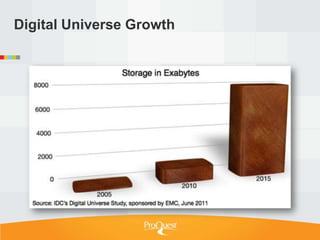
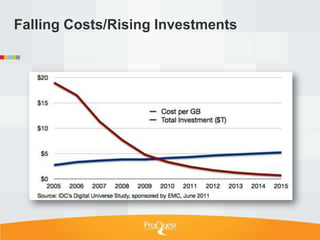
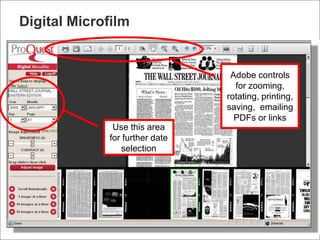
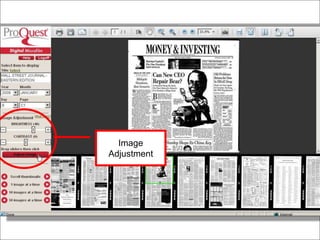
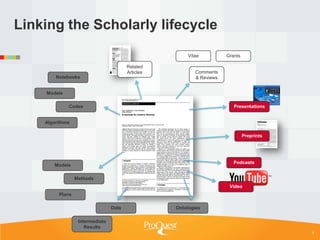
Tim Babbitt discusses the changing context of research and scholarship due to digitization and the internet. The inputs and outputs of research are increasingly digital and complex, including data, code, presentations, and more. ProQuest has a history of preserving scholarship through microfilming and is exploring how to preserve the full range of digital scholarly outputs and their linkages in a sustainable way. Key questions include balancing new and old preservation methods and moving beyond preserving individual objects to also preserving networks and linkages between scholarly works.







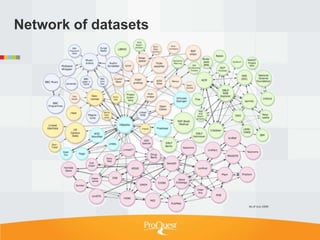
![Examples of text as data
Changes in word sense ( e.g. consumption( TB )
, moot, oratio1 ) and spelling (e.g. 18th C. ſ to s ,
*re *er )
Bibliometrics and other usage analyses
Citation patterns
Institution vs. discipline
Author demographics
Pharma: Drug / Symptom correlation.
Biology: Species / date / location observations.
Social Sci: Work/life habits of undergrads based
on access patterns at different institutions [ usage
data based]
…
10](https://image.slidesharecdn.com/charlestonconf2012preservationfinalb-130113212208-phpapp02/85/Preserving-the-Inputs-and-Outputs-of-Scholarship-10-320.jpg)