JBEI Research Highlights - March 2022
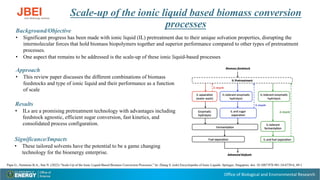
- 1. Office of Biological and Environmental Research Scale-up of the ionic liquid based biomass conversion processes Background/Objective • Significant progress has been made with ionic liquid (IL) pretreatment due to their unique solvation properties, disrupting the intermolecular forces that hold biomass biopolymers together and superior performance compared to other types of pretreatment processes. • One aspect that remains to be addressed is the scale-up of these ionic liquid-based processes Approach • This review paper discusses the different combinations of biomass feedstocks and type of ionic liquid and their performance as a function of scale Results • ILs are a promising pretreatment technology with advantages including feedstock agnostic, efficient sugar conversion, fast kinetics, and consolidated process configuration. Significance/Impacts • These tailored solvents have the potential to be a game changing technology for the bioenergy enterprise. Papa G., Simmons B.A., Sun N. (2022) “Scale-Up of the Ionic Liquid-Based Biomass Conversion Processes.” In: Zhang S. (eds) Encyclopedia of Ionic Liquids. Springer, Singapore. doi: 10.1007/978-981-10-6739-6_49-1
- 2. Office of Biological and Environmental Research Applications of targeted proteomics in metabolic engineering: advances and opportunities Background/Objective • Optimization of metabolically engineered organisms requires good understanding of producing balanced level of pathway proteins. • Targeted proteomics via selected-reaction monitoring (SRM) has been increasingly used in metabolic engineering research to detect and quantify sets of proteins with high selectivity and multiplexity. Approach • In this review, we present recent applications of targeted proteomics in metabolic engineering research and highlight several successful studies of targeted proteomics in production of high value chemicals. • We also discuss challenges and limitations of current targeted proteomics and map opportunities for future research. Results • Targeted proteomics has been a useful tool in metabolic engineering to bring a more predictable and rational engineering of biology. Significance/Impacts • Integration of high quality and accuracy protein expression data input from targeted proteomics and other omics tools to machine learning will become an avenue of interest in many metabolic engineering studies. Yunus I.S., Lee, T.S. (2022) Current Opinion in Biotechnology. doi: 10.1016/j.copbio.2022.102709
- 3. Office of Biological and Environmental Research Scalable and automated CRISPR-based strain engineering using droplet microfluidics Background/Objective • CRISPR-MAGE allows one to make many simultaneous changes to an organism’s chromosome across a population of cells, but this method is difficult to use for high-throughput efforts. • Droplet microfluidics has become an attractive option for performing such large scale synthetic biology experiments. Approach • We developed a droplet microfluidic system that miniaturizes and automates CRISPR-MAGE. The system can carry out up to 100 genetic modification reactions in parallel. Results • The system proved to be effective in two test cases: (1) to successfully disrupt the function of an enzyme found in E. coli, and (2) to modify E. coli to produce indigoidine, a sustainable alternative to traditional indigo. Significance/Impacts • This is a scalable platform for producing large numbers of engineered strains, which are needed to optimize genetic pathways and perform predictable bioengineering. Iwai, K., et al. (2022) Microsyst Nanoeng, doi: 10.1038/s41378-022-00357-3 CRISPR-MAGE steps. The ones inside the box are performed on the chip. Cells are removed from the chip after the recovery step for induction and plating. b The microfluidic chip in a 3D printed holder (left), the electrode pattern (right), a top-view of an individual well, and a side-view schematic of a well. The chip is designed to contain 100 discrete reaction chambers with individually addressable electrodes for multiplexed CRISPR-MAGE recombineering, and its 384-well format design can be interfaced with lab automation equipment. c Droplets containing plasmids and cells are dispensed into each chamber through the inlet port, mixed by electrowetting, and electroporated by applying a voltage pulse
- 4. Office of Biological and Environmental Research Continental United States may lose 1.8 petagrams of soil organic carbon under climate change by 2100 Background/Objective • High-resolution information on soils’ vulnerability to climate-induced soil organic carbon (SOC) loss can enable environmental scientists, land managers, and policy makers to develop targeted mitigation strategies Approach • We used recent SOC field observations, environmental factors, and an ensemble machine learning (ML) approach to estimate baseline SOC stocks in surface soils across the continental United States at 100-m spatial resolution, and decadal changes under the projected climate scenarios of CMIP6 earth system models (ESMs). Results • Baseline SOC projections from ML approaches captured more than 50% of variability in SOC observations, whereas ESMs represented only 6–16% of observed SOC variability (Figure 1). • Both ML and ESM predictions agree on the direction of SOC change (net emissions or sequestration) across 46–51% of continental US land area (Figure 2). Significance/Impacts • ML estimates showed a mean total loss of 1.8 Pg C from US surface soils under the high-emission scenario by 2100, whereas ESMs showed no significant change in SOC stocks with wide variation among ESMs. Gautam et al. (2022) Global Ecology & Biogeography, https://doi.org/10.1111/geb.13489 Figure 1 Figure 2
- 5. Office of Biological and Environmental Research Lignin synthesis and bioengineering approaches toward lignin modification Background/Objective • Lignin plays important roles in plants, but often represents hurdles to the utilization of cellulosic biomass for bioenergy. • Understanding lignin synthesis is a prerequisite to manipulate its content and composition in crops. Approach • The recent literature on lignin biosynthesis and engineering has been reviewed. Results • With the advance of synthetic biology, it has become feasible to fine-tune lignin content and composition by tailoring gene expression, exploiting enzyme post-translational modifications, altering enzyme cofactors, and rerouting lignin metabolic precursors. Significance/Impacts • Novel bioengineering strategies can be implemented in bioenergy crops to facilitate biomass conversion into bioproducts. Liu C.J., Eudes A. (2022) “Lignin synthesis and bioengineering approaches toward lignin modification” Book Chapter in Advances in Botanical Research, Lignin and hydroxycinnamic acids: biosynthesis and the buildup of the cell wall. doi: 10.1016/bs.abr.2022.02.002 Figure 1: Current view of the lignin biosynthetic pathway. At least thirteen enzymes have been involved in monolignol synthesis. Figure 2: Chemical structures of lignin acyl groups (pCA: p-coumarate; HBA: 4-hydroxybenzoate) and potential alternative cleavable lignin monomers.
- 6. Office of Biological and Environmental Research Nitrogen metabolism in Pseudomonas putida: functional analysis using RB-TnSeq Background/Objective • Pseudomonas putida is an attractive host for the valorization of biomass hydrolysates • Many nitrogenous target bioproducts, such as lactams and amino acids, can serve as a source of nitrogen for P. putida • To efficiently engineer the host, an extensive knowledge of its metabolism is required Approach • Using pooled mutant fitness assays, this study identifies genes involved in the assimilation of 52 different nitrogen containing compounds. Further validation experiments include biochemical enzyme characterization, biosensor development, and mutant growth assays Results • We provide evidence for known nitrogen assimilation pathways and their regulatory systems, and assign functions to genes whose exact roles in nitrogen metabolism were not previously known Significance/Impacts • The presented functional genomics data is contextualized within the literature spanning decades of research about nitrogen metabolisms. • This work underlines the utility of functional genomics in the context of metabolic engineering Schmidt and Pearson et al. (2022) Applied and Environmental Microbiology doi: 10.1128/aem.02430-21 Overview of RB-TnSeq and data processing workflow. Competitive growth assays using a pooled and barcoded transposon mutant library are conducted under different nitrogen conditions. Fitness values are calculated from barcode frequencies and clustered using the manifold learning method t-SNE