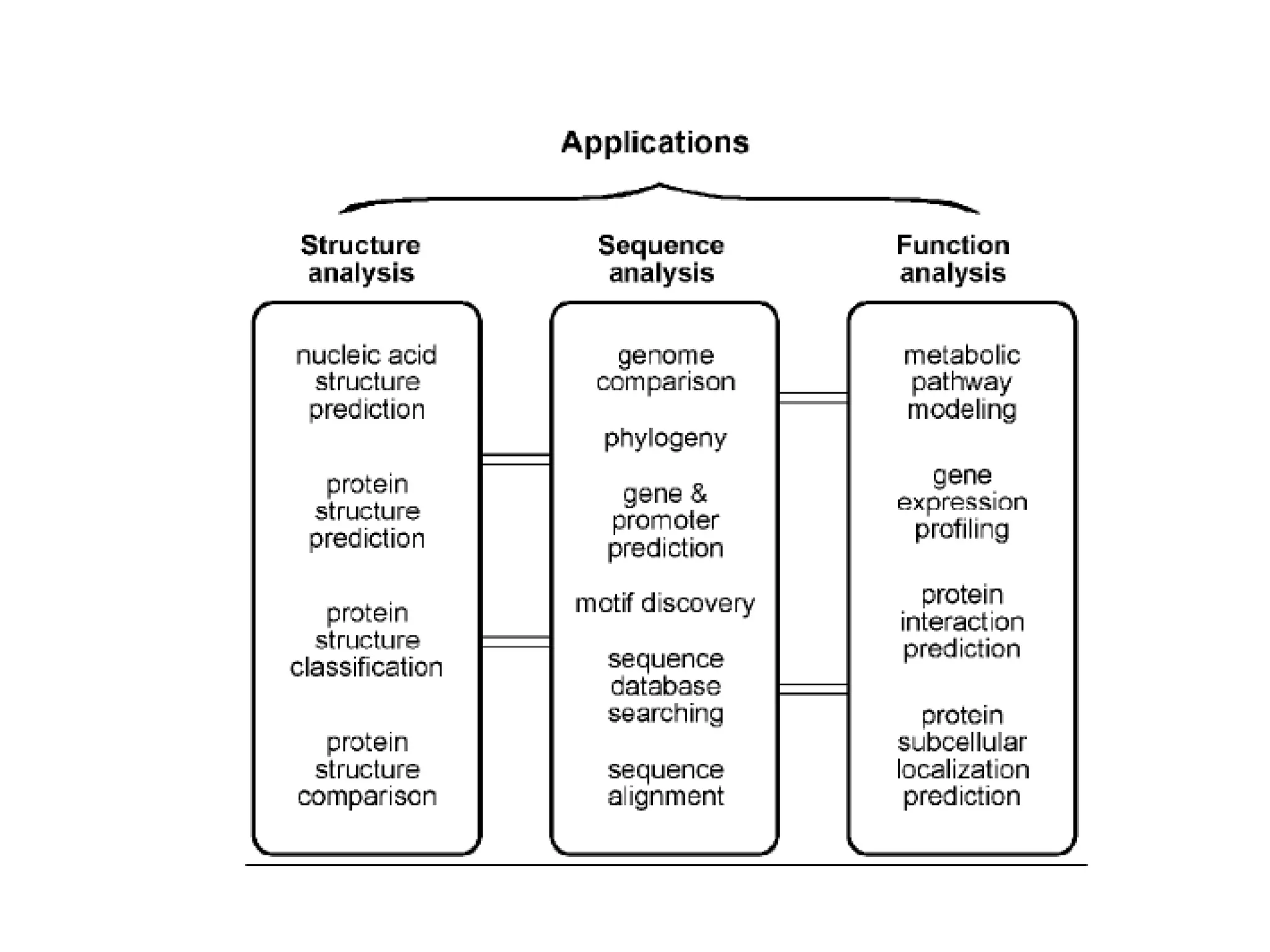
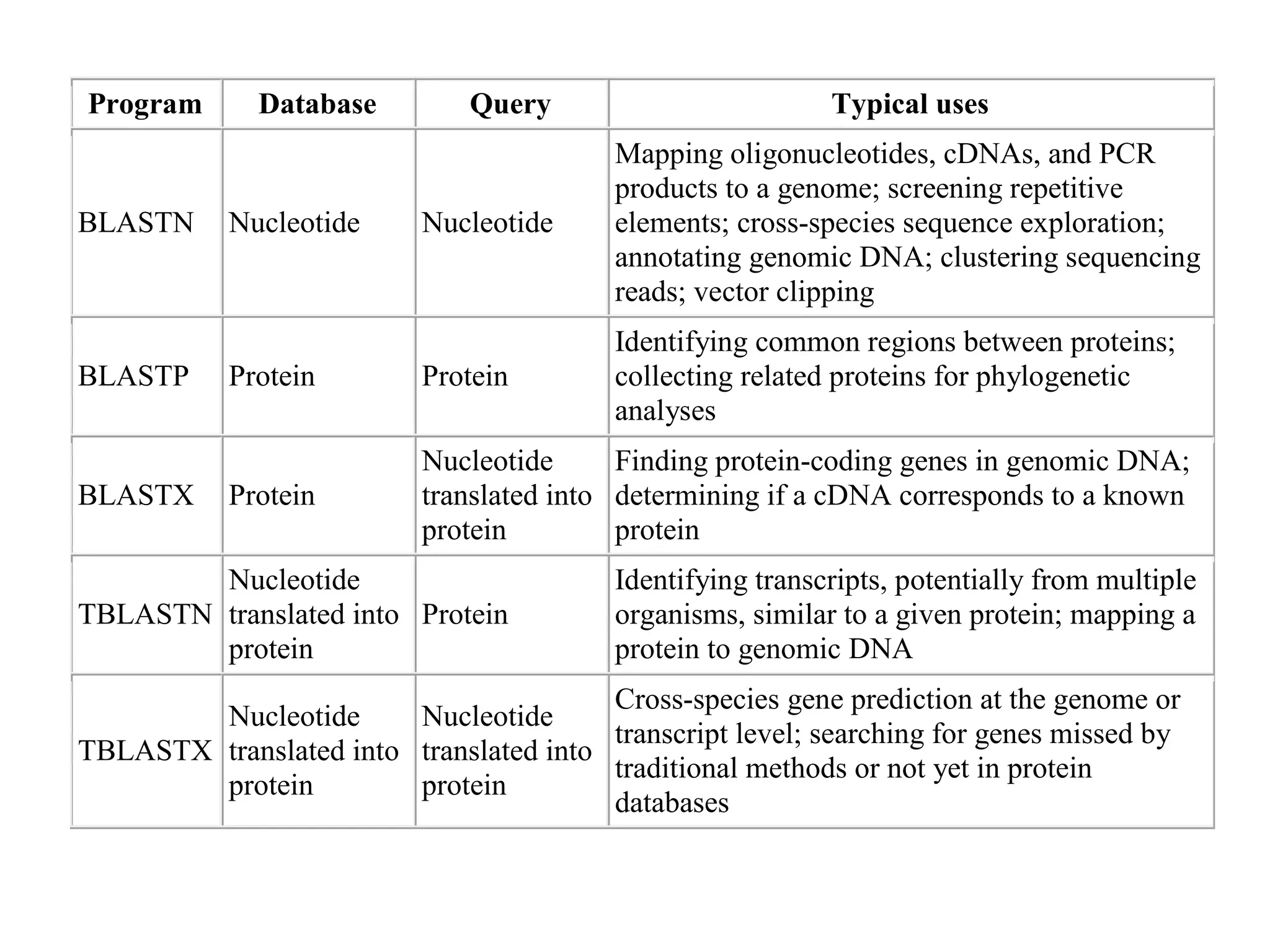
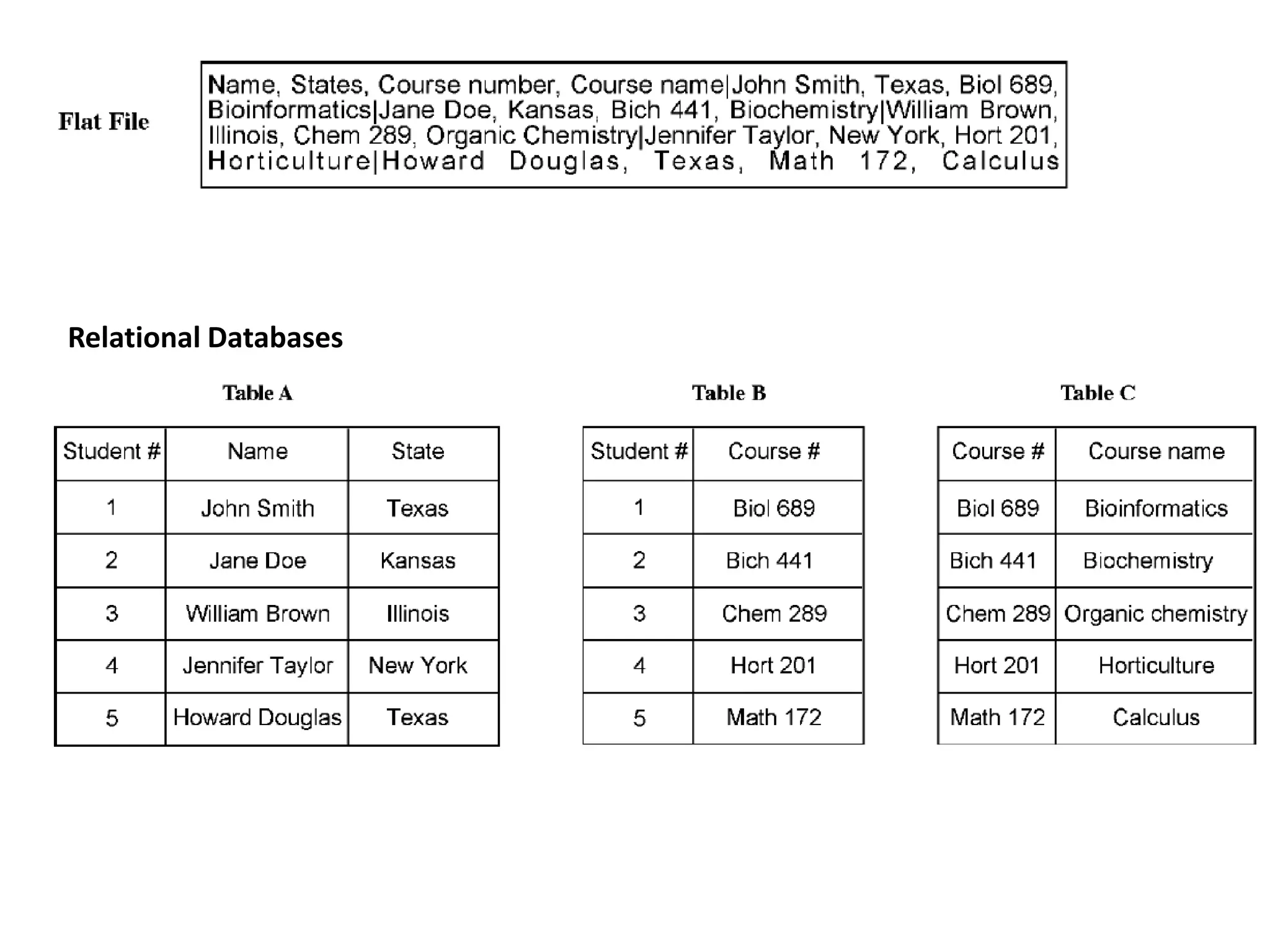
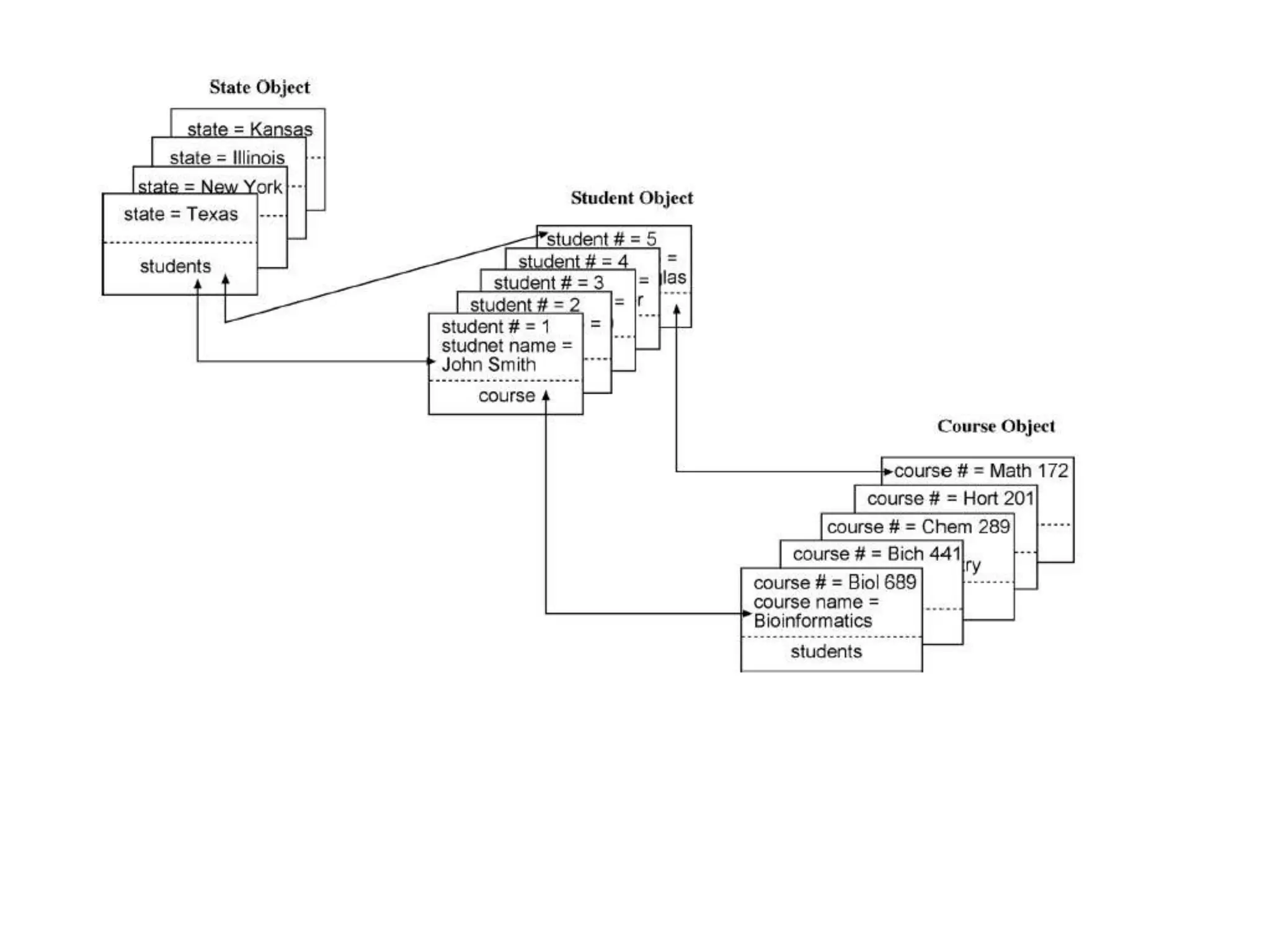
This document provides an overview of the field of bioinformatics. It defines bioinformatics as the intersection of biology and computer science, using computational tools to analyze and distribute biological information like DNA, RNA, and proteins. The goals of bioinformatics are to better understand cells at the molecular level by analyzing sequence and structure data. Key applications include drug design, DNA analysis, and agricultural biotechnology. The document also describes different types of biological databases like primary databases that contain raw sequence data, and secondary databases that provide additional annotation and analysis of sequences.