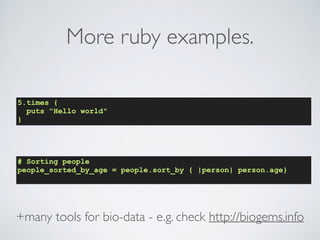
1. The document discusses best practices for scientific software development, including writing code for people rather than computers, automating repetitive tasks, using version control, and conducting code reviews.
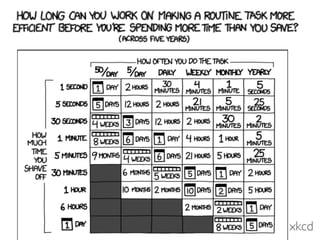
2. Specific approaches and tools recommended are planning for mistakes, automated testing, continuous integration, and using a coding style guide. R and Ruby style guides are provided as examples.
3. The benefits of following such practices are improving productivity, reducing errors, making code easier to read and maintain, and allowing scientists to focus on scientific questions rather than software issues. Reproducible and sustainable software is the overall goal.











![Best Practices for Scientific Computing
Greg Wilson ∗
, D.A. Aruliah †
, C. Titus Brown ‡
, Neil P. Chue Hong §
, Matt Davis ¶
, Richard T. Guy ∥
,
Steven H.D. Haddock ∗∗
, Katy Huff ††
, Ian M. Mitchell ‡‡
, Mark D. Plumbley §§
, Ben Waugh ¶¶
,
Ethan P. White ∗∗∗
, Paul Wilson †††
∗
Software Carpentry (gvwilson@software-carpentry.org),†University of Ontario Institute of Technology (Dhavide.Aru
State University (ctb@msu.edu),§Software Sustainability Institute (N.ChueHong@epcc.ed.ac.uk),¶ Space Telescope
(mrdavis@stsci.edu),∥University of Toronto (guy@cs.utoronto.ca),∗∗Monterey Bay Aquarium Research Institute
(steve@practicalcomputing.org),††University of Wisconsin (khuff@cae.wisc.edu),‡‡University of British Columbia (mi
Mary University of London (mark.plumbley@eecs.qmul.ac.uk),¶¶University College London (b.waugh@ucl.ac.uk),∗∗
University (ethan@weecology.org), and †††University of Wisconsin (wilsonp@engr.wisc.edu)
Scientists spend an increasing amount of time building and using
software. However, most scientists are never taught how to do this
efficiently. As a result, many are unaware of tools and practices that
would allow them to write more reliable and maintainable code with
less effort. We describe a set of best practices for scientific software
development that have solid foundations in research and experience,
and that improve scientists’ productivity and the reliability of their
software.
Software is as important to modern scientific research as
telescopes and test tubes. From groups that work exclusively
on computational problems, to traditional laboratory and field
scientists, more and more of the daily operation of science re-
volves around computers. This includes the development of
new algorithms, managing and analyzing the large amounts
of data that are generated in single research projects, and
combining disparate datasets to assess synthetic problems.
Scientists typically develop their own software for these
purposes because doing so requires substantial domain-specific
and open source software development [61
ical studies of scientific computing [4, 31,
development in general (summarized in
practices will guarantee efficient, error-fr
ment, but used in concert they will red
errors in scientific software, make it easie
the authors of the software time and effo
focusing on the underlying scientific ques
1. Write programs for people, not c
Scientists writing software need to write
cutes correctly and can be easily read and
programmers (especially the author’s fut
cannot be easily read and understood it is
to know that it is actually doing what it i
be productive, software developers must t
aspects of human cognition into account
arXiv:1210.0530v3[cs.MS]29Nov2012
1. Write programs for people, not computers.
2. Automate repetitive tasks.
3. Use the computer to record history.
4. Make incremental changes.
5. Use version control.
6. Don’t repeat yourself (or others).
7. Plan for mistakes.
8. Optimize software only after it works correctly.
9. Document the design and purpose of code rather than its mechanics.
10. Conduct code reviews.
Scientists spend an increasing amount of time building and using
software.
However, most scientists are never taught how to do this
efficiently.
We describe a set of best practices for scientific software
development that have solid foundations in research and
experience, and that improve scientists' productivity and the
reliability of their software.](https://image.slidesharecdn.com/2015-10-7-11am-reproducibleresearch-151007091948-lva1-app6892/85/2015-10-7-11am-reproducible-research-12-320.jpg)





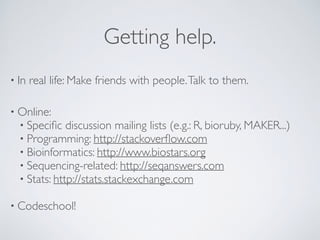
![knitr (sweave)Analyzing & Reporting in a single file.
analysis.Rmd
### in R:
library(knitr)
knit(“analysis.Rmd”)
# --> creates analysis.md
### in shell:
pandoc analysis.md -o analysis.pdf
# --> creates MyFile.pdf
A minimal R Markdown example
I know the value of pi is 3.1416, and 2 times pi is 6.2832. To c
library(knitr); knit( minimal.Rmd )
A paragraph here. A code chunk below:
1+1
## [1] 2
.4-.7+.3 # what? it is not zero!
## [1] 5.551e-17
Graphics work too
library(ggplot2)
qplot(speed, dist, data = cars) + geom_smooth()
●
●
●
●
●
●
●
●
●
●
●
●
●●● ●
●●
●
●
●
●
●
●
●
●
●
●
●
●
●
●
●
●
●
●
●
●
●
●●●
● ●
0
40
80
120
5 10 15 20
speed
dist
Figure 1: A scatterplot of cars](https://image.slidesharecdn.com/2015-10-7-11am-reproducibleresearch-151007091948-lva1-app6892/85/2015-10-7-11am-reproducible-research-22-320.jpg)