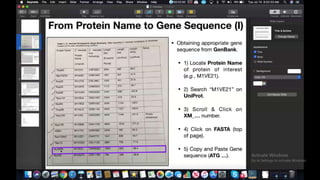
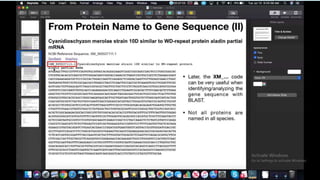
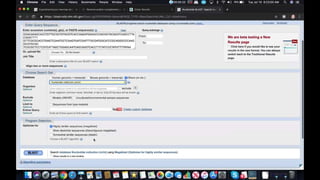
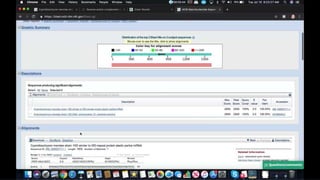
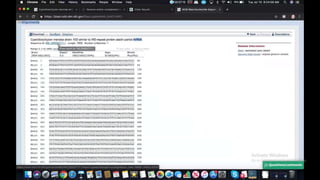
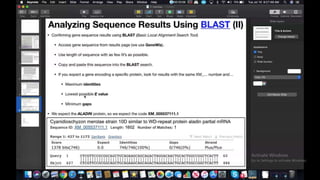
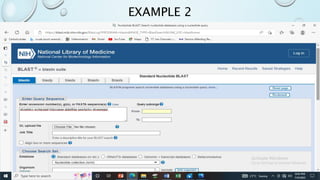
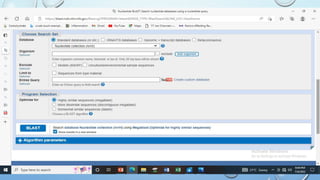
This document provides information about two BLAST searches that were conducted to identify unknown DNA and protein sequences. The results of the first BLAST search found a DNA sequence with a high percent identity and low E value, indicating it was a significant match. The second BLAST search found a protein sequence with a high percent identity and query cover, also representing a strong match. The document then explains how to interpret the key fields in a BLAST search result, such as E value, percent identity, and query cover.






















![REFERENCES
AFIQAH-ALENG N, MOHAMED-HUSSEIN ZA. CONSTRUCTION OF PROTEIN
EXPRESSION NETWORK. METHODS MOL BIOL. 2021;2189:119-132. DOI:
10.1007/978-1-0716-0822-7_10. PMID: 33180298.
STRUYF P, DE MOOR S, VANDEVIVER C, RENARD B, VANDER BEKEN T. THE
EFFECTIVENESS OF DNA DATABASES IN RELATION TO THEIR PURPOSE AND CONTENT:
A SYSTEMATIC REVIEW. FORENSIC SCI INT. 2019 AUG;301:371-381. DOI:
10.1016/J.FORSCIINT.2019.05.052. EPUB 2019 JUN 5. PMID: 31212144.
KANZ,C. ET AL. (2005) THE EMBL NUCLEOTIDE SEQUENCE DATABASE. NUCLEIC
ACIDS RES., 33, D29–D33.
ALTSCHUL SF, GISH W, MILLER W, MYERS EW, LIPMAN DJ: BASIC LOCAL ALIGNMENT
SEARCH TOOL. J MOL BIOL 1990, 215:403-410.2.
NCBI BLAST [HTTP://WWW.NCBI.NLM.NIH.GOV/BLAST/]](https://image.slidesharecdn.com/group3presentation-220826080808-d743d970/85/Group-3-presentation-pptx-31-320.jpg)