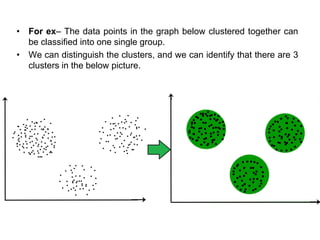
This document provides an overview of clustering and k-means clustering algorithms. It begins by defining clustering as the process of grouping similar objects together and dissimilar objects separately. K-means clustering is introduced as an algorithm that partitions data points into k clusters by minimizing total intra-cluster variance, iteratively updating cluster means. The k-means algorithm and an example are described in detail. Weaknesses and applications are discussed. Finally, vector quantization and principal component analysis are briefly introduced.










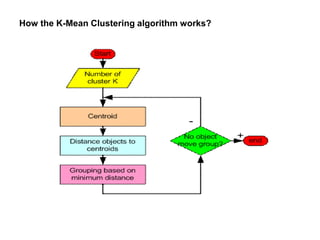
![The algorithm works as follows:
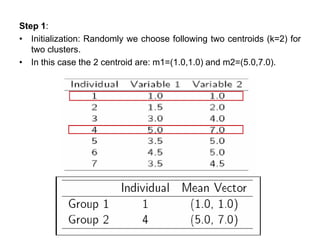
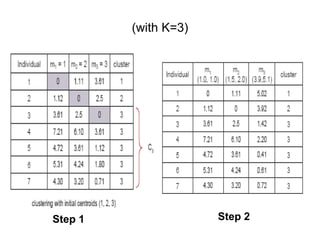
1.First we initialize k points, called means, randomly.
2.We categorize each item to its closest mean and we update the
mean’s coordinates, which are the averages of the items categorized in
that mean so far.
3.We repeat the process for a given number of iterations and at the
end, we have our clusters.
The “points” mentioned above are called means, because they hold the
mean values of the items categorized in it.
-To initialize these means, we have a lot of options. An intuitive
method is to initialize the means at random items in the data set.
- Another method is to initialize the means at random values
between the boundaries of the data set (if for a feature x the items have
values in [0,3], we will initialize the means with values for x at [0,3]).](https://image.slidesharecdn.com/csa3702machinelearningmodule-3-201106074109/85/CSA-3702-machine-learning-module-3-12-320.jpg)



















![Implementing PCA on a 2-D Dataset
Step 1: Normalize the data (get sample code)
• First step is to normalize the data that we have so that PCA works
properly. This is done by subtracting the respective means from the
numbers in the respective column. So if we have two dimensions
X and Y, all X become 𝔁- and all Y become 𝒚-. This produces a
dataset whose mean is zero.
Step 2: Calculate the covariance matrix (get sample code)
• Since the dataset we took is 2-dimensional, this will result in a 2x2
Covariance matrix.
• Please note that Var[X1] = Cov[X1,X1] and Var[X2] = Cov[X2,X2].](https://image.slidesharecdn.com/csa3702machinelearningmodule-3-201106074109/85/CSA-3702-machine-learning-module-3-41-320.jpg)

















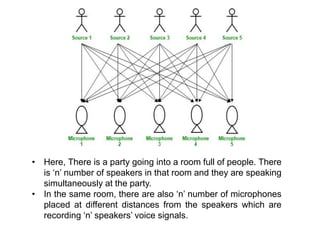
![• Hence, the number of speakers is equal to the number must of
microphones in the room.
• Now, using these microphones’ recordings, we want to separate all
the ‘n’ speakers’ voice signals in the room given each microphone
recorded the voice signals coming from each speaker of different
intensity due to the difference in distances between them.
• Decomposing the mixed signal of each microphone’s recording into
independent source’s speech signal can be done by using the
machine learning technique, independent component analysis.
[ X1, X2, ….., Xn ] => [ Y1, Y2, ….., Yn ]
- where, X1, X2, …, Xn are the original signals present in the
mixed signal and Y1, Y2, …, Yn are the new features and are
independent components which are independent of each other.](https://image.slidesharecdn.com/csa3702machinelearningmodule-3-201106074109/85/CSA-3702-machine-learning-module-3-61-320.jpg)






![Input X: D by N matrix consisting of N data items in D dimensions.
Output Y: d by N matrix consisting of d < D dimensional embedding
coordinates for the input points.
1. Find neighbours in X space [b,c].
for i=1:N
compute the distance from Xi to every other point Xj
find the K smallest distances
assign the corresponding points to be neighbours of Xi
end
2. Solve for reconstruction weights W
for i=1:N
create matrix Z consisting of all neighbours of Xi [d]
subtract Xi from every column of Z
compute the local covariance C=Z'*Z [e]
solve linear system C*w = 1 for w [f]
set Wij=0 if j is not a neighbor of i
set the remaining elements in the ith row of W equal to
w/sum(w);](https://image.slidesharecdn.com/csa3702machinelearningmodule-3-201106074109/85/CSA-3702-machine-learning-module-3-71-320.jpg)
![3. Compute embedding coordinates Y using weights W.
create sparse matrix M = (I-W)'*(I-W)
find bottom d+1 eigenvectors of M
(corresponding to the d+1 smallest eigenvalues)
set the qth ROW of Y to be the q+1 smallest eigenvector
(discard the bottom eigenvector [1,1,1,1...] with
eigenvalue zero)
Advantages of LLE
Better computational time
Consideration of the non-linearity of the structure
Applications
Data visualization
Information retrieval
Image process
Pattern recognition](https://image.slidesharecdn.com/csa3702machinelearningmodule-3-201106074109/85/CSA-3702-machine-learning-module-3-72-320.jpg)





![Possible issues
• The connectivity of each data point in the neighborhood graph is
defined as its nearest k Euclidean neighbors in the high-dimensional
space.
• This step is vulnerable to "short-circuit errors" if k is too large with
respect to the manifold structure or if noise in the data moves the
points slightly off the manifold.
• Even a single short-circuit error can alter many entries in the
geodesic distance matrix, which in turn can lead to a drastically
different (and incorrect) low-dimensional embedding.
• Conversely, if k is too small, the neighborhood graph may become
too sparse to approximate geodesic paths accurately. But
improvements have been made to this algorithm to make it work
better for sparse and noisy data sets.[5]](https://image.slidesharecdn.com/csa3702machinelearningmodule-3-201106074109/85/CSA-3702-machine-learning-module-3-78-320.jpg)