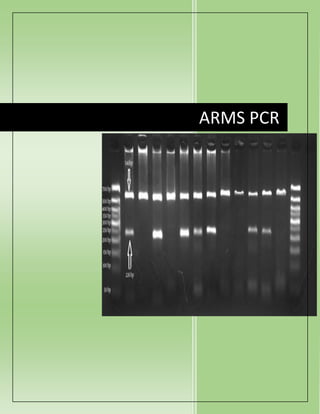
Detect Genetic Mutations with ARMS PCR
- 1. ARMS PCR
- 2. CONTENTS Introduction Principle ARMS Primer designer Procedure Importance of PCR in the Diagnosis of Genetic Disease Advantages of ARMS PCR Limitations of ARMS PCR
- 3. INTRODUCTION The allele-specificPCRalsocalledasan ARMS- PCR (amplification refractory mutation system) orPASA (PCRamplification of specificalleles) orAS-PCRusedtodetectthe SNPs. More specifically, itis adopted todetectthe known SNPs(single nucleotide polymorphism). The term suggeststhatthe technique usedinthistype of PCRisspecifictothe particulartype of allele. An allele isthe alternativeformof a gene. If one allele hasanSNPandthe otheralternative formis normal,we can analyse boththe allelesbydesigningspecificprimersforeachallele.Forthat,we have to modifythe single base atthe 3’ endof the primer(one primermatchesthe normal allele andone primermatchesthe mutantallele). The PCR is performedsimultaneouslyinasingle reaction.If amutantallele ispresent,thenthe PCR amplifiesthe mutantallele orif the normal allele ispresent,the normal allelewill amplify. AmplificationRefractoryMutationSystem(ARMS) isanapplicationof PCRinwhichDNA isamplifiedby allele specificprimers.InPCRmismatchatthe 3’ endof the primercan dramaticallyreduce the annealingandhence the amplification.Thisisdue tothe absence of 3’ to 5’ exonuclease proofreading activityof Taq polymerase.HighfidelityDNA polymerases,thathave thisactivity,cannotbe usedin ARMS. It isan extremelyusefulmethodforidentificationof pointmutationsorpolymorphisms.Since the ARMS PCR ismostlydone toidentifyamutationora polymorphismitisalsoimportantthatitshould be able to identifywhetherthe change inDNA isheterozygousorhomozygous.A heterozygoteor homozygote isdifferentiatedbyusingARMSprimersforthe mutant/polymorphicandthe normal (wild type) alleles.The reactionsforthe mutantandthe normal allelesare usuallycarriedoutinseparate tubes.Butthese maybe done inthe same tube after labellingthe twoprimerswithdifferent fluorescentdyes. Kary Mullis described the technique of in vitro amplification inthe year1983. Aftera few yearsof the discovery of the actual PCR technique, C. R. Newton and co-workers discovered the ARMS-PCRor allele-specificPCRtechnique. The technique ismajorlyused forthe genotyping of the single nucleotide polymorphismwiththe help of the refractoryprimers. ARMS-PCR,amplificationrefractorymutationsystempolymerase chainreaction
- 4. PRINCIPLE The mechanismof the ARMS PCR isbasedon the modification of the primersfordifferentalleles. Here, the 3’ end of the primers ismodified insuchaway that one setof the primercan amplify the normal allele andothers canamplify the mutantallele. The mismatchsingle base isintroduced atthe 3’ endof the primer. Thismismatchallowsthe primertoamplify one single allele. The concept of mismatch: Here the mismatchbetween the primerandthe template DNA playsacrucial role inachievingthe amplification. Introduction of a mismatchat the 3’ endof the primeraltersthe annealingtemperature forthat particularallele. Strong mismatch: G/A, C/T, T/T Mediummismatch: A/A, G/G,C/C, Weakmismatch: C/A, G/T Due to the absence of the exonuclease activityof TaqDNA polymerase,the mismatchcannotbe repaired.
- 5. ARMS PRIMER DESIGN 1. The ARMS PCRrequiresapair of primersincludingacommonandan ARMS primer.The commonprimerislike anyotherPCR primer.Butthe ARMS primerhas the followingspecial features: 2. The primeris usually30 basesin length. 3. The nucleotide atthe 3’ endof the primershouldbe complementarytothe targetnucleotide i.e., G forC or C forG and T for A or A forT. Mismatch at thispositioncandramaticallyreduce the amplification. A:G,G: A,and C:C mismatcheshave the worsteffectwhereasthe other mismatcheshave varyingdegreesof effect.For example, inamutationwithA-Tsubstitution the ARMS primerfor the mutantallele shouldhave the lastnucleotide complementarytothe nucleotide Ti.e., itshouldhave A.The primerforthe normal allele atthe same positionshould be complementarytothe nucleotide A i.e., itshouldhave Tasshowninthe figure below. 4. An additional mismatchatone of the lastfive nucleotidesof the ARMSprimerfurther increasesits specificity. 5. It iscustomaryto include aninternal PCRcontrol inARMS reactions.A pair of primersis designed toamplifyaregionof the gene of interestthatusuallyisfree of mutations.An amplificationof the internal control regionandnoamplificationbythe ARMSprimerindicate a true negative.Ina false negativeresultneitherthe ARMSprimernorthe internal control showsanyamplification.There couldbe several reasons for the false negative result e.g., too little or too much DNA, poorqualityof DNA template,failure toaddprimer,Taq,or other reagentsandpresence of PCRinhibitors. 6. The sensitivityandspecificityof anARMS reactioncanbe controlledbystringentreaction conditions.Goodprimerdesign,higherannealingtemperature andlimitednumberof cycles are importantinavoidingfalse results.The numberof cyclesshouldbe justenoughtogive a clearpositive result.Increasingthe numberof cyclesun-necessarilycancause false positives. The usual lengthof ARMS primeris30 bases.Primersof thislengthhave ahighTm and annealingtemperature andare therefore more specific.
- 6. ARMS primersforthe normal and the mutant allele of apointmutation,IVSI-5(G-C),inthe β-globin gene.SegmentA showsthe matchedARMSprimerforthe mutation(C),segmentBshowsthe mismatchbetweenthe mutantARMSprimerandthe normal sequence (G), segmentCshowsthe matchedARMS primerforthe normal sequence (G),segmentDshowsthe mismatchbetweenthe normal ARMS primerandthe mutation(C).A deliberate mismatch(T:T) isalsoaddedat position enclosedbyblue box. PCR conditions for ARMS Locus: β-globingene GenBank accession NG_000007.3 Allele: β-thalassemiamutationIVSI-5(G-C) o Forward primer: 5’-ACCTCACCCTGTGGAGCCAC o Reverse primer(ARMS) 5’-CTCCTTAAACCTGTCTTGTAACCTTGTTAG o Amplified product 285 bp o Control primer(forward): 5'-CAATGTATCATGCCTCTTTGCACC o Control primer(reverse): 5'-GAGTCAAGGCTGAGAGATGCAGGA o Amplified product: 861 bp Reaction volume: 25μl PCR mix: 22μl Primerconcentration: 1μl (5 p mol each/ μl) (Chapter3) Taq polymerase: 0.5 units(0.1μl) Template DNA: 2μl (~300ng) Thermal cycling: o Initial denaturation: 1 minute at94o C o No. of cycles: 25 o Denaturation: 1 minute at94o C o Annealing: 1 minute at65o C o Extension: 1 minute 30 secondsat 72o C o Final extension: 3 minutesat72o C Electrophoresis: 10 X 10 cm 6% polyacrylamide,40minutesat150 volts. Staining: 0.1% silvernitrate Result: below
- 7. Silver-stainedpolyacrylamidegel electrophoresisafterARMSPCR.All lanesshow 861bp internal control fragment.Lanes1-3 shows285bp fragmentof IVS1-5 mutation.Lanes4-5 are negative forthe same mutations.Lane 7 showsallelicladderforvariousthalassaemiamutations. PROCEDURE The process of the ARMS PCR issimple andvery effective. Noharmful radiolabellingisinvolved inthe ARMS allele-specificPCR. The procedure of ARMS PCR isdivided into4steps: 1. Primerdesigning 2. Amplification 3. Agarose gel electrophoresis 4. Resultsinterpretations PRIMER DESIGNING FOR ARMS PCR: The primermust be allele-specific. Suppose ourDNA sequence hasG-A pointmutation viz, Gin normal allele andA inplace of G inthe mutantallele. We have to designaforwardprimerinsuch a mannerthat for normal allele the primercontains C (complementary toG) at 3’ endand the mutantprimercontains T in place of C. The magic of the presenttechniquehashappened when we addamismatchbase nearto our SNP at the 3’ end. The mismatchis the keyfactor inachievingthe amplification. If the mismatchisweakthe chance of amplification ishigher. Addstrongmismatchnearthe 3’ endof the primer(at -2 position ideally) hence, inthe non-complementary alleleitcannotbindtothe DNA sequences andtherefore terminates amplification. C: T, G: A andA: G are the strongmismatchbase pairswhichreduce the amplification process upto 100-fold.The reverse primeroranotherprimer(which isnotmodified) generally remains the same. The primerlengthof the allele-specificprimermustbe between 20 to 30 nucleotides.
- 8. Once our primerisready, we have to modify the amplification conditions forachievinghigher amplification. AMPLIFICATION CONDITIONS: The annealingtemperatureshould higher(donotcompromise the annealingtemperature). The concentration of the PCR components isgivenintothe table below, The PCR reaction preparation recipe. Importantly,the PCRcyclesforARMS-PCRare lowerthanthe normal PCR reaction.Seta PCRcycle between22to 25 but not 35. Asthe PCR cyclesincrease the chance of false-positive resultsincreases. “An additional mismatchtothe 3’ endafterthe firstmismatchincreasesthe specificityof the ARMS-PCRreaction.” In the ARMS PCR,the internal control playsa crucial role,itfacilitatesadditionalaccuracyinthe allele-specificPCRbyreducingthe chance of the false-positive results. AGAROSE GEL ELECTROPHORESIS: The beauty of the ARMS PCR isthat it does notrequire anyhybridization steps. Whichwill make this technique more suitable andmore reliable forthe diagnosis. The amplified fragmentsare directly loadedonthe 2% agarose gel for gettingthe results. The samples are loaded sequentially onthe agarose gel andrun for 45 minutes.
- 9. RESULTS AND INTERPRETATION: First, observe the internal control. If the internal controls bandsare presentinall the wellsindicates that our reactioniscompletelyfine, nofalse-positive resultsare present. Now analyseeachbandin normal as well asinmutant allele. The graphical representation of the agarose gel electrophoresis resultsisshown inthe figure below.Suppose we have three samplesone normal, one heterozygous carrier andone homozygous dominant. We have prepared 2tubes foreach sample, atotal of 6 tubes withone positive control andtwonegative control.
- 10. As shownintothe figure above,anonlysingle bandisobservedinthe homozygousnormal sample, twobands (one fornormal and one formutant) are observedinthe heterozygouscarriersample anda single bandof a mutantallele isobservedinthe homozygousdiseasecondition.The wells7,8 and 9 are the controls. IMPORTANCE OF ARMS-PCR IN THE DIAGNOSIS OF GENETIC DISEASE: The ARMS PCRis one of the importanttoolsinthe geneticdisease diagnosis inrecentdays. The restriction digestion methodisnot100% accurate and also, not all the sequences have the restriction site. Therefore,restrictiondigestionisnotapplicableinall typesof mutationorpolymorphism. The allele-specificPCRisan accurate methodforsingle-gene disordershavingthe SNPs.Itisalsoa gold standardmethodforSickle cell anaemiaandthalassemialikeinherited disorder. Also,itisapplicable inmutationdetectionof JAK2and HIV.Commercial kitsare available forthe differentrange of disorderssuchasHbS and JAK2gene SNPsdetectionkits. ADVANTAGE OF ARMS PCR: The technique isexclusively forthe SNP genotyping. Further, homozygous andheterozygous canbe detected bythistechnique It iswidely applicable inthe single gene pointmutationdetection suchassickle cell anaemia and thalassemia. It isfast, reliable accurate andrapid. LIMITATION OF ARMS PCR: Deletion/othermajorduplication andchromosomal abnormalitiescannotbe detected. Only knownSNPsare detected byARMS-PCR. Internal control isrequired because of the chance of the false-negative results. It istemperature-sensitive. A minute fluctuationinthe temperatureleadstofalse-positive results. Thousandsof SNPscannot be detectedinasingle assay.
- 11. CONCLUSION The amplification-refractorymutationsystem(ARMS) isasimple methodfordetectinganymutation involvingsinglebase changesorsmall deletions.ARMSisbasedonthe use of sequence-specificPCR primersthatallowamplificationof testDNA onlywhenthe targetallele iscontainedwithinthe sample.Followingan ARMS reaction, the presence orabsence of aPCR product isdiagnosticforthe presence orabsence of the targetallele. The amplification-refractorymutationsystem(ARMS) isasimple methodfordetectinganymutation involvingsinglebase change. The ARMS-PCRmethodispopularfor the detection of known SNPs;however, itisa very tedious and time-consumingprocess toencountermore SNPsatonce. Because itis allele-specific, the accuracy of ARMS-PCRishigher. ARMS-PCR can be utilizedasacost-effective,rapidandreproduciblemethodforSNPgenotyping especiallywhile performinglarge-scale epidemiological/associationatstudies. Some of the deletions canalsobe screened usingARMS-PCRmethod. REFERENCES Dr. Tushar Chauhan. 27Feb.2019, GeneticEducation.co.in Tushar Chauhan, Tushar Kachhadiya.19Aug.2018, Genetic.co.in Dr. Satyendra Kumar.30Dec.2013,Technology Health & Medicine Baoyu Peng. July 2017, reseachgate.net
