JBEI April 2021 - Research Highlights
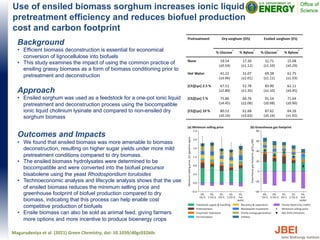
- 1. Use of ensiled biomass sorghum increases ionic liquid pretreatment efficiency and reduces biofuel production cost and carbon footprint Background • Efficient biomass deconstruction is essential for economical conversion of lignocellulose into biofuels • This study examines the impact of using the common practice of ensiling grassy biomass as a form of biomass conditioning prior to pretreatment and deconstruction Approach • Ensiled sorghum was used as a feedstock for a one-pot ionic liquid pretreatment and deconstruction process using the biocompatible ionic liquid cholinium lysinate and compared to non-ensiled dry sorghum biomass Outcomes and Impacts • We found that ensiled biomass was more amenable to biomass deconstruction, resulting on higher sugar yields under more mild pretreatment conditions compared to dry biomass. • The ensiled biomass hydrolysates were determined to be biocompatible and were converted into the biofuel precursor bisabolene using the yeast Rhodosporidium toruloides • Technoeconomic analysis and lifecycle analysis shows that the use of ensiled biomass reduces the minimum selling price and greenhouse footprint of biofuel production compared to dry biomass, indicating that this process can help enable cost competitive production of biofuels • Ensile biomass can also be sold as animal feed, giving farmers more options and more incentive to produce bioenergy crops Magurudeniya et al. (2021) Green Chemistry, doi: 10.1039/d0gc03260c
- 2. A SWEET surprise: Anaerobic fungal sugar transporters and chimeras enhance sugar uptake in yeast Background • Membrane-embedded transport proteins govern carbon uptake into cell factories such that the activity of these transporters can dictate the productivity of production strains. • This study addresses the transport challenge of lignocellulosic hydrolysate fermentation by the yeast Saccharomyces cerevisiae, namely poor co-utilization of the abundant xylose sugar, by evaluating the performance of heterologously expressed anaerobic fungal SWEETs. Approach • Anaerobic fungal SWEET transporters were heterologously expressed in a transporter-deficient strain to screen sugar transport activity by growth and sugar consumption. • Single crossover chimeras sampled the functional qualities of SWEETs that failed to functionally expressed. Outcomes and Impacts • The anaerobic fungal transporter NcSWEET1 was functionally expressed in S. cerevisiae, demonstrating broad activity on hexose sugars and novel activity on xylose. • A leading NcSWEET1 expression enabling domain recovered robust chimeras that improved performance on hexose sugars • By manipulating the crossover junction location, a narrow set of residues that control substrate specificity were identified. • NcSWEET1 mediated co-utilization of glucose and xylose outperformed the engineered HXT7(F79S) control, highlighting the utility of SWEETs as yeast strain engineering tools. Podolsky et al. (2021) Metabolic Engineering, doi: https://doi.org/10.1016/j.ymben.2021.04.009 SWEET chimeras sampled characteristics of poorly trafficked homologues by using a leading NcSWEET1 domain. The wild- type NcSWEET1 demonstrated robust xylose consumption. 1 3 2 5 7 6 4 Domain 1 Domain 2 5 µm NcSWEET1 PfSWEET2 10 20 30 0 4 8 12 OD600 [Glucose] [Xylose] 10 20 30 0 4 8 12 0 24 48 72 96 120 Concentration (g/L) OD 600 Time (h) NcSWEET1
- 3. Integrating systems and synthetic biology to understand and engineer microbiomes Background • Microbiomes, diverse communities of microorganisms, inhabit every known environment, from oceans and soil to the intestines of humans and animals, and perform myriad functions including biogeochemical nutrient cycling and conversion of dietary substrates within multicellular hosts. • Leveraging microbiomes for environmental and societal benefit in agriculture, medicine, and bioproduction demands holistic understanding of the various biochemical, molecular, spatial, and ecological factors that dictate their composition and function. Approach • This review highlights recent developments in meta-omics, modeling, and engineering tools that enable us to understand and manipulate microbiome composition and function, with emphasis on the ecological parameters that drive community behavior and must be considered for predictive microbiome design. Leggieri et al. (2021) Annual Review in Biomedical Engineering, doi: 10.1146/annurev-bioeng-082120-022836 Microbiome engineering tools can be used to modify existing functions within the community or to introduce novel functions on many different scales, from editing a single genome to an introducing an entirely new engineered community. Development and deployment of these tools is facilitated by integrating multiple meta- omics analyses to develop causal links between microbiome composition and function. Outcomes and Impacts • Continuously improving meta-omics tools allow comprehensive characterization of environmental and cultured samples and elucidate strategies for tailoring the performance of microbial communities. • Computational models of microbiomes synthesize vast empirical data to predict new strategies to accentuate desired microbiome functions. Experimental validation is irreplaceable for model validation and improvement; obtaining key metrics such as temporal stability, growth rates, and fluxes remains a challenge. • Owing to its complexity, microbiome engineering requires versatile tools that allow the precise introduction of targeted traits into complex communities with predictable outcomes.
- 4. Reprogramming sphingolipid glycosylation is required for endosymbiont persistence in Medicago truncatula Background • Plant endosymbiosis relies on the development of specialized membranes that encapsulate the endosymbiont and facilitate nutrient exchange. • However, the identity and function of lipids within these membrane interfaces is largely unknown. Approach • Here, we identify GLUCOSAMINE INOSITOL PHOSPHORYLCERAMIDE TRANSFERASE1 (GINT1) as a sphingolipid glycosyltransferase highly expressed in Medicago truncatula root nodules and roots colonized by arbuscular mycorrhizal (AM) fungi and further demonstrate that this enzyme functions in the synthesis of N-acetyl-glucosamine-decorated glycosyl inositol phosphoryl ceramides (GIPCs) in planta. Outcomes and Impacts • MtGINT1 expression was developmentally regulated in symbiotic tissues associated with the development of symbiosome and periarbuscular membranes. • RNAi silencing of MtGINT1 did not affect overall root growth but strongly impaired nodulation and AM symbiosis, resulting in the senescence of symbiosomes and arbuscules. • Our results indicate that, although M. truncatula root sphingolipidome predominantly consists of hexose-decorated GIPCs, local reprogramming of GIPC glycosylation by MtGINT1 is required for the persistence of endosymbionts within the plant cell. • Understanding communication between plants and AM fungi is important for the engineering of bioenergy crops with environmental resilience and increased carbon sequestration Moore et al. (2021) Current Biology, doi: 10.1016/j.cub.2021.03.067 Glycosylinositol phosphoryl ceramides (GIPCs) are major plasma membrane lipids in plants. GINT1 adds N-acetylglucosamine to GIPCs. In M. truncatula this form of GIPC is found in endosymbiotic tissues whereas other tissues have GIPCs with mannose. Additional sugars can be added to GIPCs (not shown) Cer Ins IPUT1 GINT1 GMT1 P Ceramide Ins Cer Ins P Cer Ins P GlcN(Ac) Man GlcA P Dowregulation of GINT1 results in inability of the plant to develop functional symbioses with N-fixing Rhizobia (left) and arbuscular mycorrhizal fungi (right). The symbionts will in both cases enter the roots, i.e. the initial signaling processes are functioning, but the symbiotic structures do not fully develop. Nodules cannot fix nitrogen (measured as C2H4 production from acetylene), and arbuscles are few and senesce quickly. * * * 0 0.05 0.1 0.15 0.2 0.25 0.3 0.35 14 dpi 28 dpi C 2 H 4 (nmol/h) GUS-RNAi MtGINT1-RNAi
- 5. In-planta production of the biodegradable polyester precursor 2-pyrone- 4,6-dicarboxylic acid (PDC): Stacking reduced biomass recalcitrance with value-added co-product Background • 2-Pyrone-4,6-dicarboxylic acid (PDC) is a promising building block for the manufacturing of performance-advantaged polymers. • Reduction of lignin and in-planta production of chemicals in bioenergy crops are two desired traits to improve the economics of second-generation biofuels and bioproducts. • This study demonstrates an approach to concomitantly reduce lignin and produce PDC in the model Arabidopsis. Approach • A five-gene construct was designed for overproduction of protocatechuate (PCA) and its conversion into PDC at the expense of lignin. • Bacterial genes encoding AroG, QsuB, PmdA, PmdB, and PmdC were fused to signal peptide sequences for targeting to plastids where the shikimate pathway occurs. • Several independent transgenic lines were obtained and analyzed for lignin content, PDC production, and yields of sugars after saccharification. Outcomes and Impacts • Most transgenic lines are healthy and show no yield penalty. • These lines accumulate PDC (up to 3% DW), have 30-45% less lignin, and release more sugars upon cellulase and xylanase treatment (up to +75%). • A construct containing the same bacterial genes placed under the control of monocot-specific promoters has been designed for sorghum transformation. • This study will guide the design of bioenergy crops with reduced recalcitrance and accumulation of co-products Lin et al. (2021) Metabolic Engineering, doi: 10.1016/j.ymben.2021.04.011 WT Transgenic lines High PCA / PDC WT Transgenic lines Low lignin PDC de-novo pathway WT Transgenic lines No yield penalty **P < 0.001
- 6. Background • Lignin peroxidases (LiP) catalyze cleavage of both β-O-4’ ether and C–C bonds in lignin and are essential for depolymerizing lignin into fragments amendable to biological upgrading to valuable products. • This study reported on the effect of pH on the LiP-catalyzed cleavage of β-O-4’ ether C–C bonds in LiP isozyme H8 from Phanerochaete chrysosporium and an acid stabilized variant of it. Approach • Nanostructure initiator mass spectrometry assay to quantify bond breaking in a phenolic model lignin dimer (LigNIMS assay). • Ab initio molecular dynamics simulations and climbing-image nudge elastic band based transition state searches to test hypothesis that the effect of lower pH is via protonation of aliphatic hydroxyl groups of the lignin-like dimer. Outcomes and Impacts • Degradation of the lignin like dimer to products by an acid- stabilized variant of LiP isozyme H8 increased from 38.4 % at pH 5 to 92.5% at pH 2.6. At pH 2.6, the observed product distribution resulted from 65.5% β-O-4’ ether bond cleavage, 27.0% Ca-C1 carbon bond cleavage. • Quantum calculations confirmed protonation of aliphatic hydroxyl groups under extremely acidic conditions resulted in lower energetic barriers for bond-cleavages. • Lignin protonation implies catalysis of lignin depolymerization will be more efficient at low pH, independent of the catalyst. Pham L., Deng K., Northen T. R., Singer S. W., Adams P. D., Simmons B. A., Sale K. L. (2021) “Experimental and theoretical insights into the effects of pH on catalysis of bond-cleavage by the lignin peroxidase isozyme H8 from Phanerochaete chrysosporium.” Biotechnology for biofuels, 14(1), 108. doi: 10.1186/s13068-021-01953-7 Experimental and theoretical insights into the effects of pH on catalysis of bond-cleavage by the lignin peroxidase isozyme H8 from Phanerochaete chrysosporium pH dependence of lignin peroxidase from Phanerochaete chrysosporium Proposed mechanism for b-O-4’ ether bond cleavage from protonated Ca-OH, GGE cationic radical
- 7. Papers Enabled by JBEI Technologies
- 8. Leveling the cost and carbon footprint of circular polymers that are chemically recycled to monomer Background • Mechanical recycling of polymers downgrades them such that they are unusable after a few cycles • Chemical recycling to monomer offers a means to recover the embodied chemical feedstocks for remanufacturing • Designing novel polymers that can be made with bio-based materials can reduce the energy footprint of chemical recycling to virgin-quality monomers Approach • We use systems analysis to quantify the costs and life-cycle carbon footprints of virgin and chemically recycled polydiketoenamines (PDKs), next-generation polymers that depolymerize under ambient conditions in strong acid • By developing process simulations and exploring long term shifts in system-wide costs and emissions, we can establish targets for making PDKs competitive with conventional plastics Outcomes and Impacts • We find that the cost of producing virgin PDK resin using unoptimized processes is ~30-fold higher than recycling them • The cost of recycled PDK resin ($1.5 kg−1 ) is on par with PET and HDPE, and below that of polyurethanes • Virgin resin production is carbon intensive (86 kg CO2e kg−1 ), while chemical recycling emits only 2 kg CO2e kg−1 • This cost and emissions disparity provides a strong incentive to recover and recycle future polymer waste • Future adjustments in the chemistry and a shift toward greater bio- derived content will be required to reduce the GHG footprint and cost of virgin PDK Vora et al. (2021) Science Advances, doi: 10.1126/sciadv.abf0187 Figure 2. Minimum selling price and life-cycle GHG footprint for different conventional and PDK scenarios. Figure 1. System boundary for chemically recyclable PDK
- 9. Further engineering of R. toruloides for the production of terpenes from lignocellulosic biomass Background • Terpenes encompass a wide variety of molecules with a diverse range of commercial applications, including biofuels • Efficient production of terpenes from cheap carbon sources, such as lignocellulose, is essential for them to be economical • This study focuses on optimizing terpene production in a conversion host capable of converting a wide range of renewable carbon, Rhodosporidium toruloides Approach • To improve terpene production, strains producing the biofuel precursor bisabolene was subjected to multi-omic analysis, and the data generated was leveraged to identify specific enzymes in the mevalonate pathway for optimization Outcomes and Impacts • Proteomic, metabolomic, and transcripomic analysis were conducted on several bisabolene-producing strains and several mevalonate enzymes were identified for optimization, including HMG-CoA reductase, mevalonate kinase, phosphomevalonate kinase • The optimization was conducted for two terpenes, 1,8-cineole and bisabolene, and titers were improved for both terpenes to 1.4 g/L and 2.6 g/L, respectively Kirby et al. (2021) Biotechnology for Biofuels, doi: 10.1186/s13068-021-01950-w