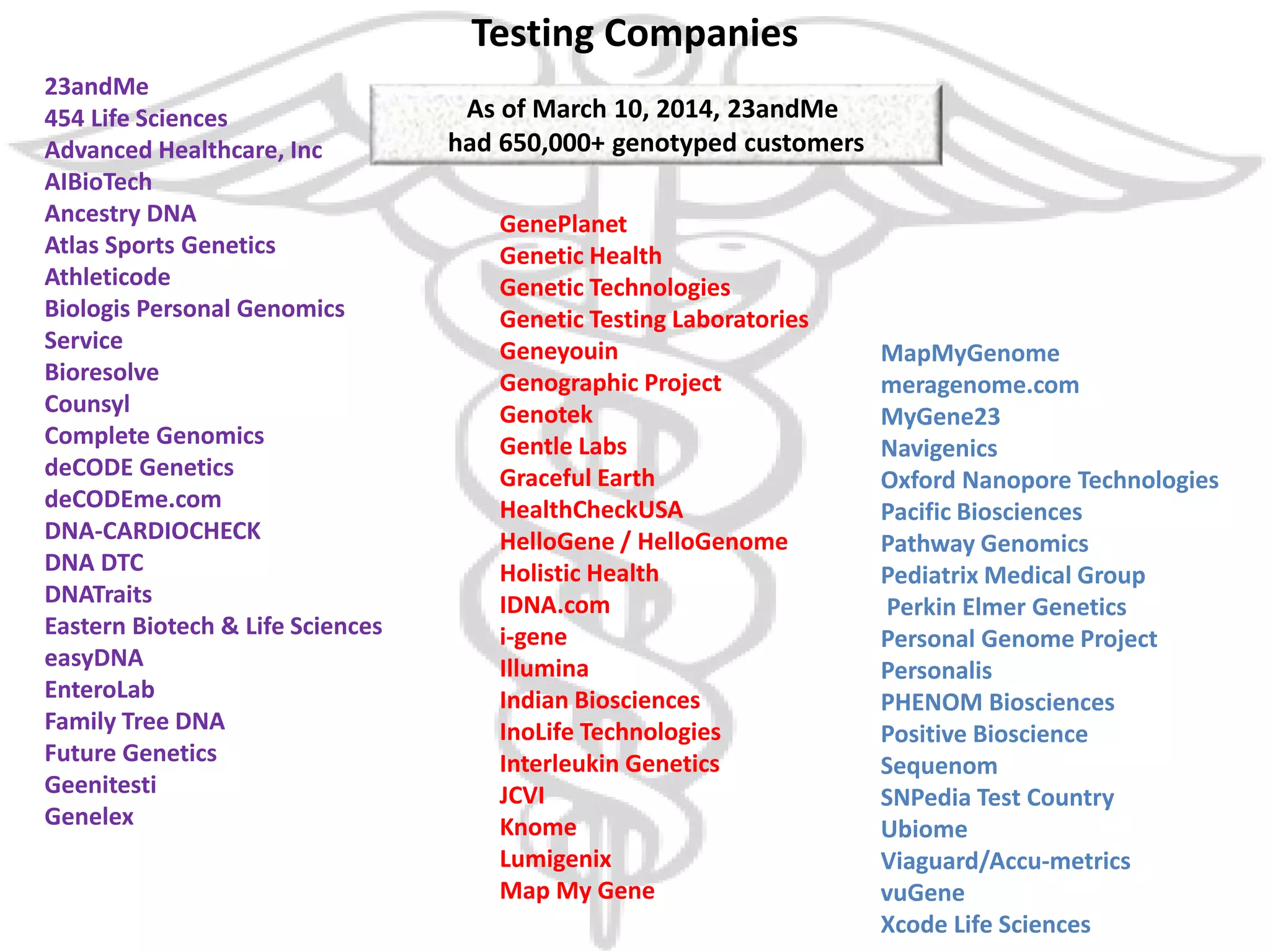
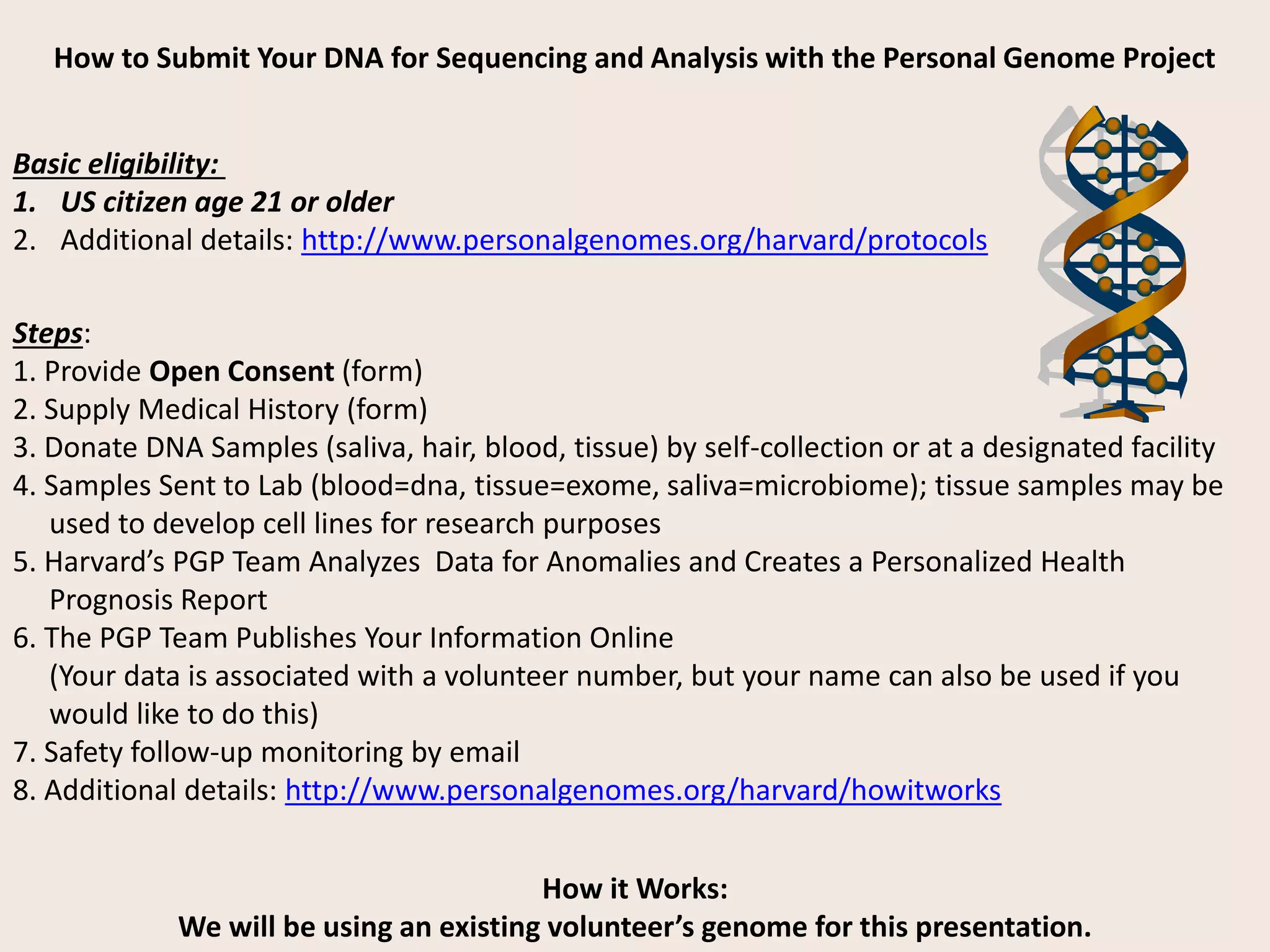
The document presents an overview of genetic inheritance, bioinformatics, and the implications of personal genome sequencing, emphasizing advancements in medical technology and personalized medicine. It discusses the impact of genetic and epigenetic factors on disease, the importance of genetic counseling, and the ethical considerations surrounding genetic testing. Additionally, it highlights various bioinformatics tools used for analyzing genetic data and the role of emerging industries in genomic sciences.















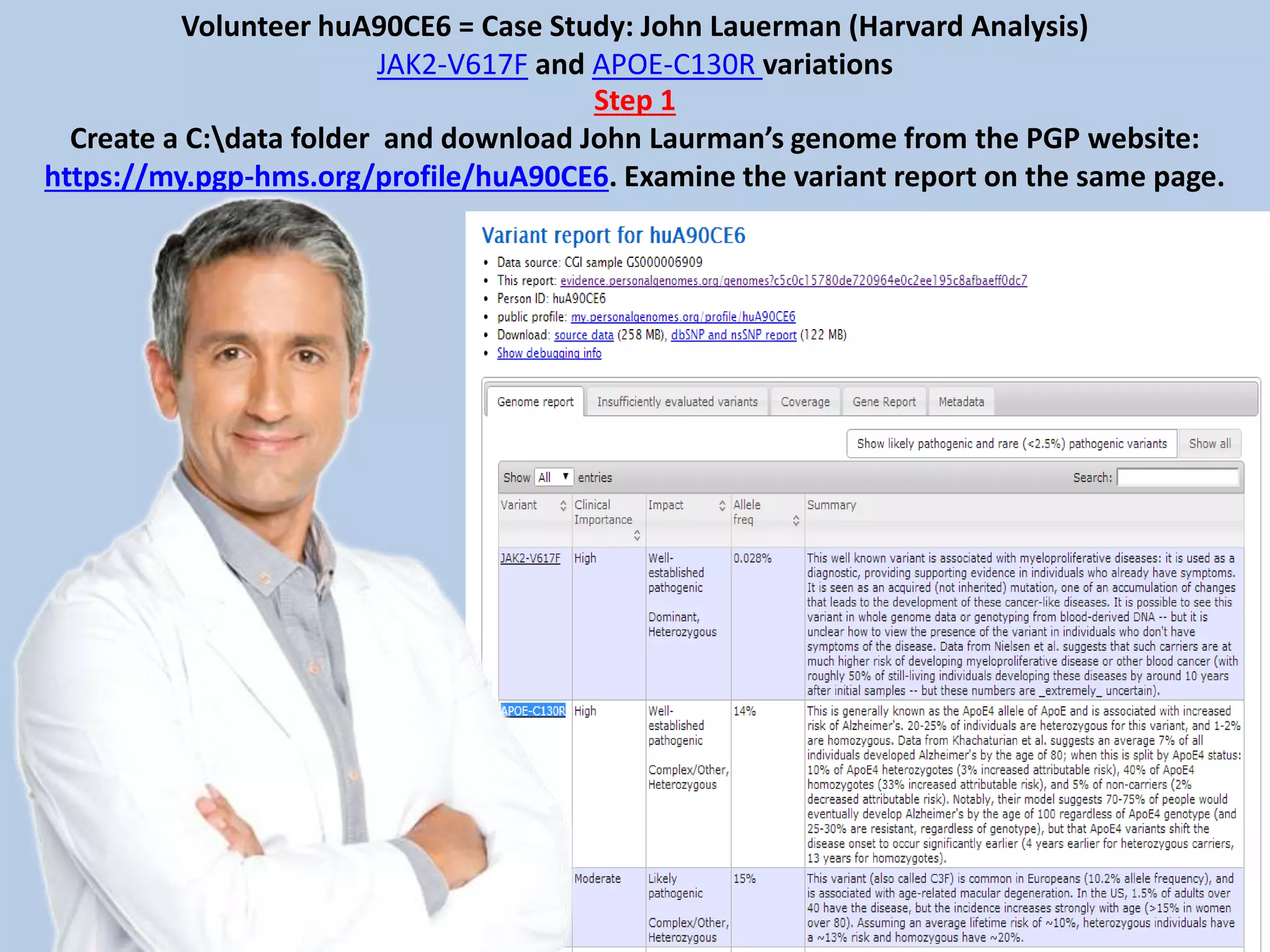

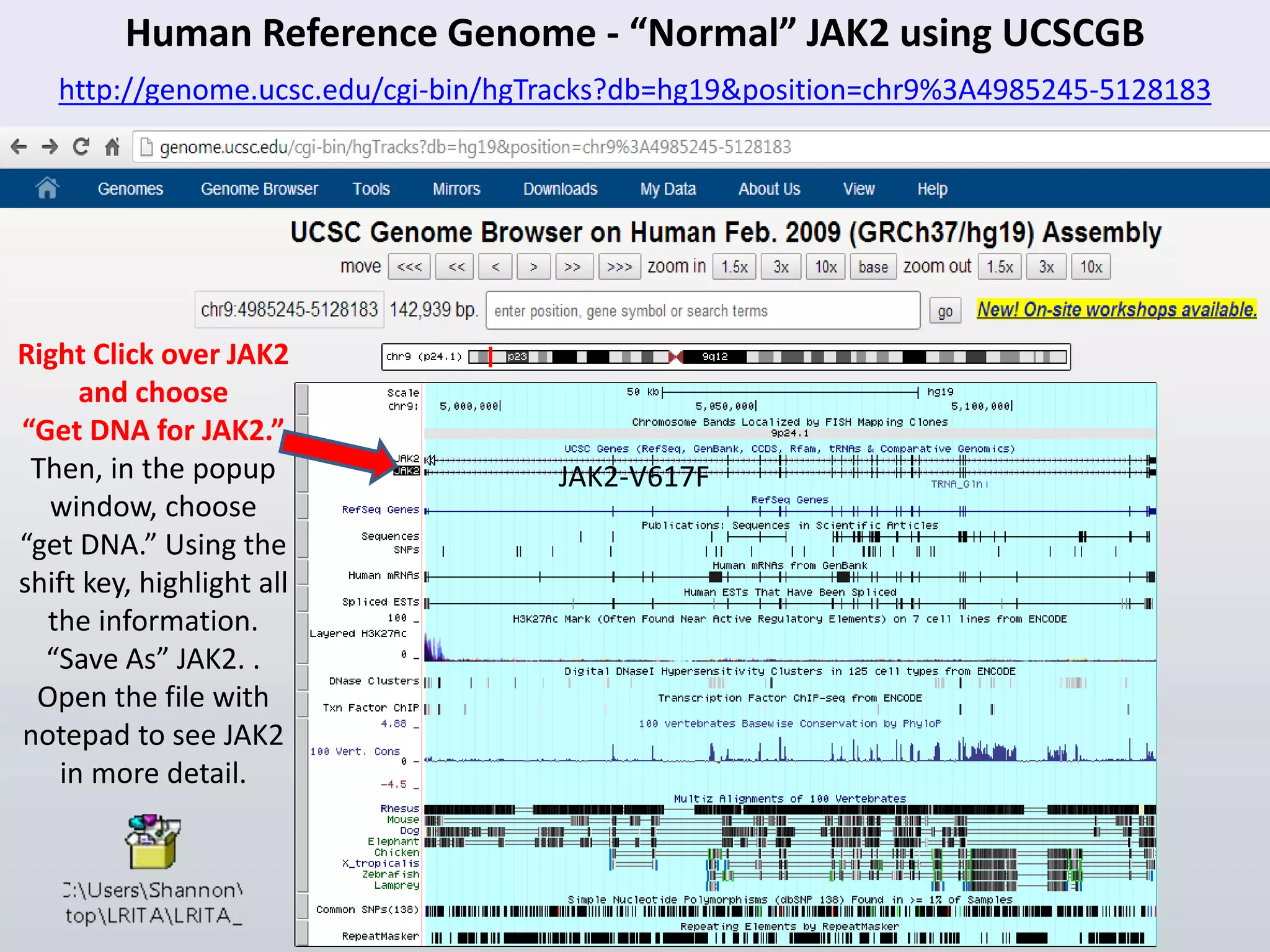

![Volunteer huA90CE6 -- Case Study: John Lauerman (Looking Closer with PGA & PyMOL)
Step 1
Download and install Python https://www.python.org/download/releases/2.7.5
Windows X86-64 MSI Installer (2.7.5) [1] (sig)
Step 2
Download and install the PyMOL extension for a free 3D molecule viewer
http://www.lfd.uci.edu/~gohlke/pythonlibs/#pymol (pymol-1.7.1.0.win-amd64-py2.7.exe)
Find application file C:Python27PyMOL
Find PyMOL application file in the list and create shortcut.
Drag shortcut to the desktop.
Double click icon on desktop to run PyMOL.
SETTING UP PYTHON (Win 7, 64-bit)
Note: Installing the extension may open
a C prompt window to compile.
Step 3
Download and install the wxPython extension:
http://downloads.sourceforge.net/
wxpython/wxPython3.0-win64-3.0.0.0-py27.exe](https://image.slidesharecdn.com/exploringyourpersonalgenomewithfreeonlinebioinformaticstoolsbackup3-140416173646-phpapp02/75/Exploring-your-personal-genome-with-free-online-bioinformatics-tools-35-2048.jpg)
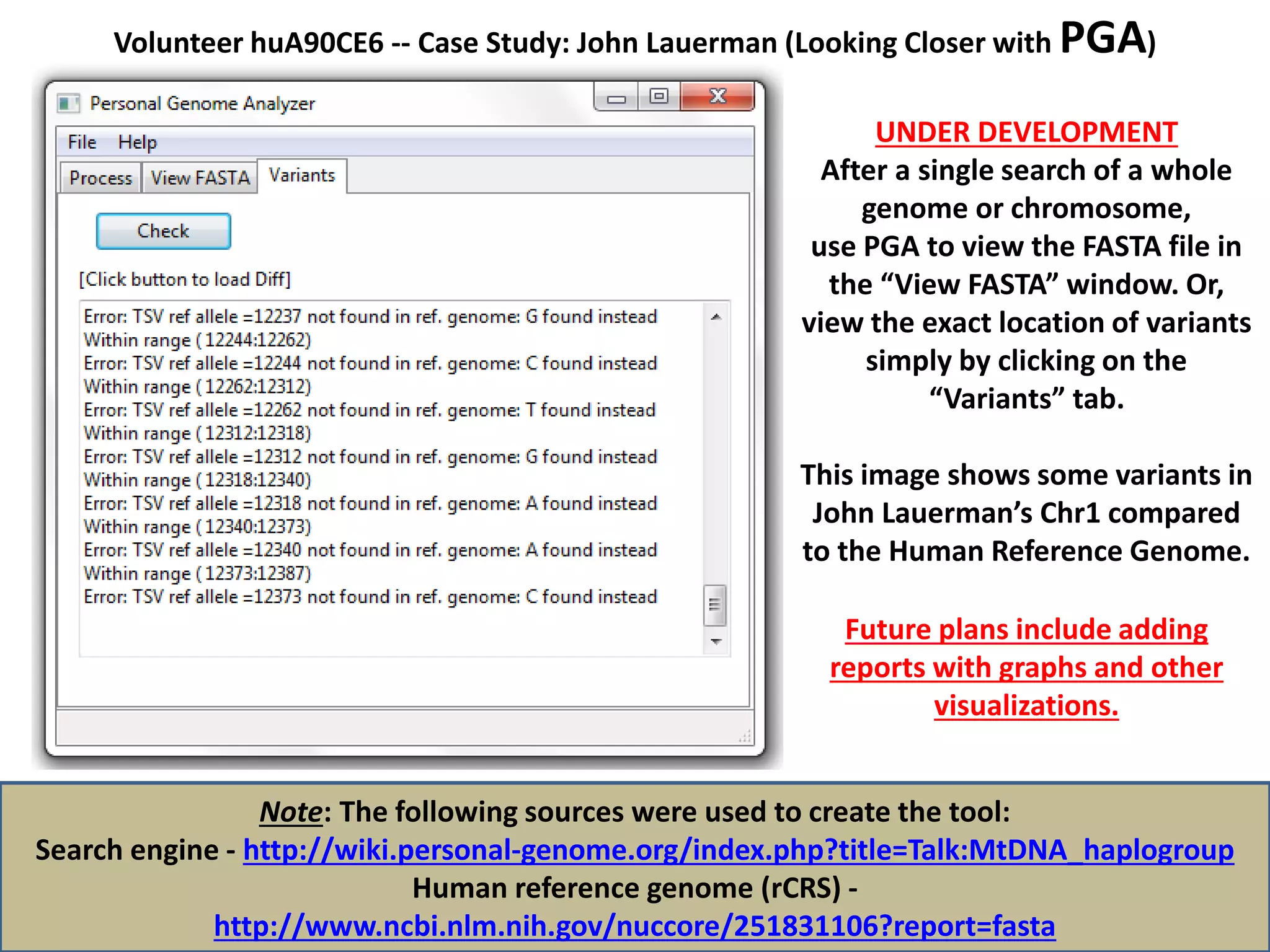
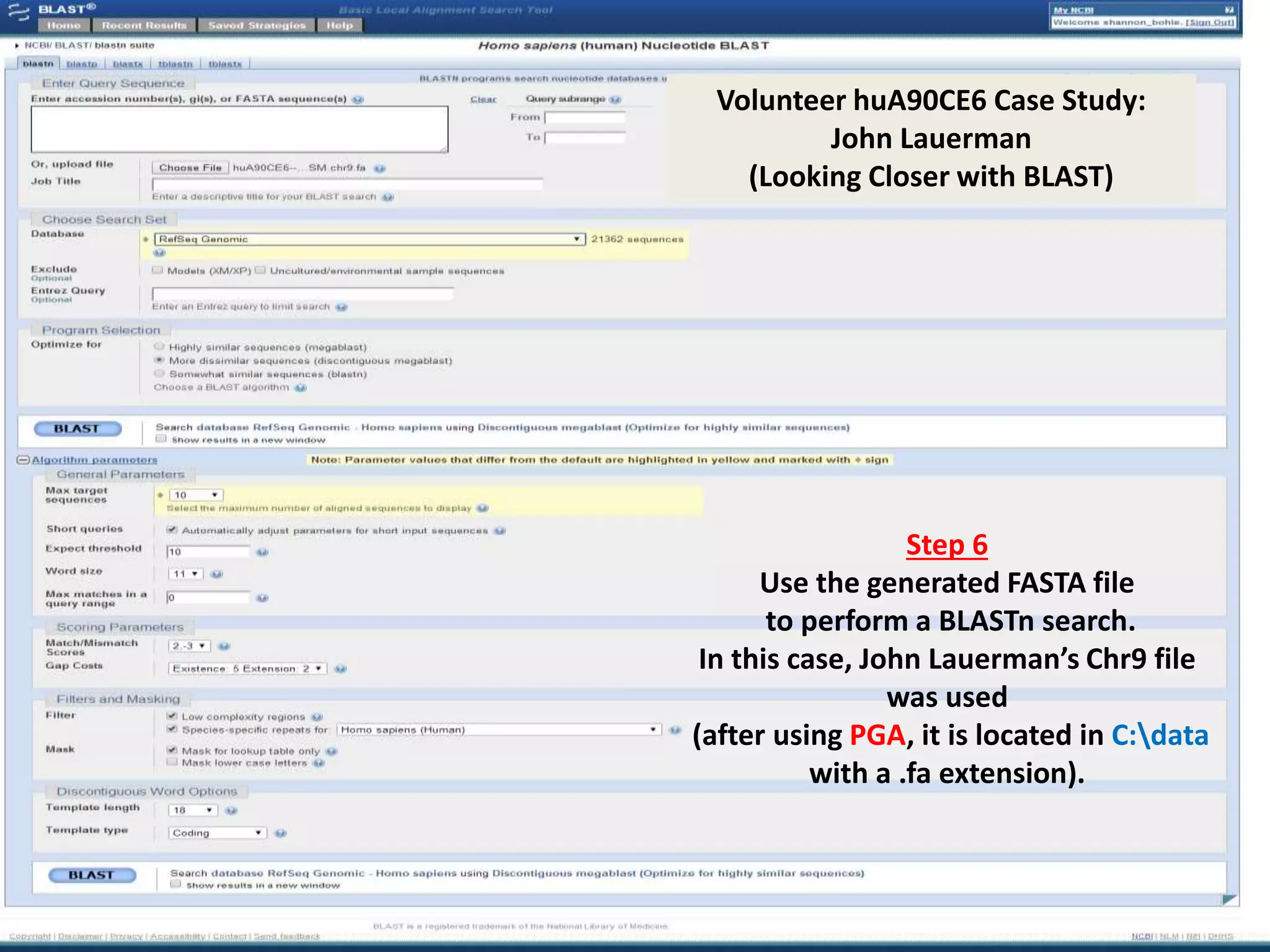
![Volunteer huA90CE6 -- Case Study: John Lauerman (Looking Closer with PGA)
Step 4
Use the Python-driven tool designed for this project to convert an isolated chromosome in
your whole genome sequence from TSV to FASTA and SQL formats in under 5 minutes.
Note: The following sources were used to create the tool:
Search engine - http://wiki.personal-genome.org/index.php?title=Talk:MtDNA_haplogroup
Human reference genome (rCRS) -
http://www.ncbi.nlm.nih.gov/nuccore/251831106?report=fasta
Insert
• Browse your hard drive for the Volunteer’s Whole Genome Sequence
• File name: huA90CE6--GS000006909-ASM.tsv
Insert
• Enter a single chromosome number you wish to examine
• 1-22, X, or Y; or leave blank for whole genome. [Enter 9]
Insert
• Enter an exact location or leave at defaults if you wish to scan the
whole chromosome or whole genome. [Use Default]
Check mark “Generate FA” for FASTA
Check mark “Generate SQL” for SQL
Click the PROCESS BUTTON.
Go to C:data
for the converted files in
FASTA and SQL formats.](https://image.slidesharecdn.com/exploringyourpersonalgenomewithfreeonlinebioinformaticstoolsbackup3-140416173646-phpapp02/75/Exploring-your-personal-genome-with-free-online-bioinformatics-tools-36-2048.jpg)


![Selected Bibliography
ALBERTS, B. (1983). Molecular biology of the cell. New York, Garland Pub.
CAREY, N. (2013). Epigenetics revolution: how modern biology is rewriting our
understanding of genetics, disease, and inheritance.
CHURCH, G. M., & REGIS, E. (2012). Regenesis: how synthetic biology will reinvent
nature and ourselves. New York, Basic Books.
SCHRÖDINGER, E. (2012). What is life?: the physical aspect of the living cell. Cambridge,
Univ. Press.
SKLOOT, R. (2010). The immortal life of Henrietta Lacks. New York, Crown Publishers.
VENTER, J. C. (2007). A life decoded: my genome, my life. New York, Viking.
WATSON, J. D. (1968). The double helix; a personal account of the discovery of the
structure of DNA. New York, Atheneum. [SIGNED FIRST EDITION]
WATSON, J. D. (2008). Molecular biology of the gene. San Francisco, Pearson/Benjamin
Cummings.
ZVELEBIL, M. J., & BAUM, J. O. (2008). Understanding bioinformatics. New York,
Garland Science.](https://image.slidesharecdn.com/exploringyourpersonalgenomewithfreeonlinebioinformaticstoolsbackup3-140416173646-phpapp02/75/Exploring-your-personal-genome-with-free-online-bioinformatics-tools-42-2048.jpg)