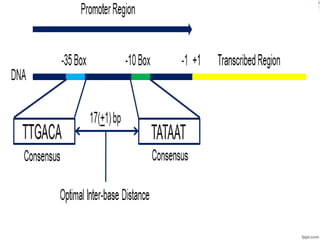
This document discusses promoters, which are DNA sequences located near genes that regulate gene expression. Promoters bind proteins like transcription factors to initiate transcription of genes into mRNA. The document covers different types of promoters in prokaryotes and eukaryotes, how promoters control gene expression through mechanisms like operons, and diseases associated with aberrant promoter function. It also discusses common laboratory promoters and why the study of promoters is important for controlling gene expression in research.