Non covalent interactions
- 1. Organic Pedagogical Electronic Network Non-Covalent Interactions Alexandra Kent & Allyson Brome University of Utah
- 2. Non Covalent Interactions Wiki Page: http://en.wikipedia.org/wiki/Non-covalent_interactions Other References: Anslyn, E. V., Dougherty, D. A. (2006). Modern Physical Organic Chemistry. Sausalito, CA. University Science. Overview: Non covalent interactions are incredibly important characteristics in supermolecular chemistry, especially in biological molecules, such as nucleic acids and enzymes. These can include pi-pi, pi-cation and electrostatic interactions. Pi-Pi Interactions Pi-Pi interactions in benzene can be observed in three different ways. These occur when pi orbitals overlap in favorable orientations. Cation-Pi Interactions Cation-pi interactions are a strong interaction between electrons from the pi system and a cation. These interactions have the potential to be as strong or stronger than H- bonding. They are detected by a large upfield shift in the H-NMR spectra, because the protons are shielded by the pi system.
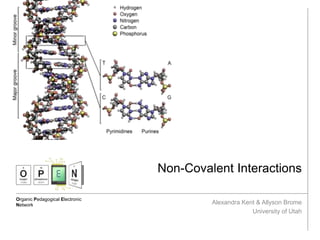
- 3. Examples DNA Helix Pi stacking interactions are incredibly important in the stabilization of a DNA double helix. The nucleobase stacking can even influence the interstrand hydrogen bonding that glues together the helical structure.
- 4. Examples G-quadruplex G-quadruplex architecture is a more complex example that incorporates cation-pi as well pi stacking interactions. These can be used for biosensing applications. Wiki Page: http://en.wikipedia.org/wiki/G-quadruplex Other References: Matta, C. F.; Castillo, N.; Boyd, R.; J. Phys. Chem. 2006, 110, 563-578
- 5. Molecular Tweezers References: Landon P. B.; Ramachandran, S.; Gillman, A.; Gidron, T; Yoon, D.; Lal, R.; Langmuir, 2012, 28, 53- 540 One of the more interesting biological examples uses the non covalent interactions in nucleic acids for non traditional applications. Molecular Tweezers are short sequences of DNA that can be used as a renewable system of detection for DNA sequences. They can be integrated with DNA nanomachines for lab-on-a-chip utilization. Molecular tweezers utilize the additive properties of intermolecular forces. Inosine, a guanine analogue, is present in the red part of the original sequence above. Therefore, in the presence of the complementary blue strand, the tweezers open because binding of the blue strand is favored over the inosine mutated red strand. However, this blue strand has a base overhang. In accordance with the additive nature of intermolecular forces, the addition of a complementary sequence that can bind to that overhang will displace the blue sequence, closing the tweezers. This opening and closing event can be detected using a fluorophore- quencher pair. This process, as shown by the data on the left, is regenerative and the tweezers can be used multiple times.
- 6. Problems Wiki Page: http://en.wikipedia.org/wiki/Ethidium_bromide Other References: Zhang, L.; Peritz, A.; Meggers, E. J. Am. Chem. Soc. 2005, 127, 4174-4175; Haeusler et. al, Nature, 2013, 507, 195-200 1. In DNA it is a well known fact that the sugar phosphate backbone contributes some stability to the alpha-helical structure. However, it has been shown that this sugar phosphate backbone can be replaced with a glycol backbone, and this GNA molecule maintains a helical structure. Propose an explanation for this phenomena. 2. Water is a known fluorescence quencher. Ethidium bromide is non-fluorescent in aqueous solution, but upon the addition of double stranded DNA, small fluorescence is observed. What is occurring to cause the fluorescence? 3. The DNA sequence, (GGGGCC)4 will naturally form a hairpin loop, however in the presence of KCl it can form a G-quadruplex structure. Why does this occur?
- 7. Solutions 1. This is observed because the helical structure is mainly stabilized through pi- stacking and hydrogen bonding interactions of the base pairs, despite the added stability of the sugar phosphate backbone. 2. The ethidium bromide is intercalating into the DNA strand which is a more hydrophobic environment. This causes it to shed the water molecules, allowing it to fluoresce. 3. This structure is stabilized by the KCl because the K+ is favorably interacting with the pi system, which favors the g-quadruplex structure more so than the hairpin loop.
- 8. This work is licensed under a Creative Commons Attribution- ShareAlike 4.0 International License. Contributed by: Alexandra Kent & Allyson Brome (Undergraduate Students) University of Utah 2014