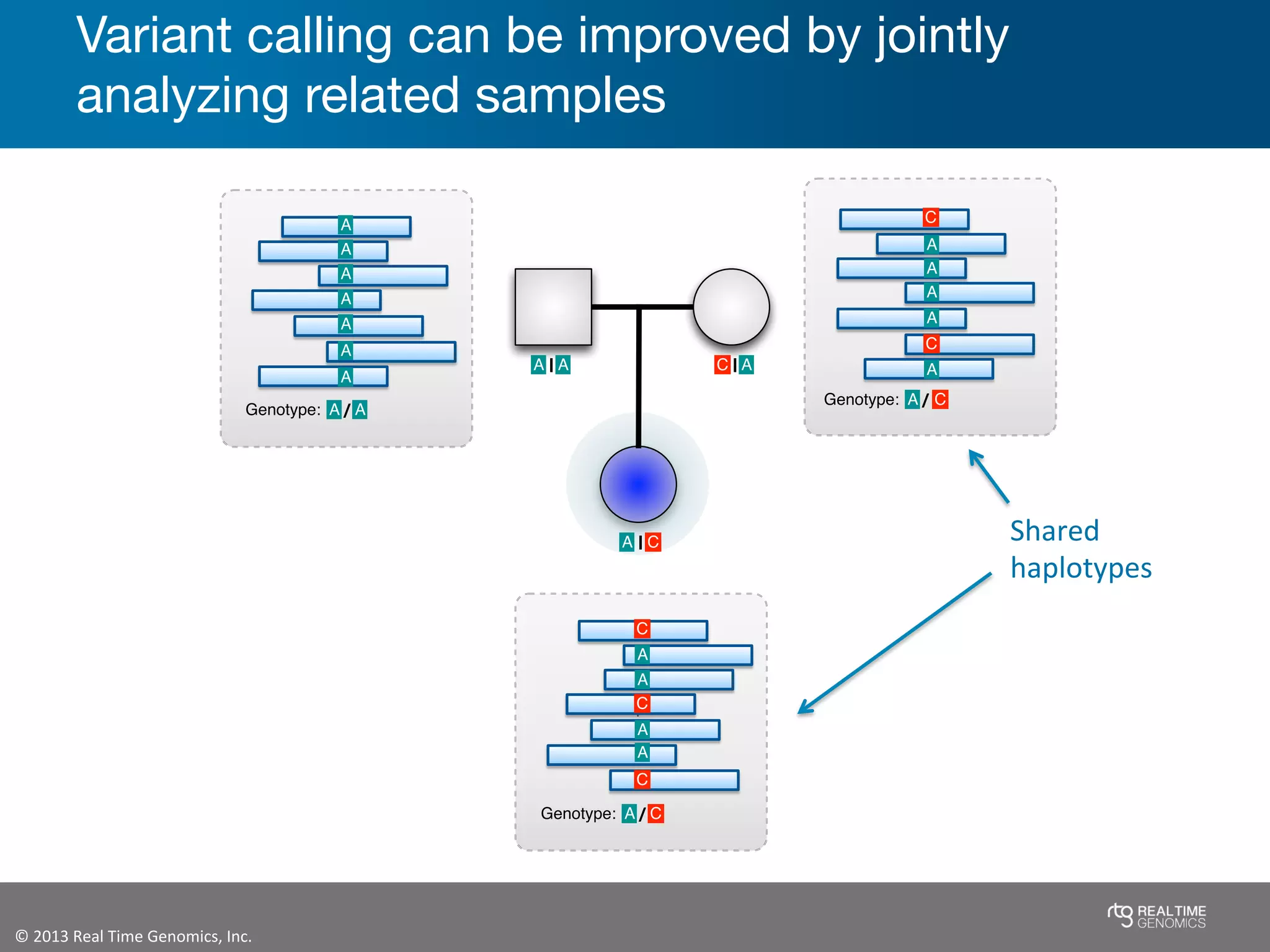
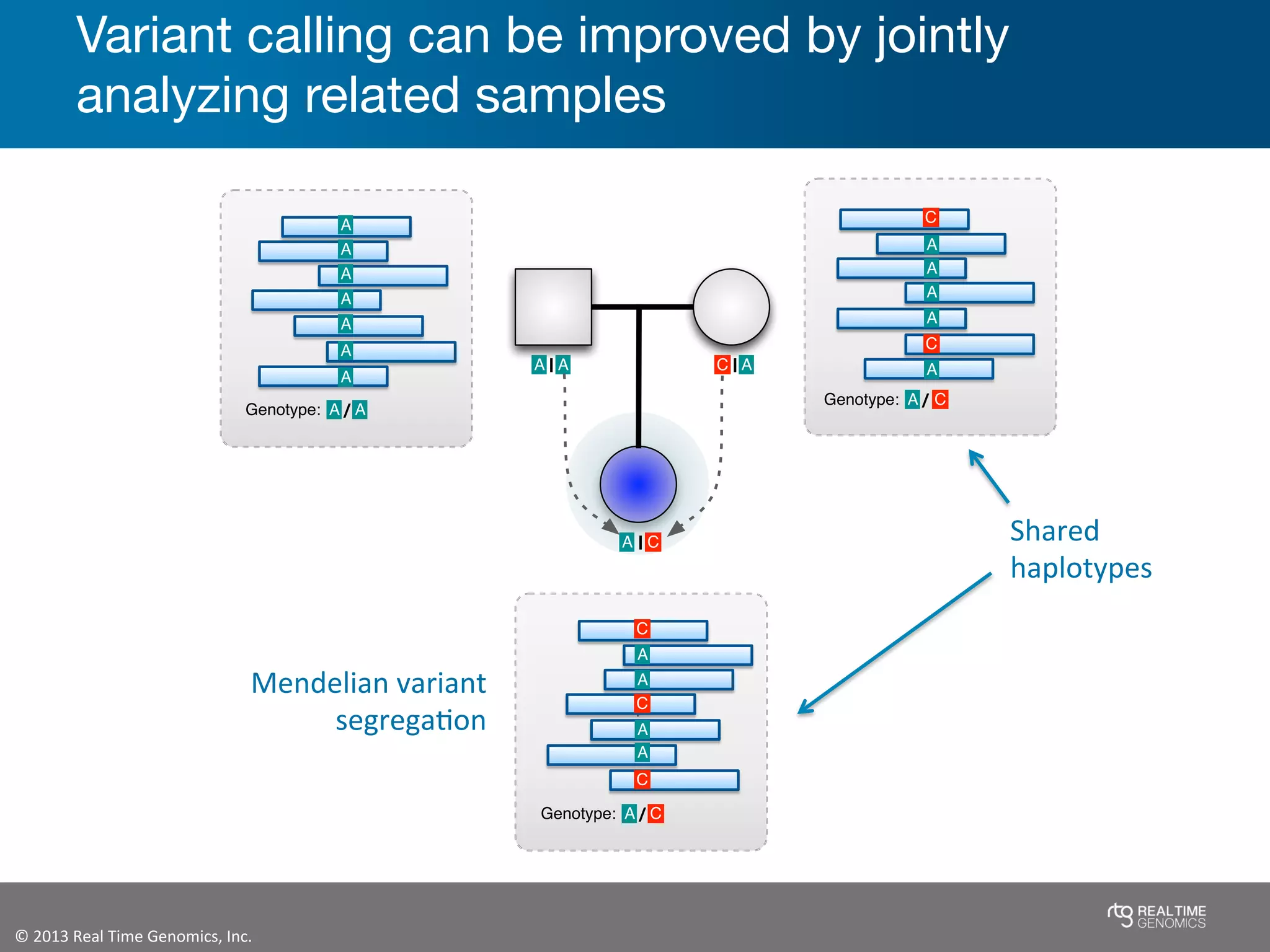
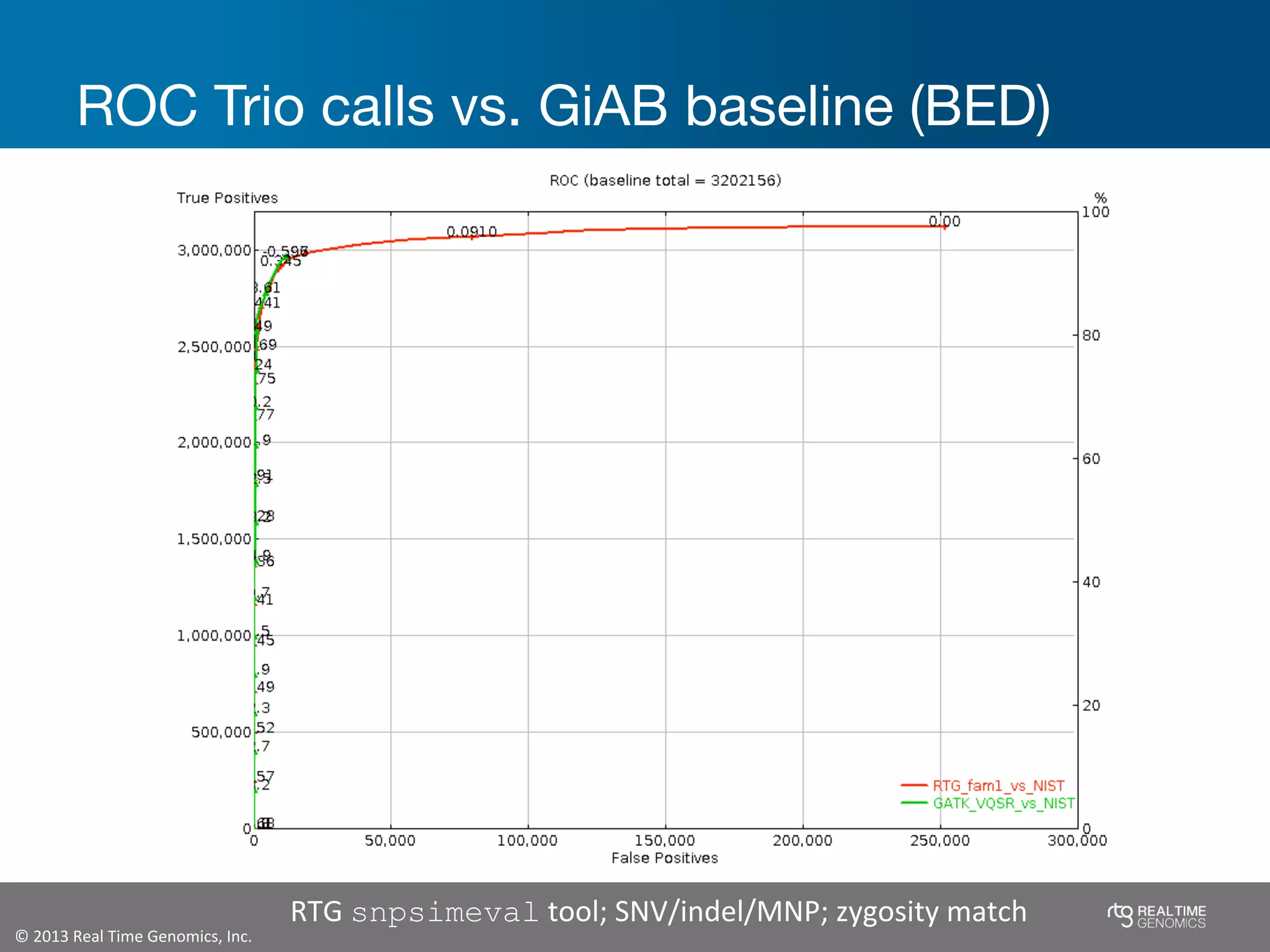
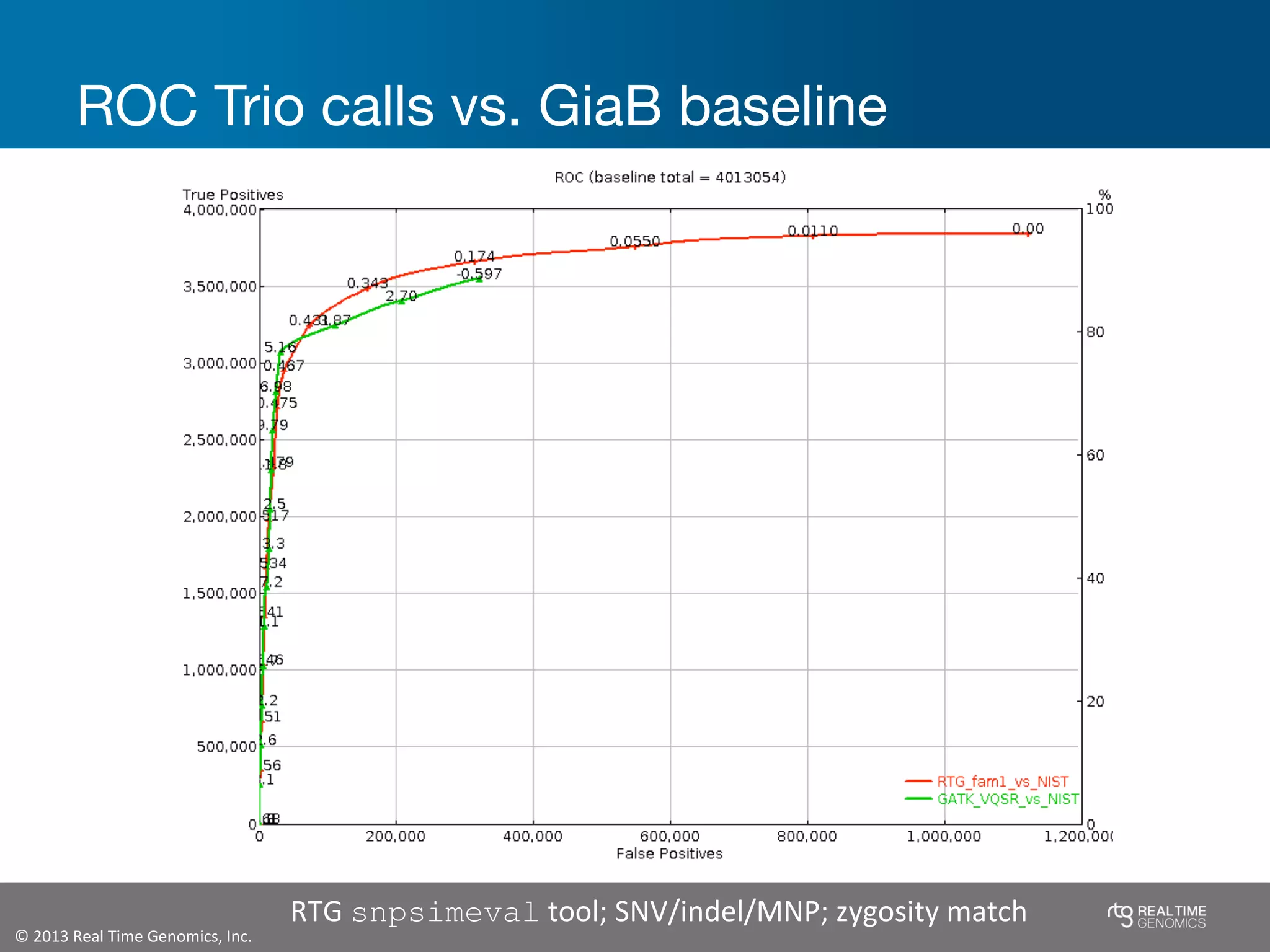
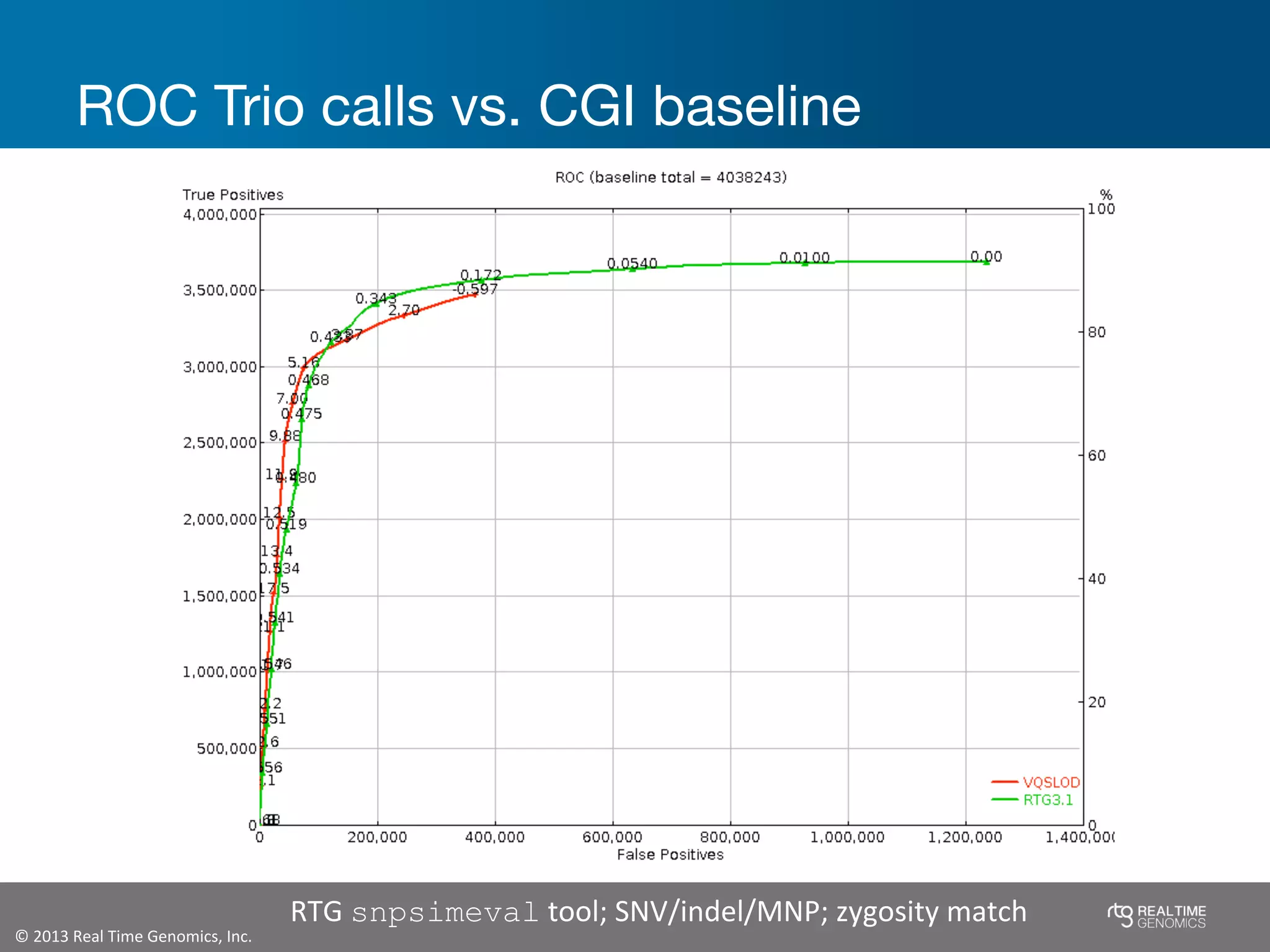
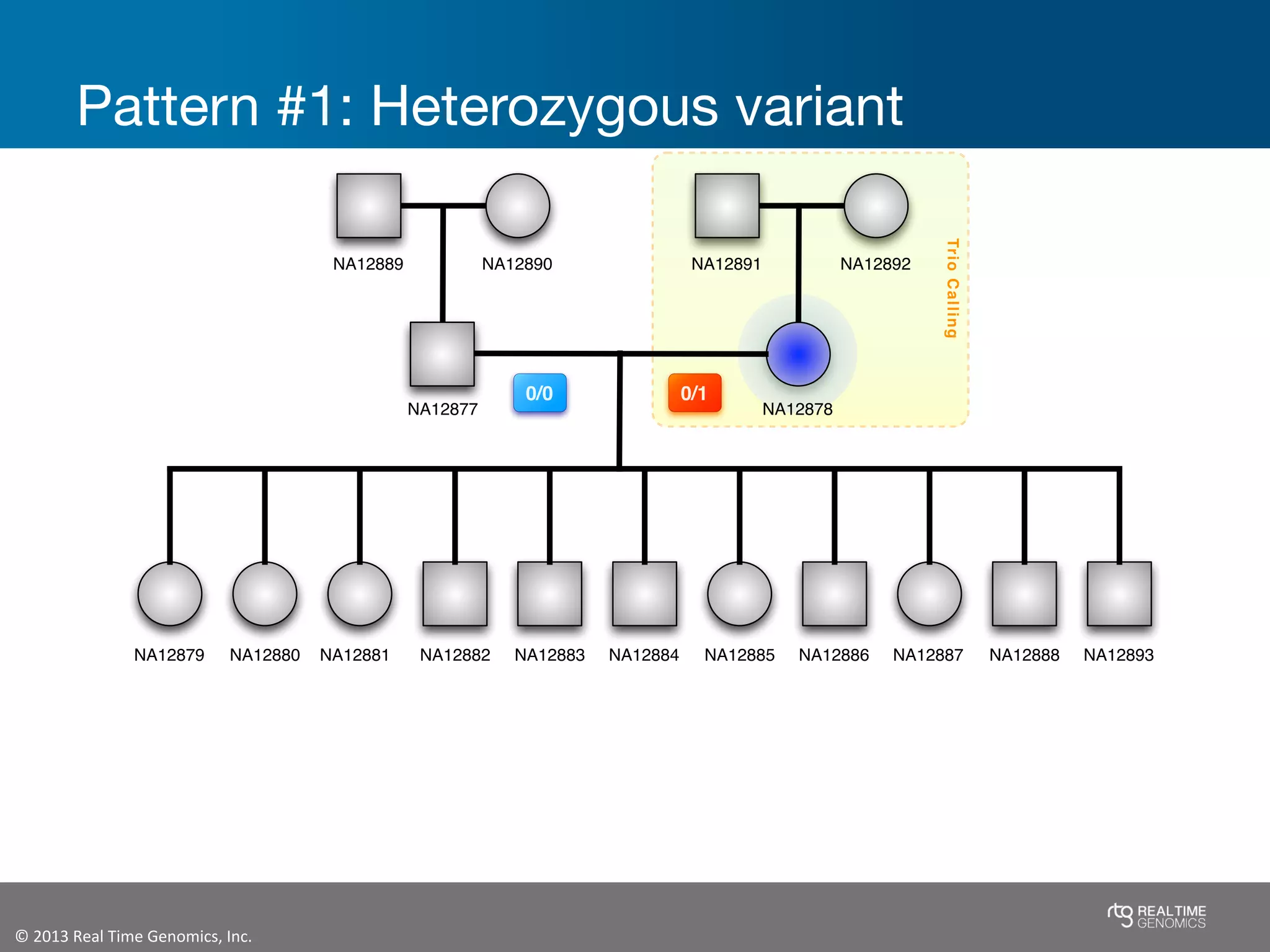
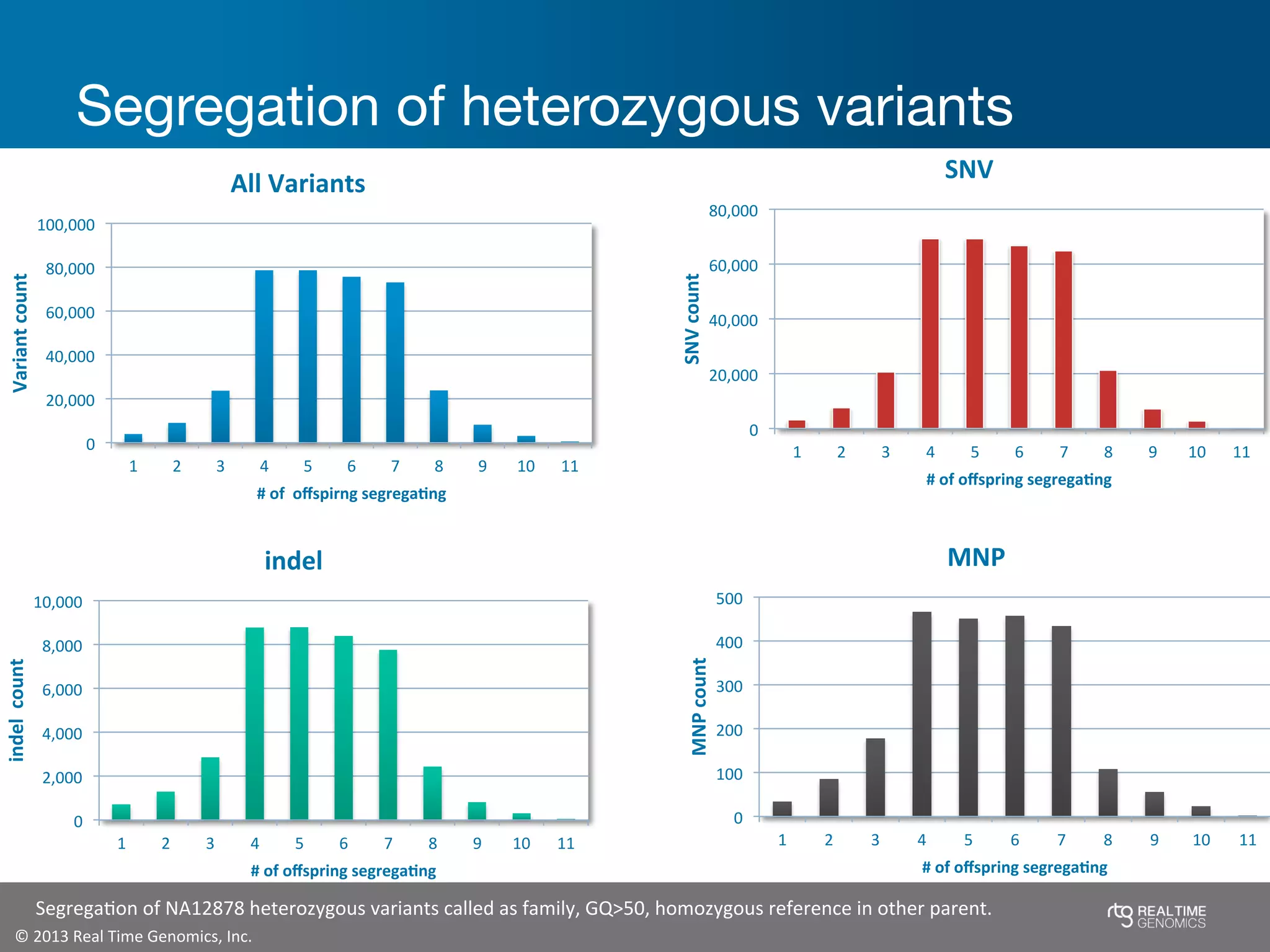
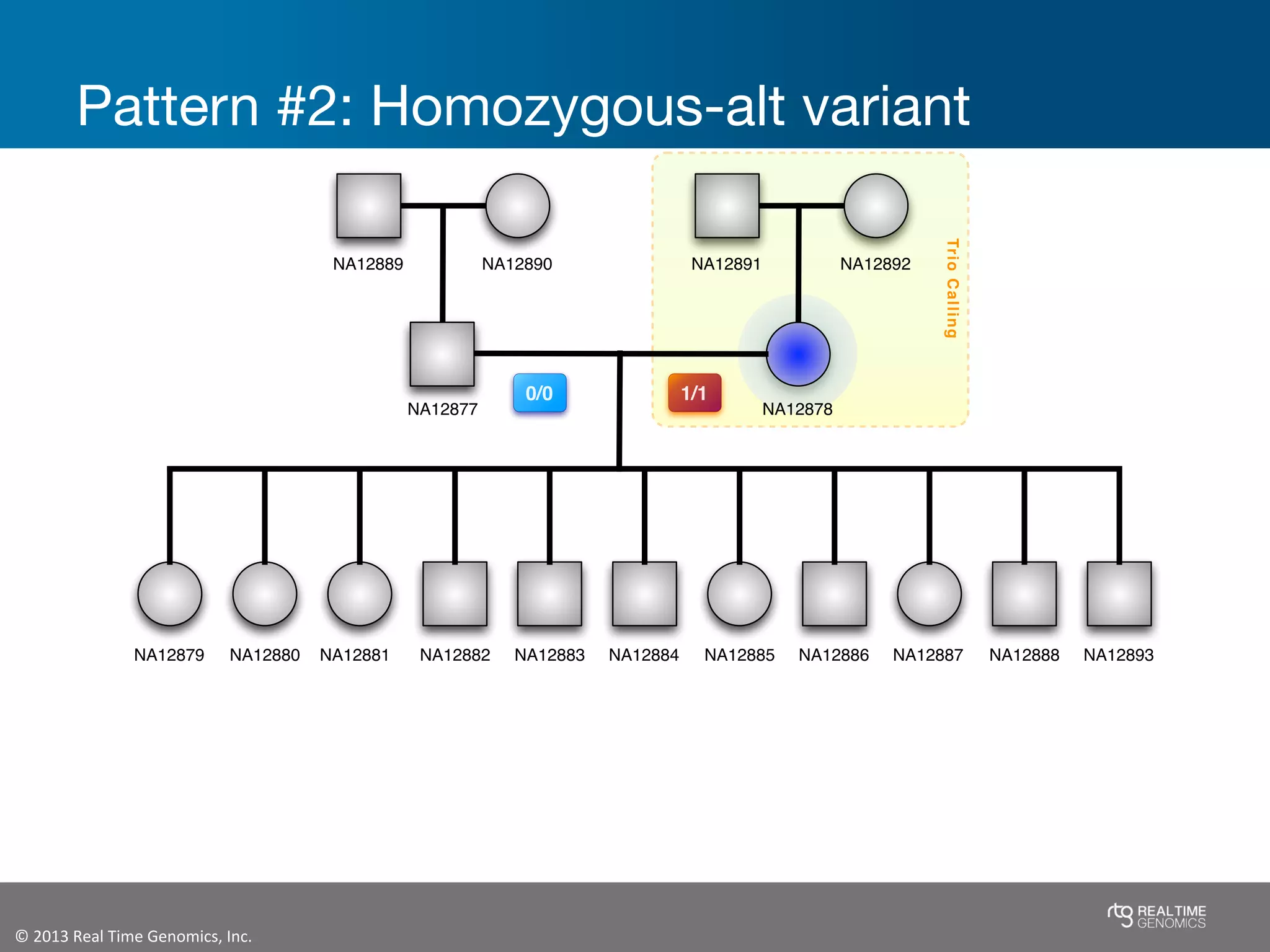
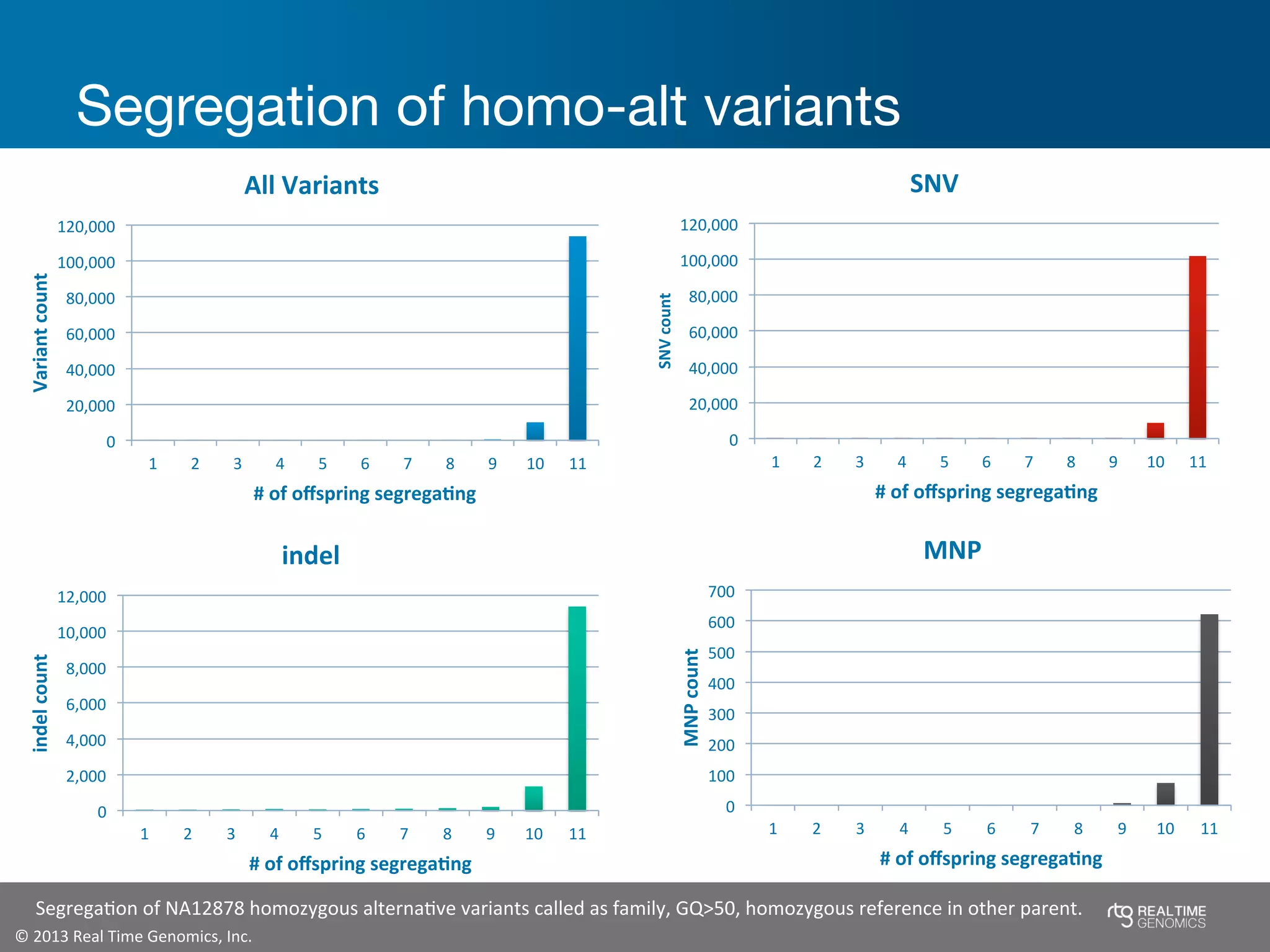
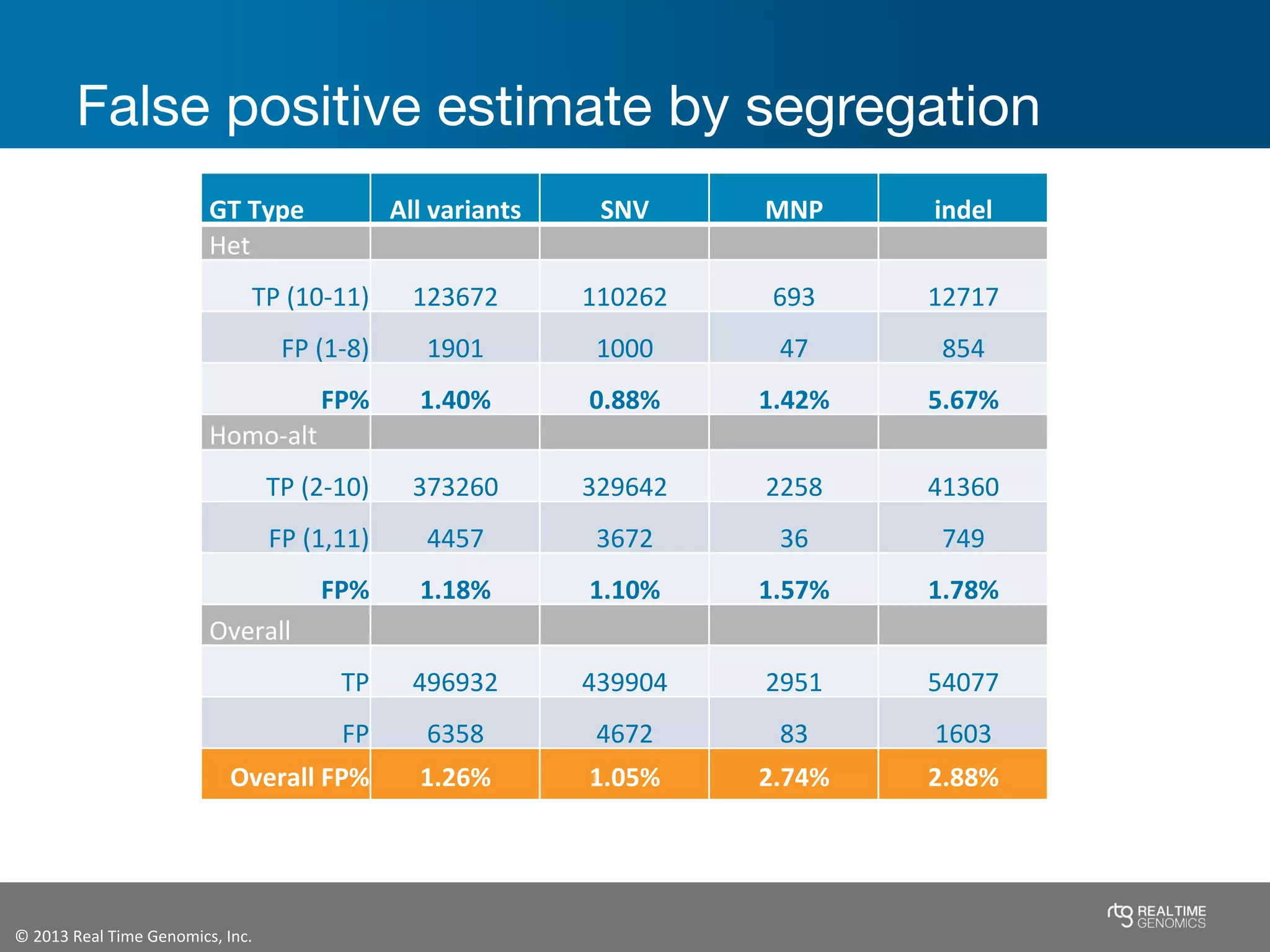
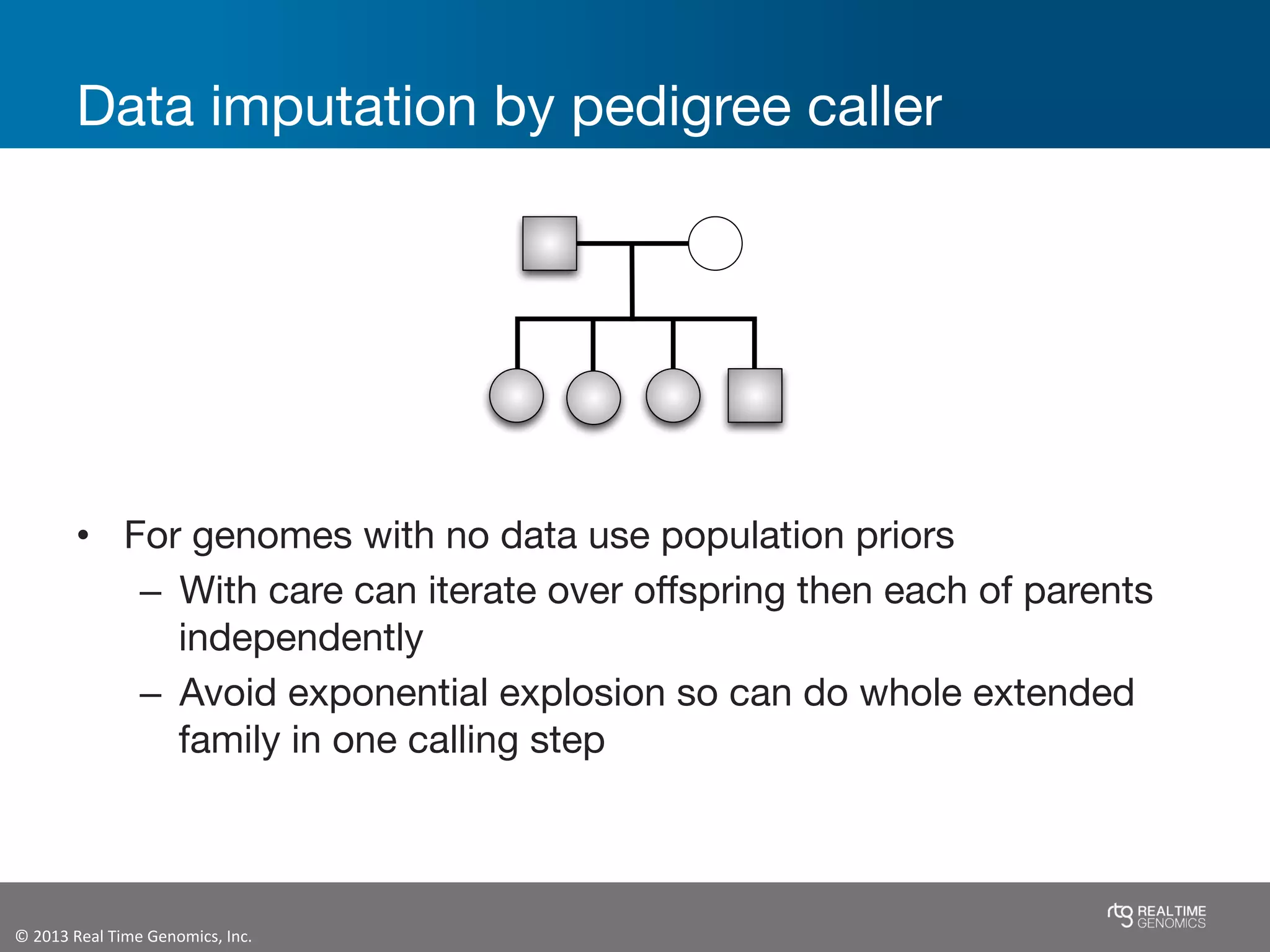
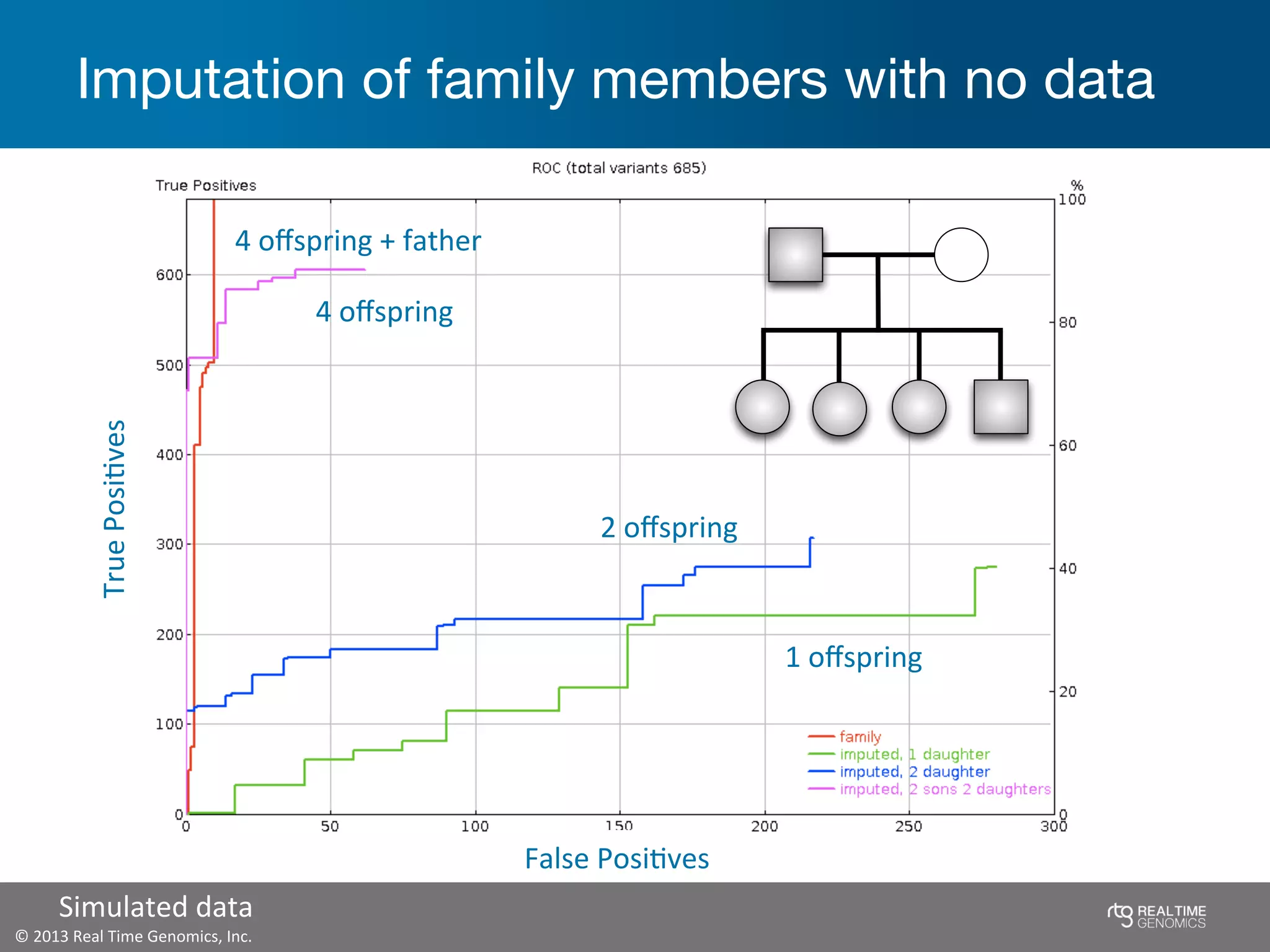
This document summarizes the analysis of variant calling using trio information from the NA12878 pedigree. It finds that jointly analyzing related samples using the RTG Family caller, which leverages Mendelian inheritance patterns and shared haplotypes, improves genotype calls by reducing errors compared to singleton calling. The trio analysis corrects over 60x more Mendelian inconsistency errors compared to singleton calling and identifies variants that segregate in offspring as expected for heterozygous and homozygous-alternate sites.