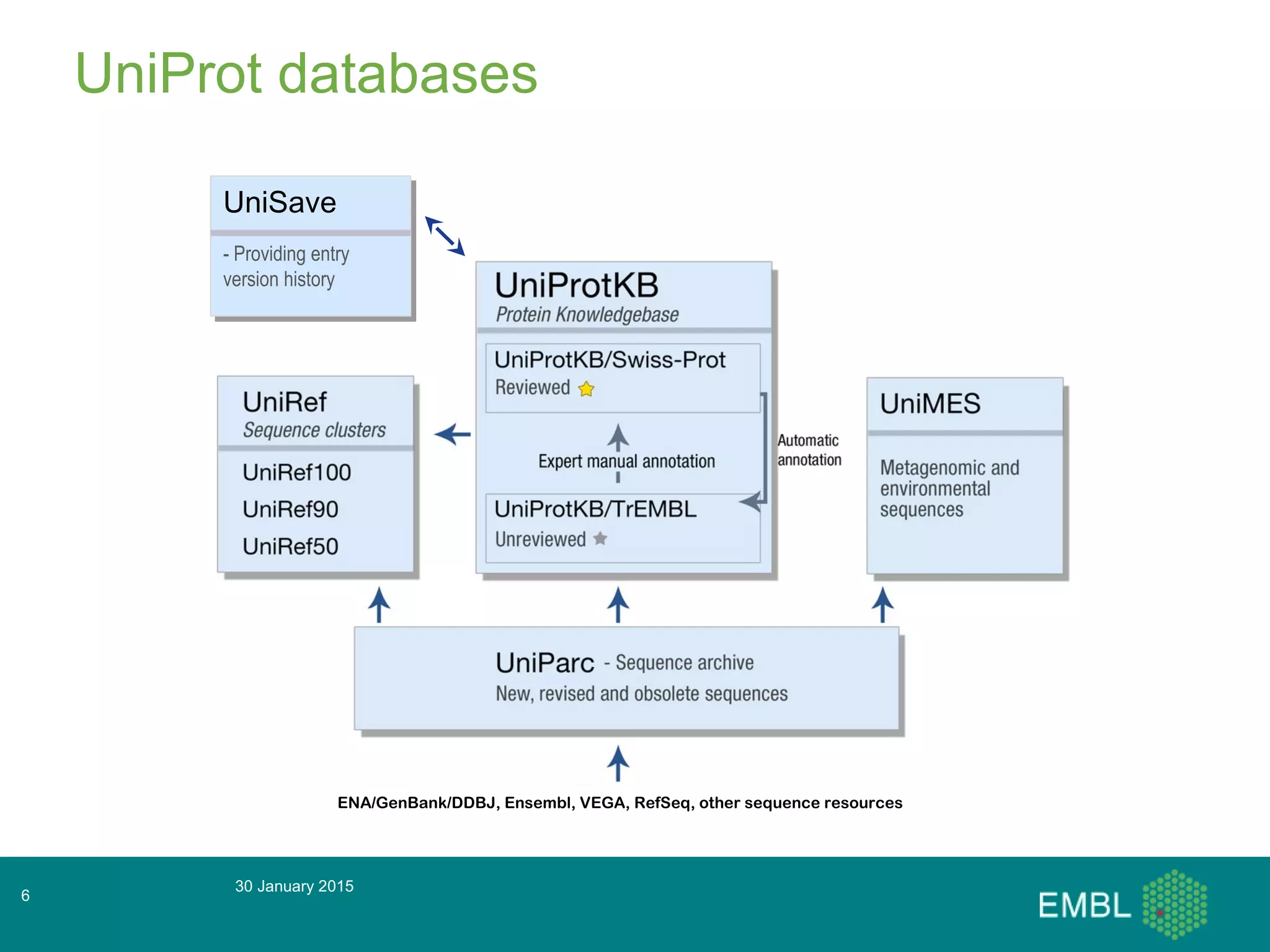
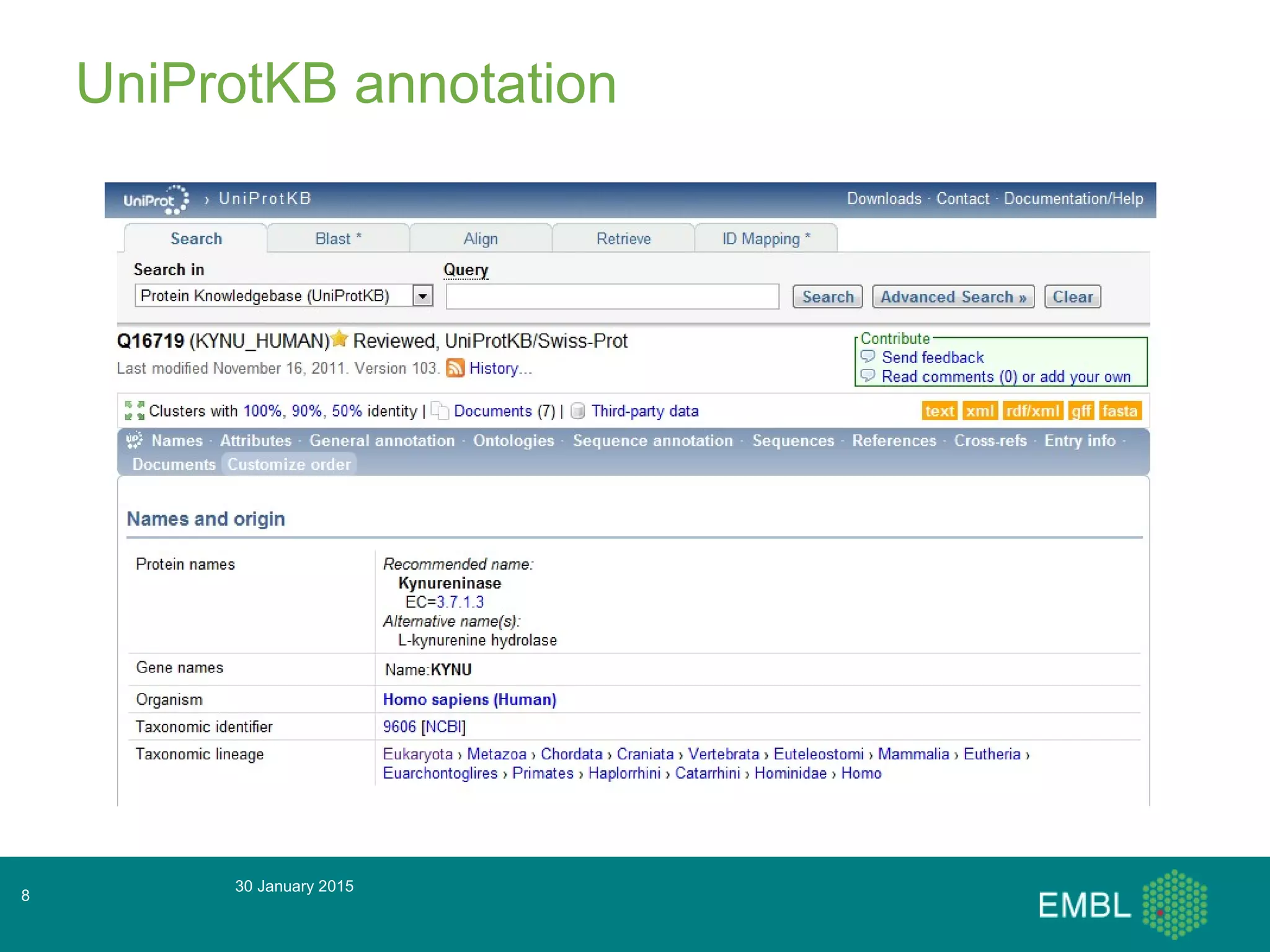
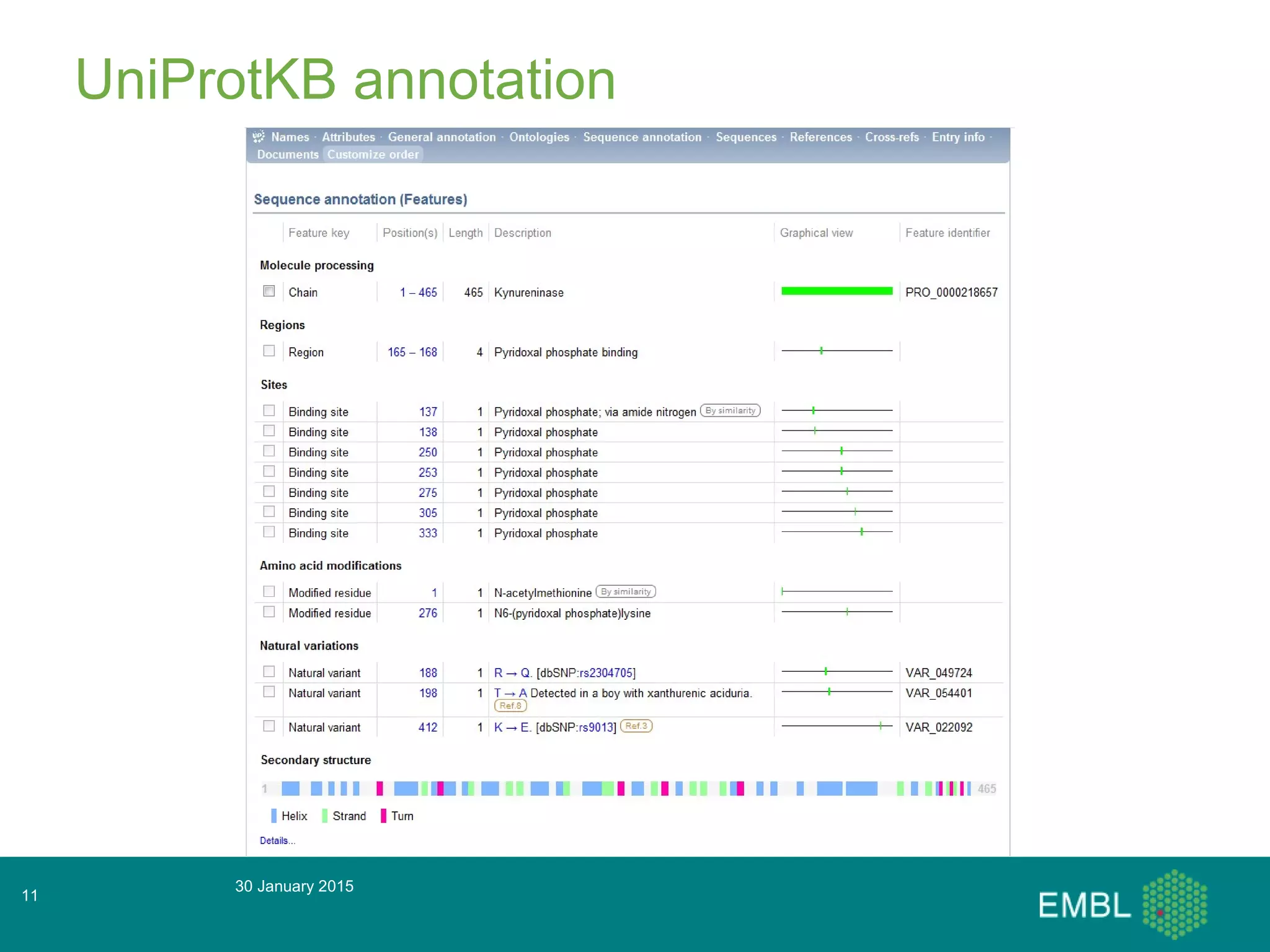
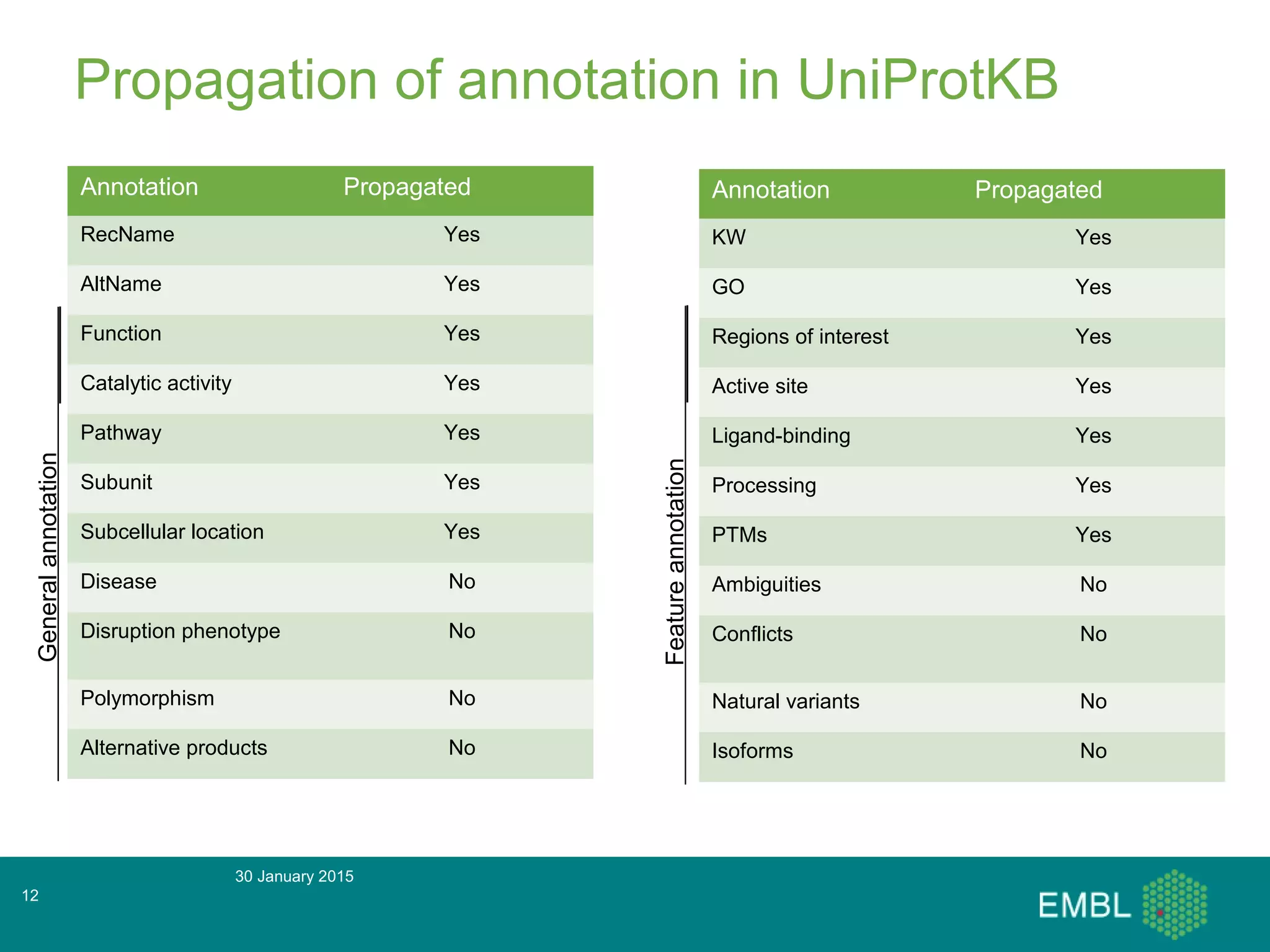
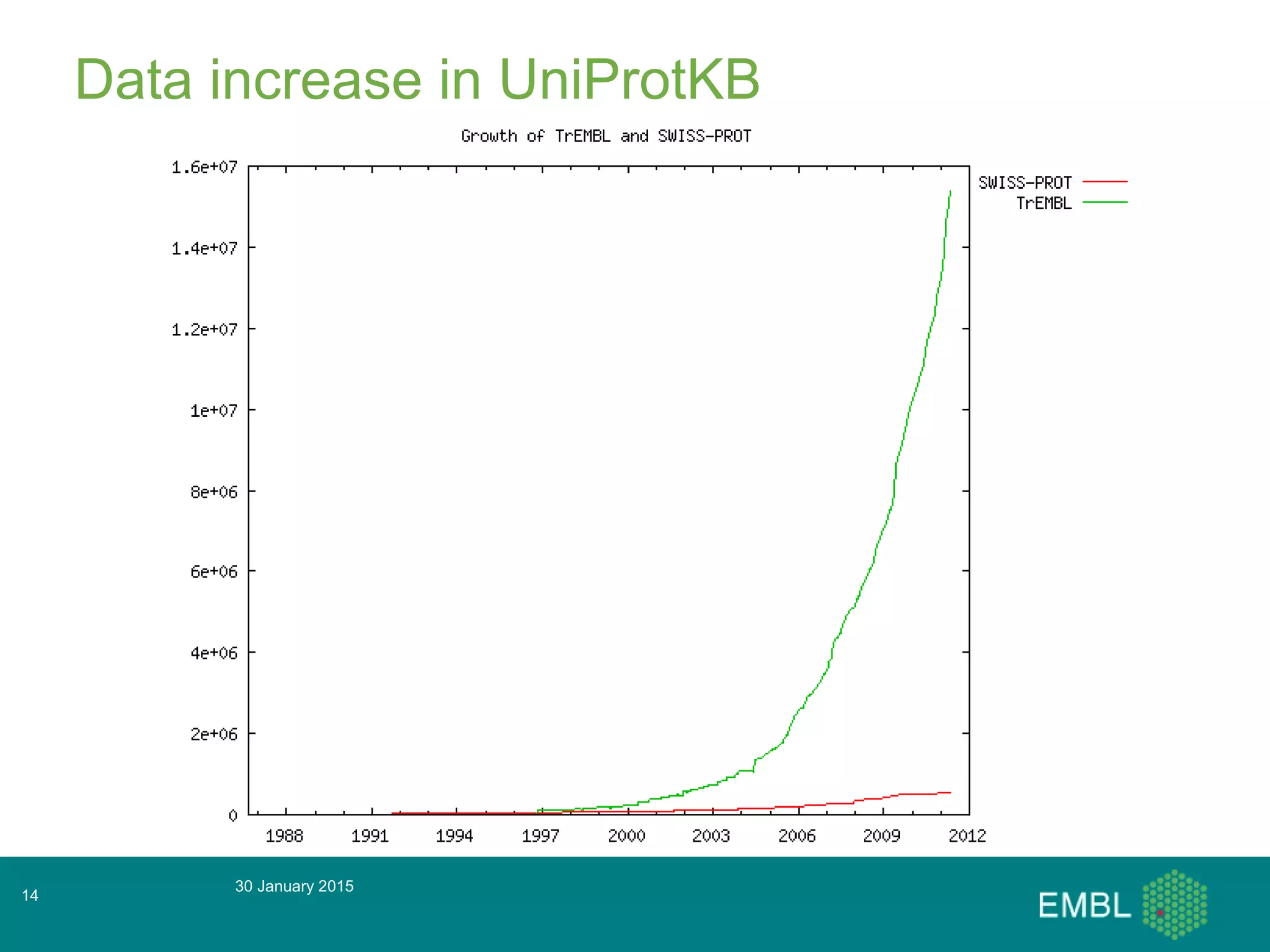
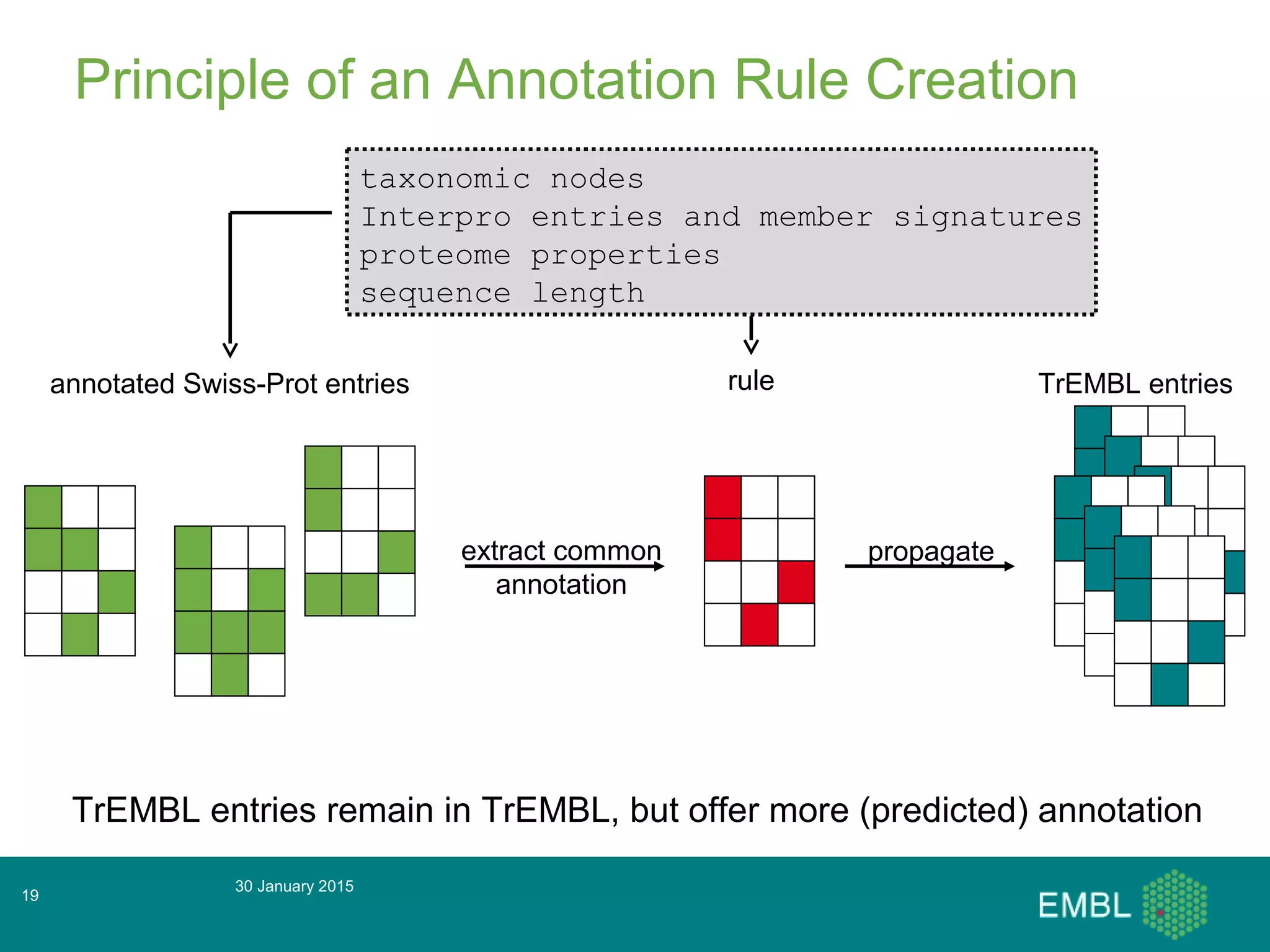
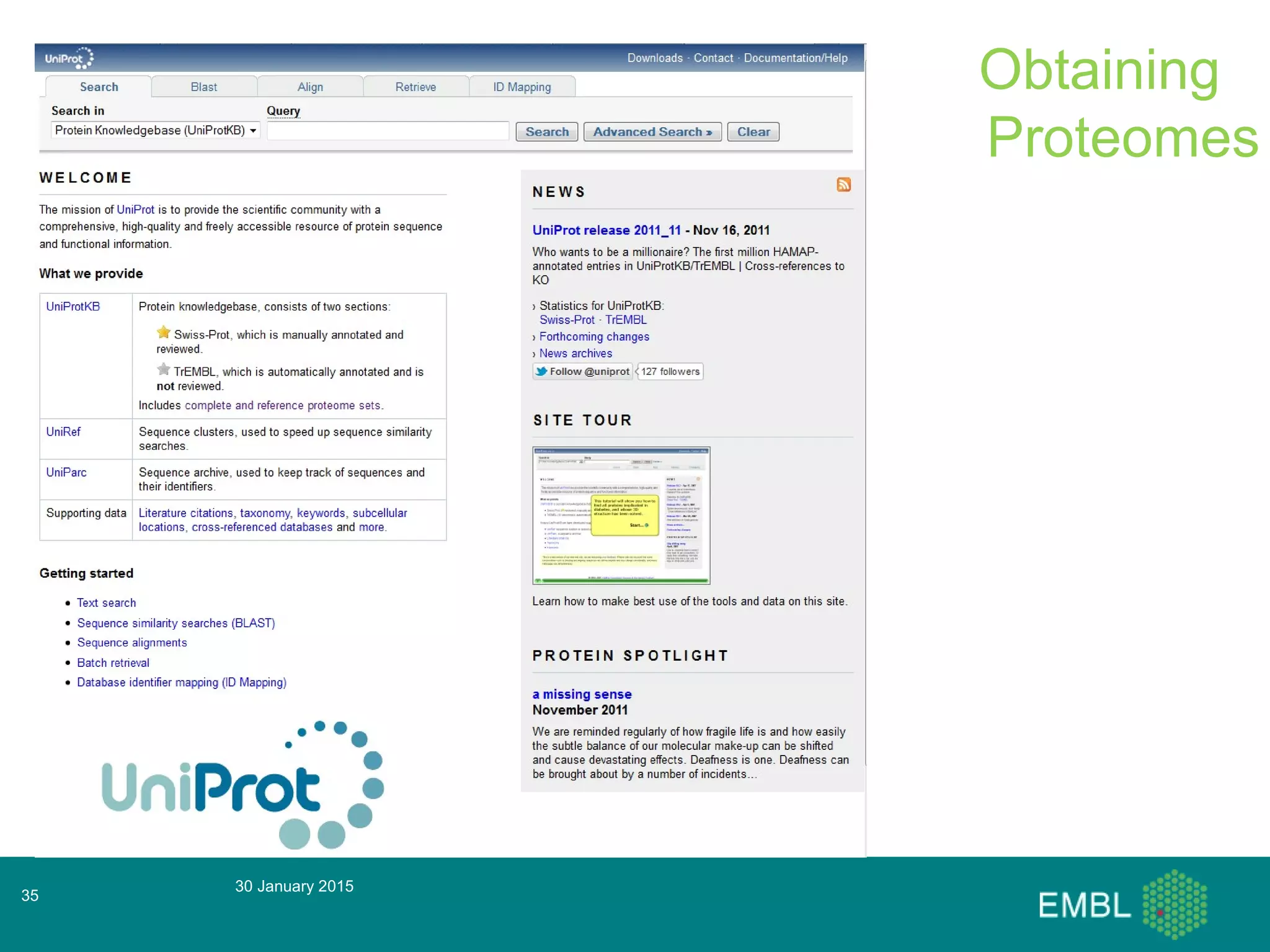
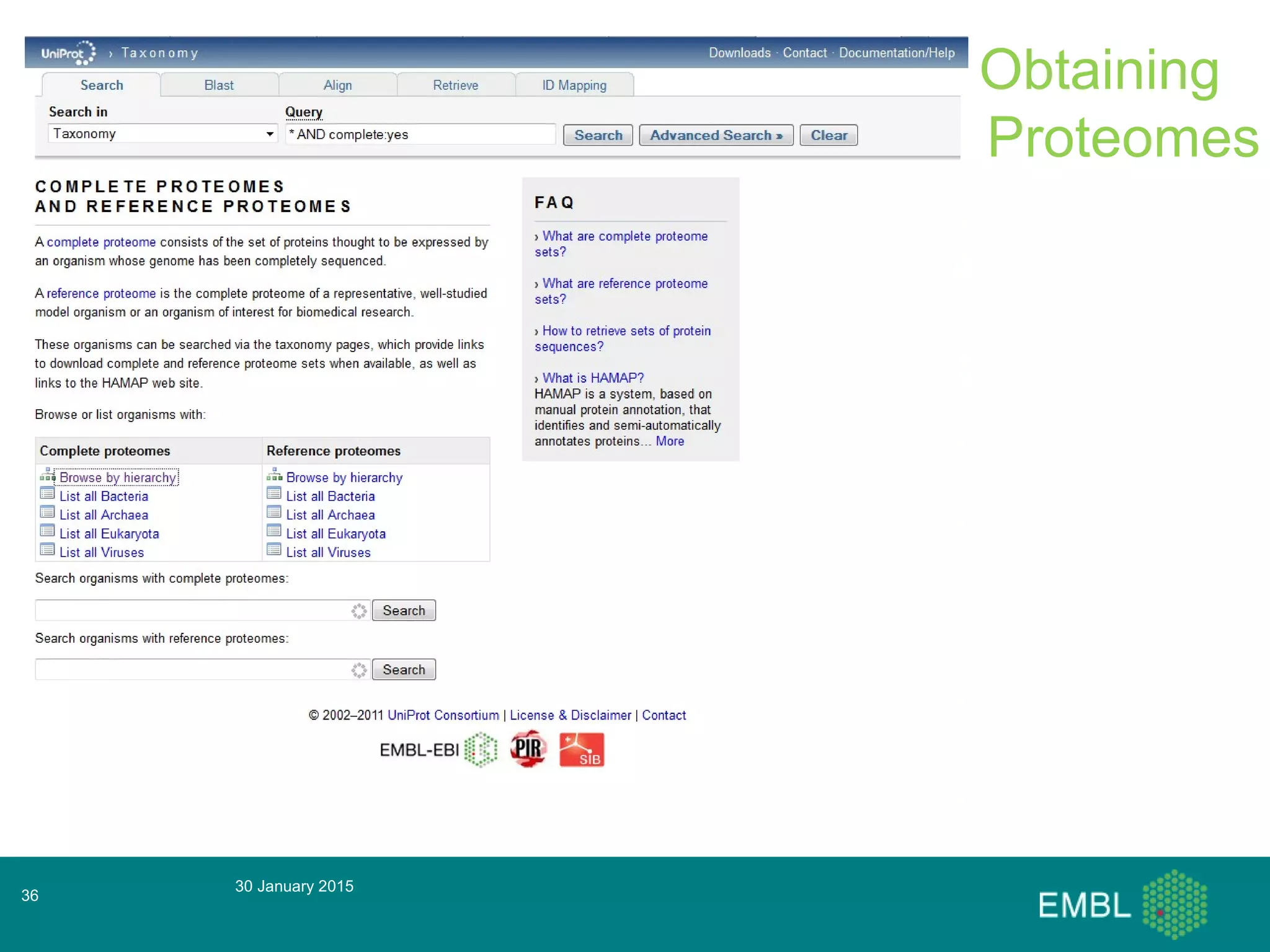
This document summarizes a presentation about automatic annotation in UniProtKB. It discusses how UniProtKB uses automatic annotation systems like UniRule and SAAS to annotate the increasing number of sequences as manual annotation cannot keep pace. UniRule allows manual creation of annotation rules while SAAS uses machine learning. It also describes how UniProtKB defines and works to annotate complete proteomes from various organisms.