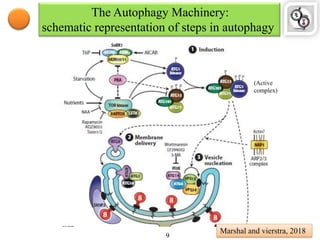
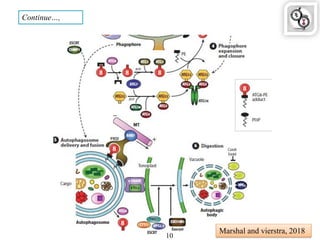
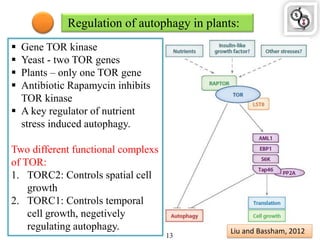
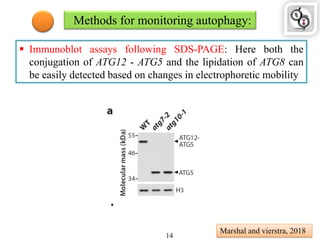
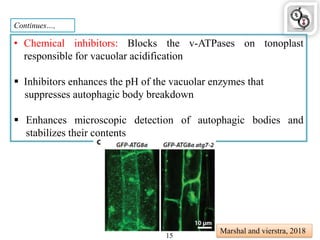
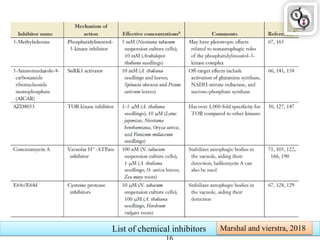
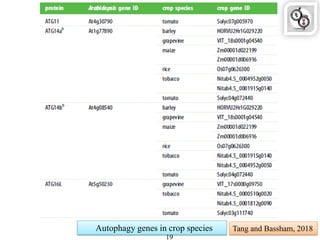
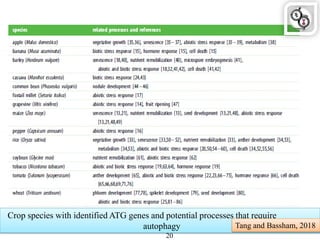
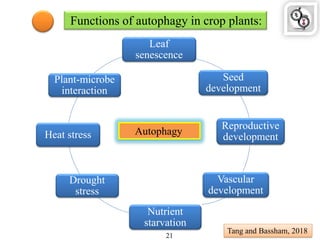
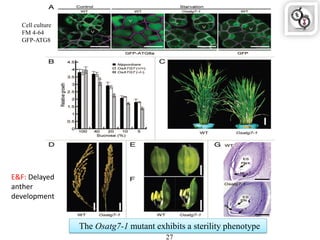
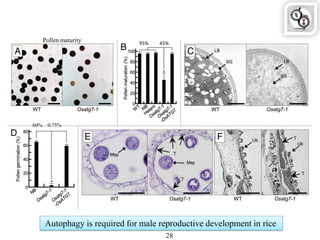
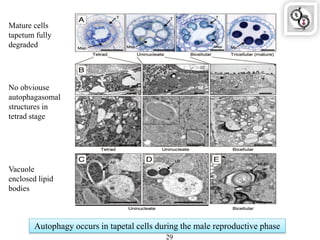
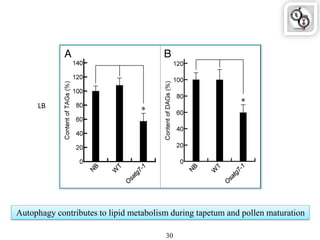
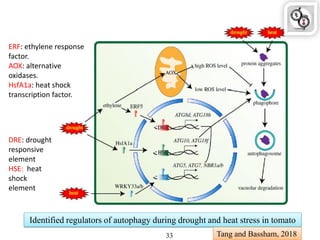
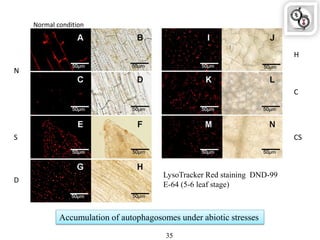
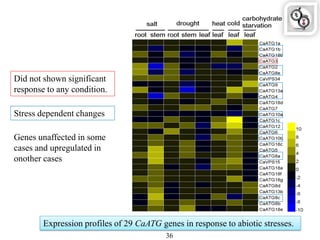
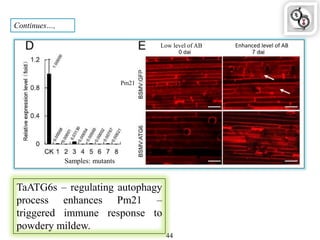
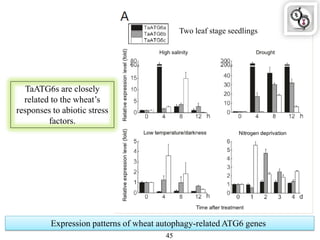
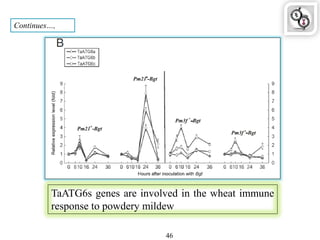
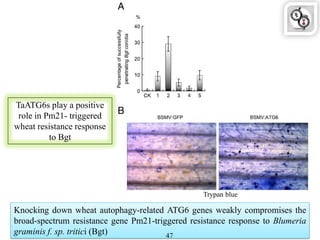
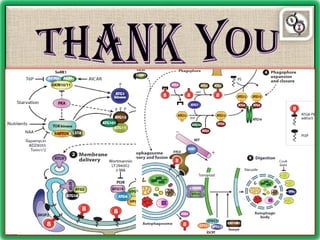
Autophagy is a process in which cells degrade and recycle cellular components. The document discusses autophagy in crop plants, including the history and discovery of autophagy genes, the autophagy machinery and mechanisms, and functions of autophagy in processes like leaf senescence, seed development, and stress responses. It also explores the role of autophagy in plant-pathogen interactions and future research prospects. Manipulating autophagy through increased expression of autophagy genes has potential for agricultural benefits like higher yield and stress tolerance in crop plants.