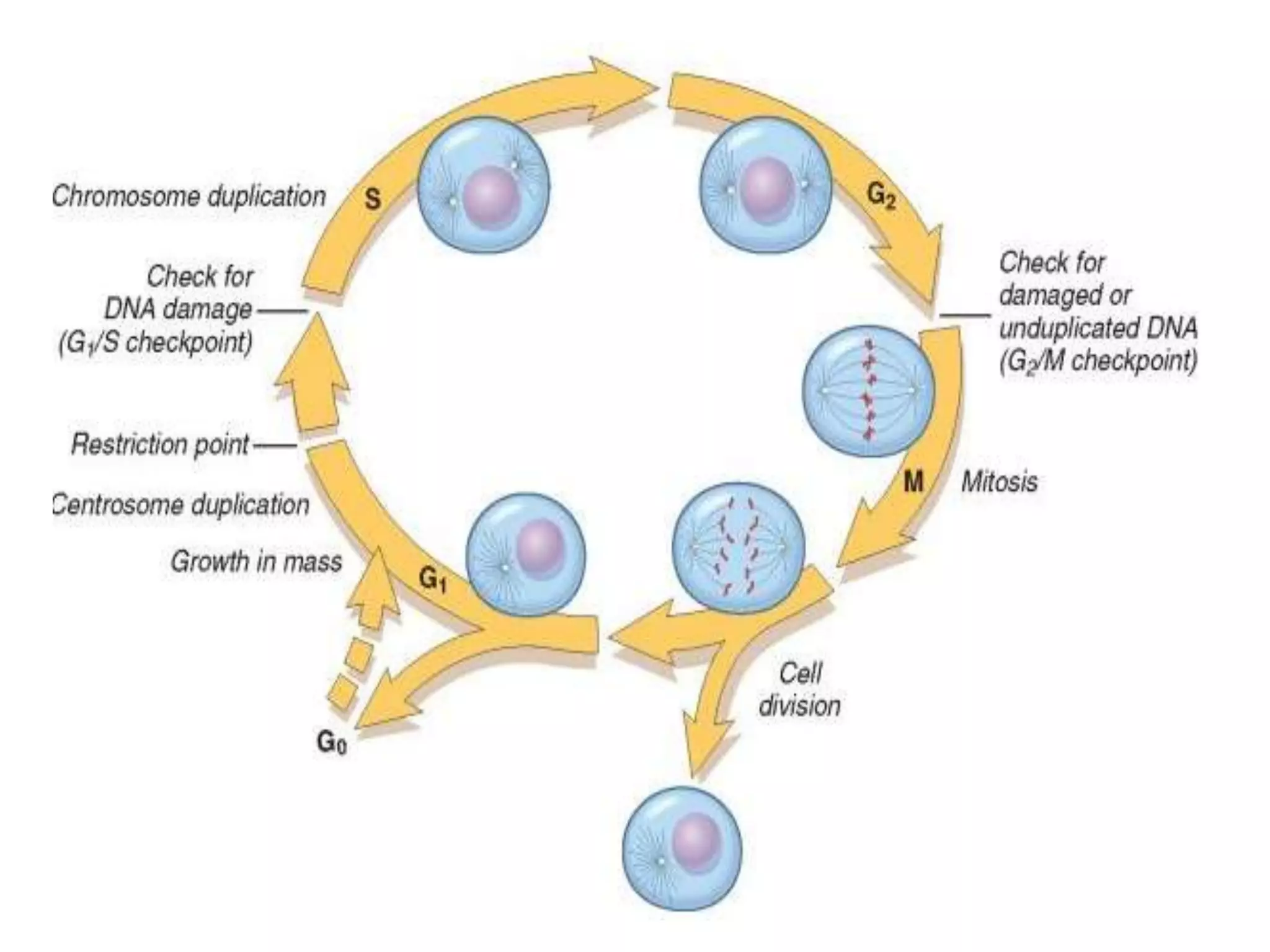
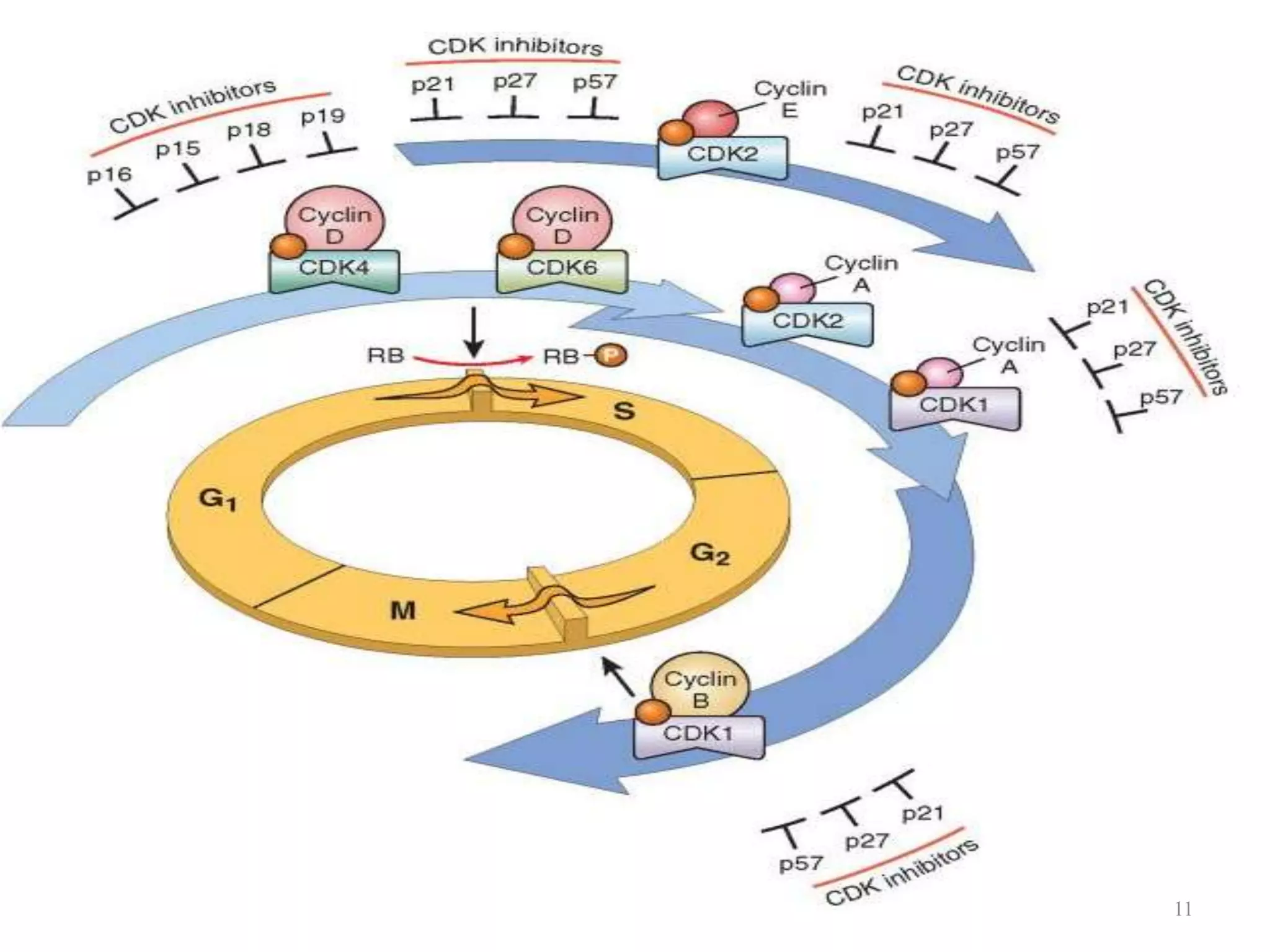
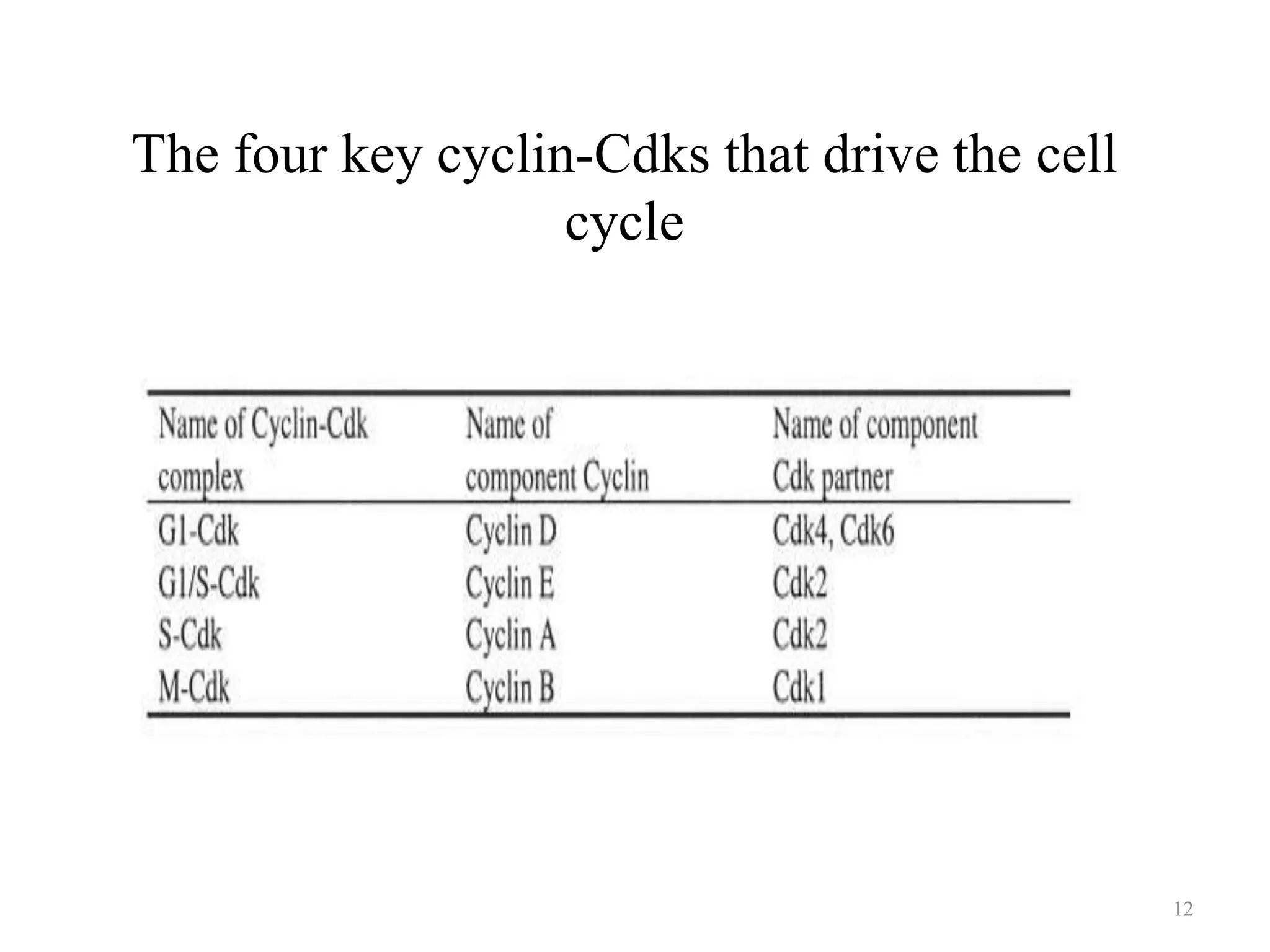
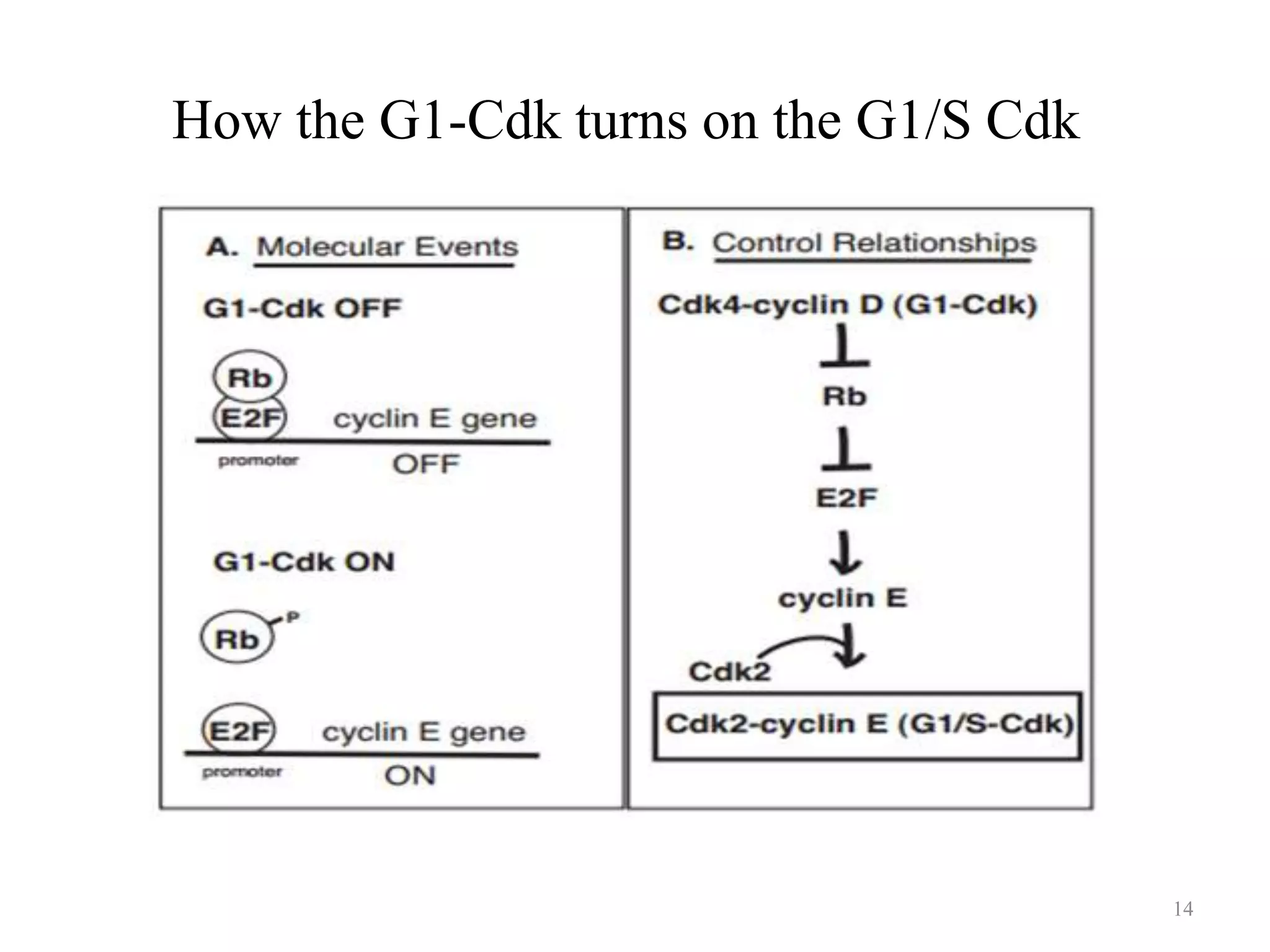
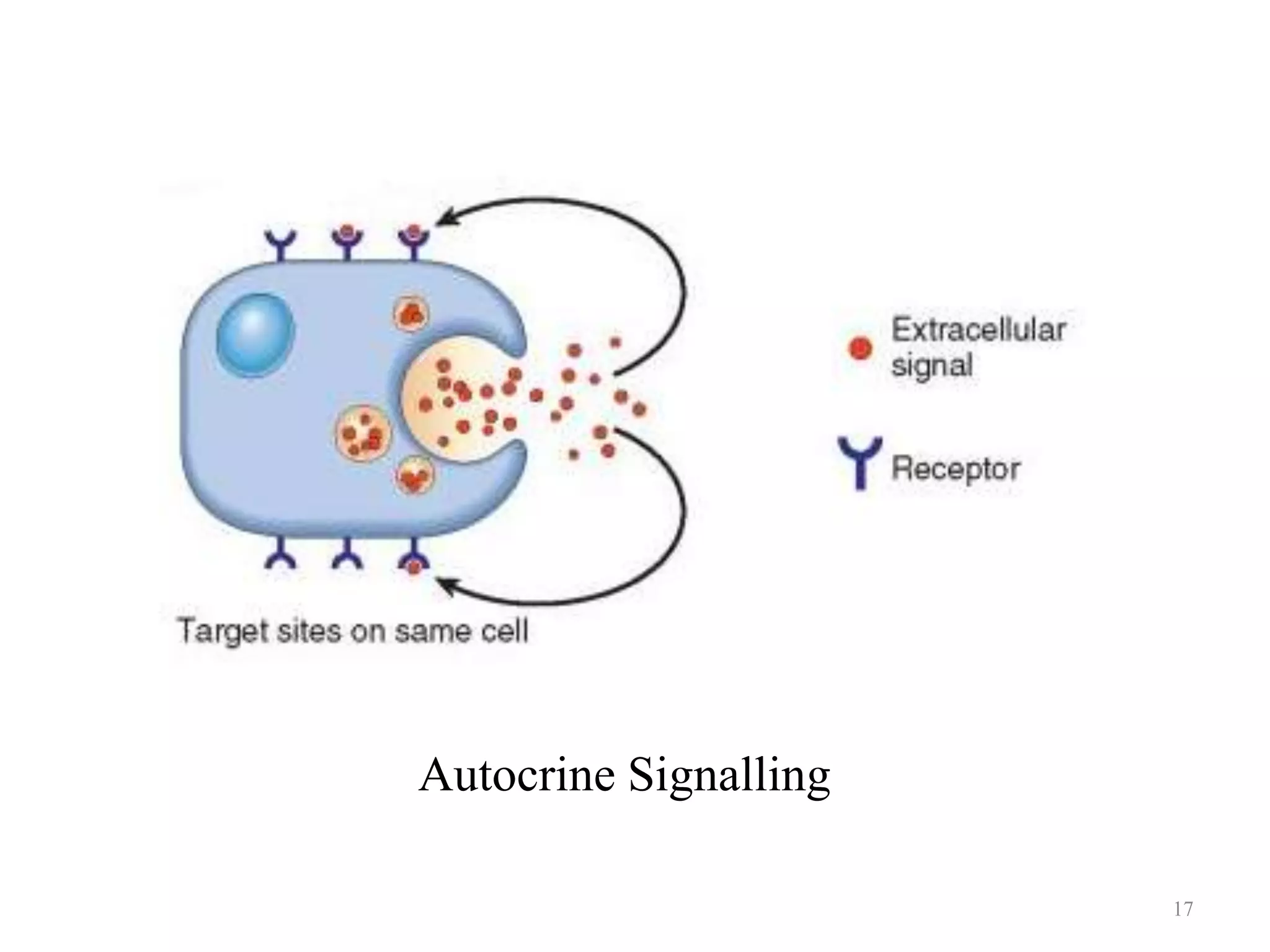
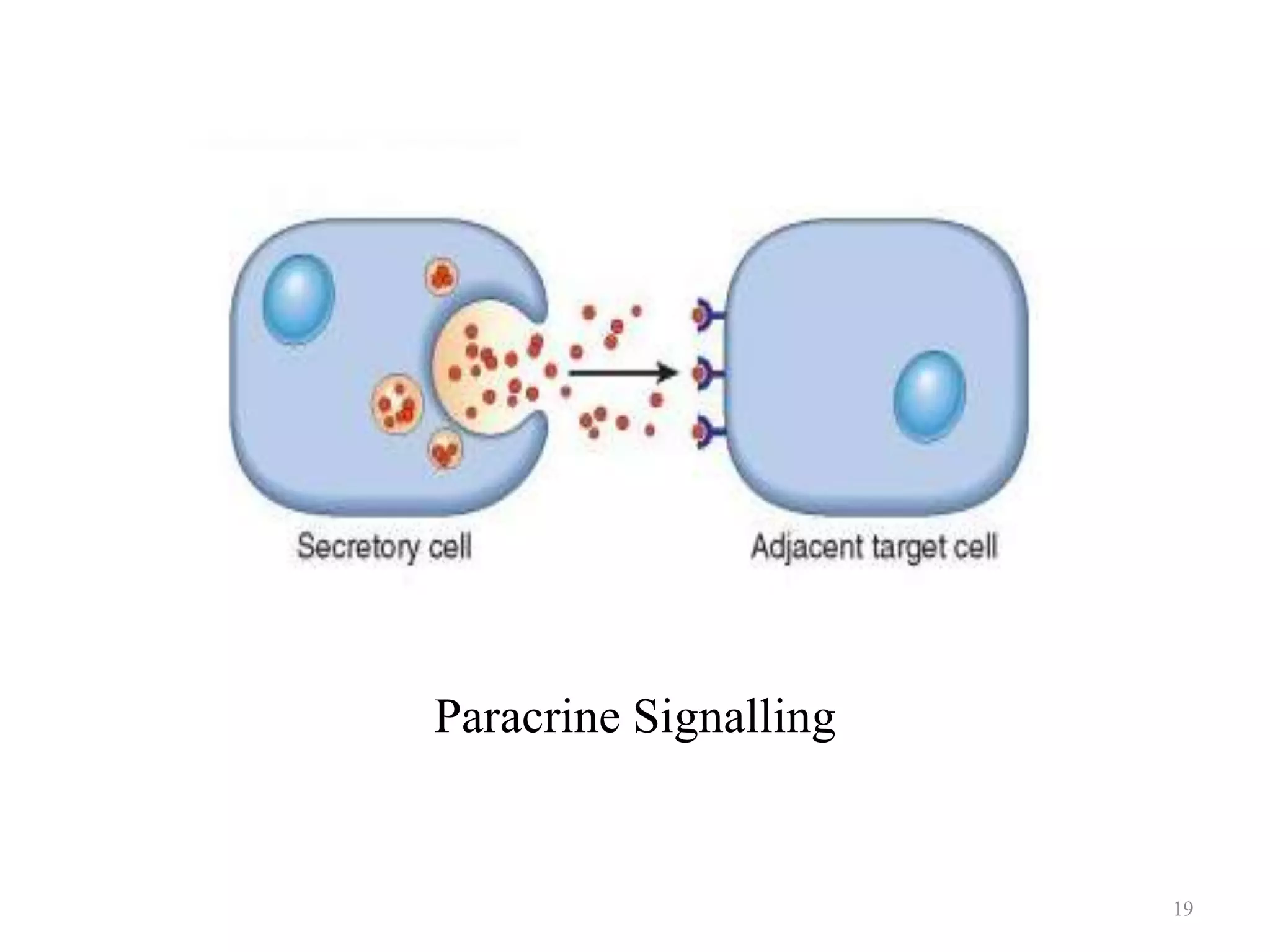
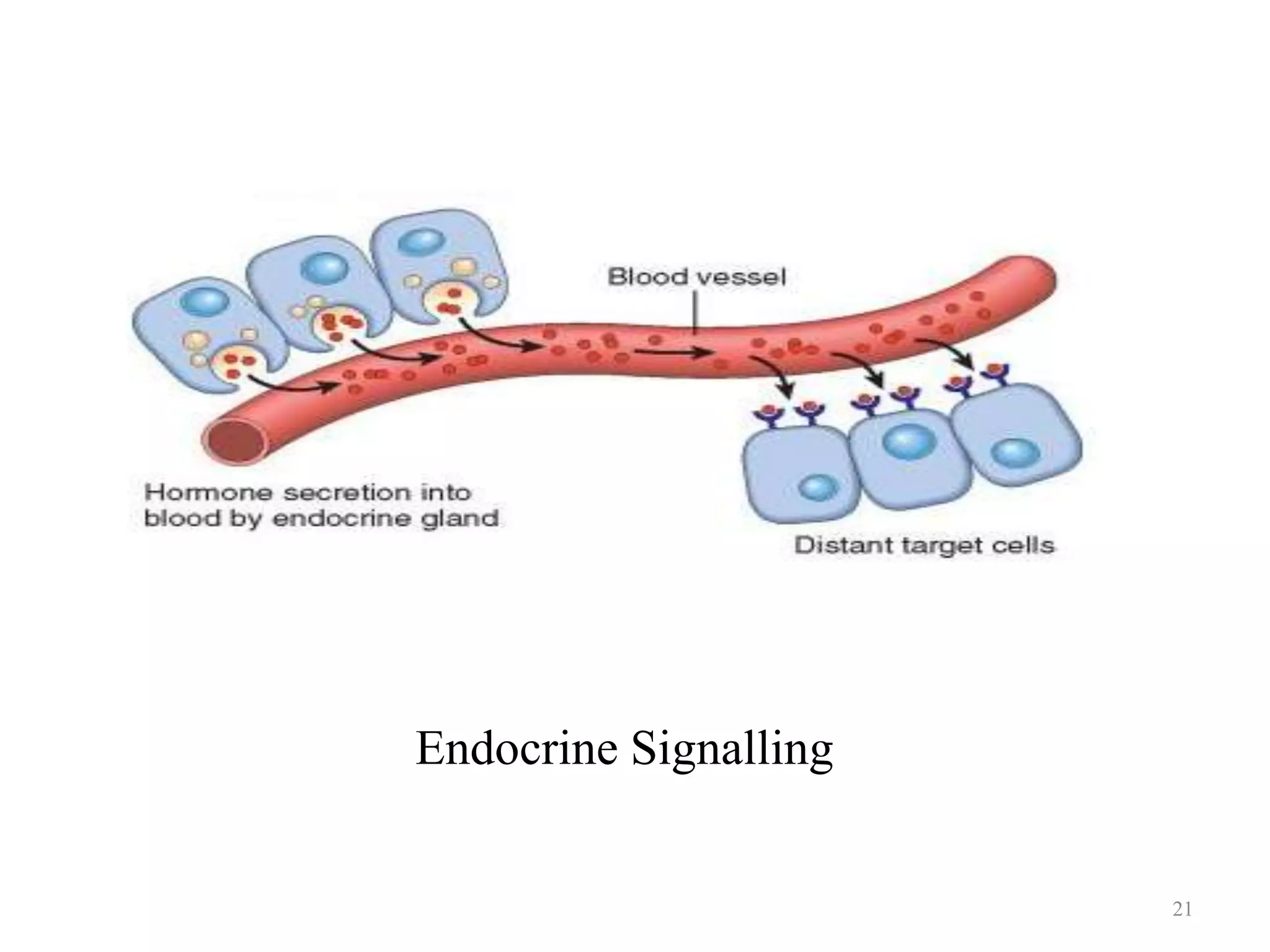
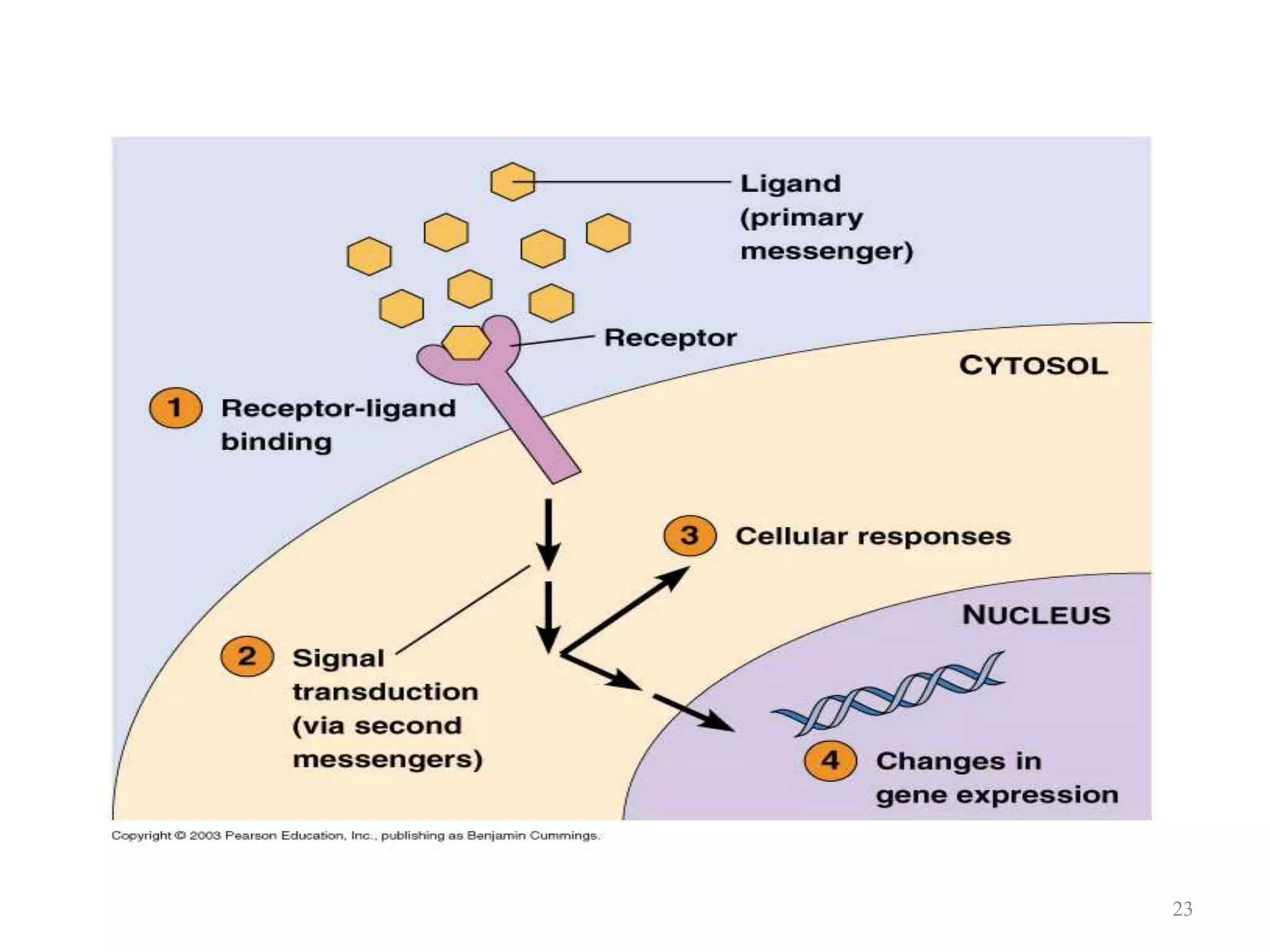
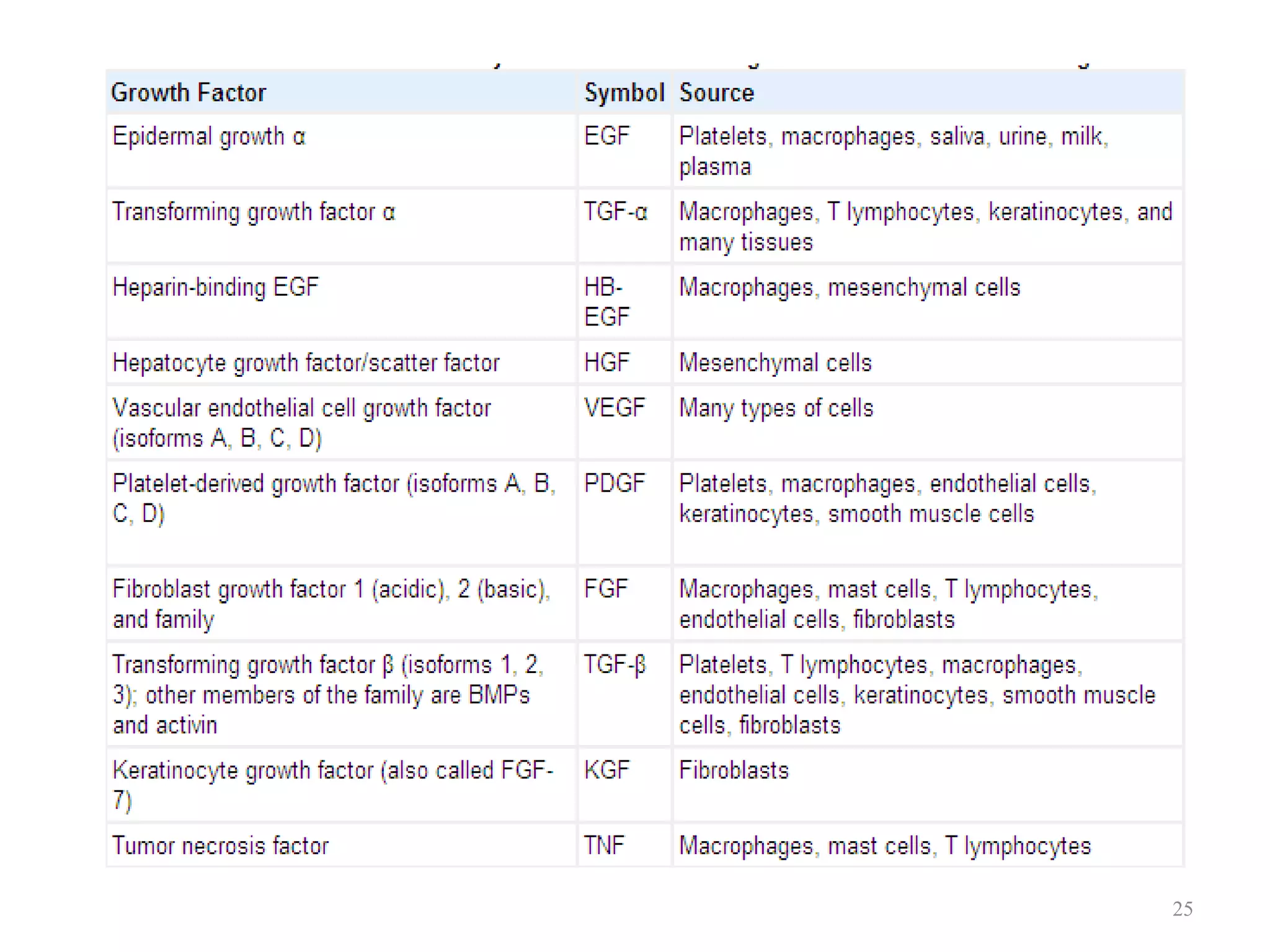
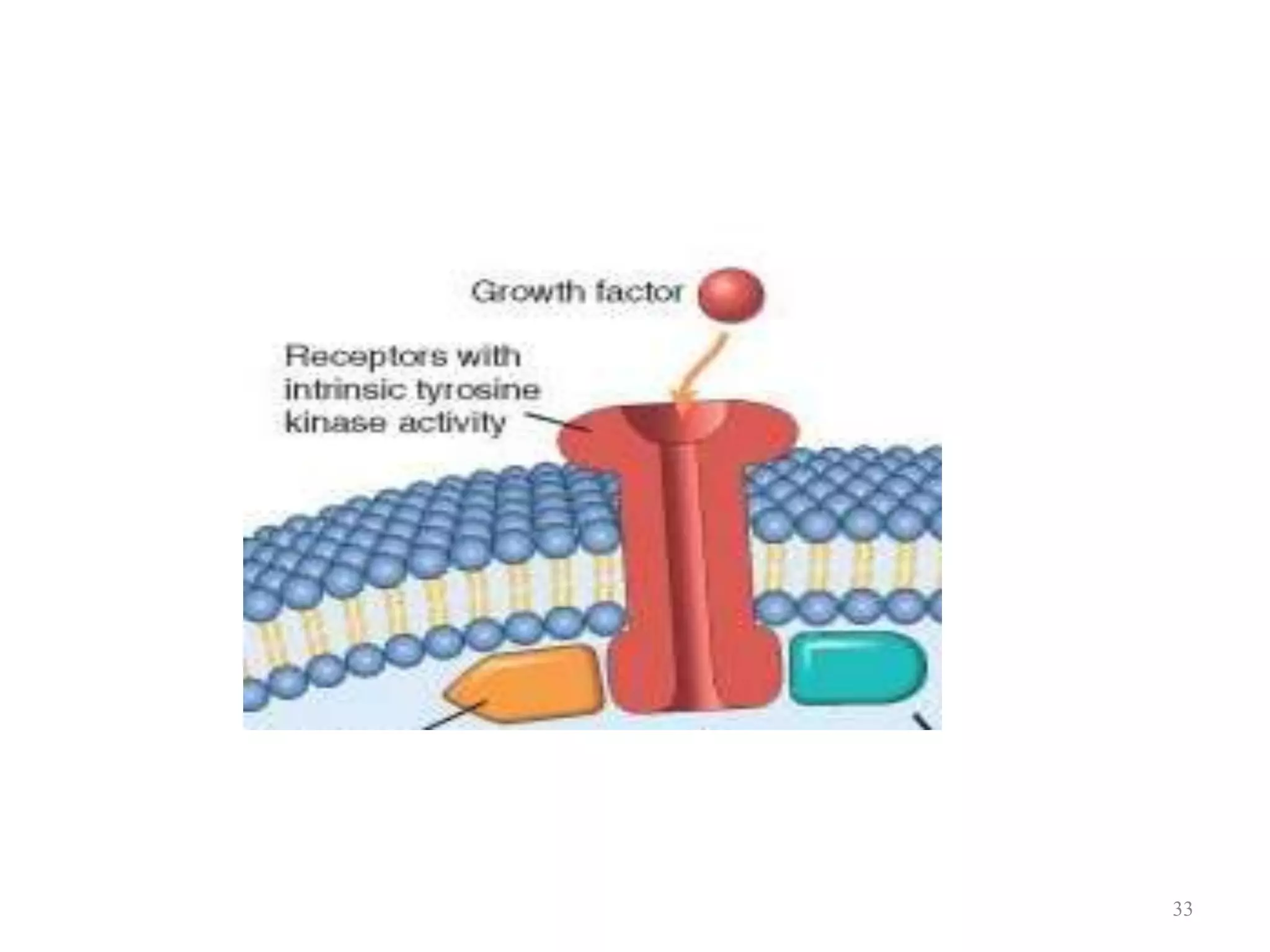
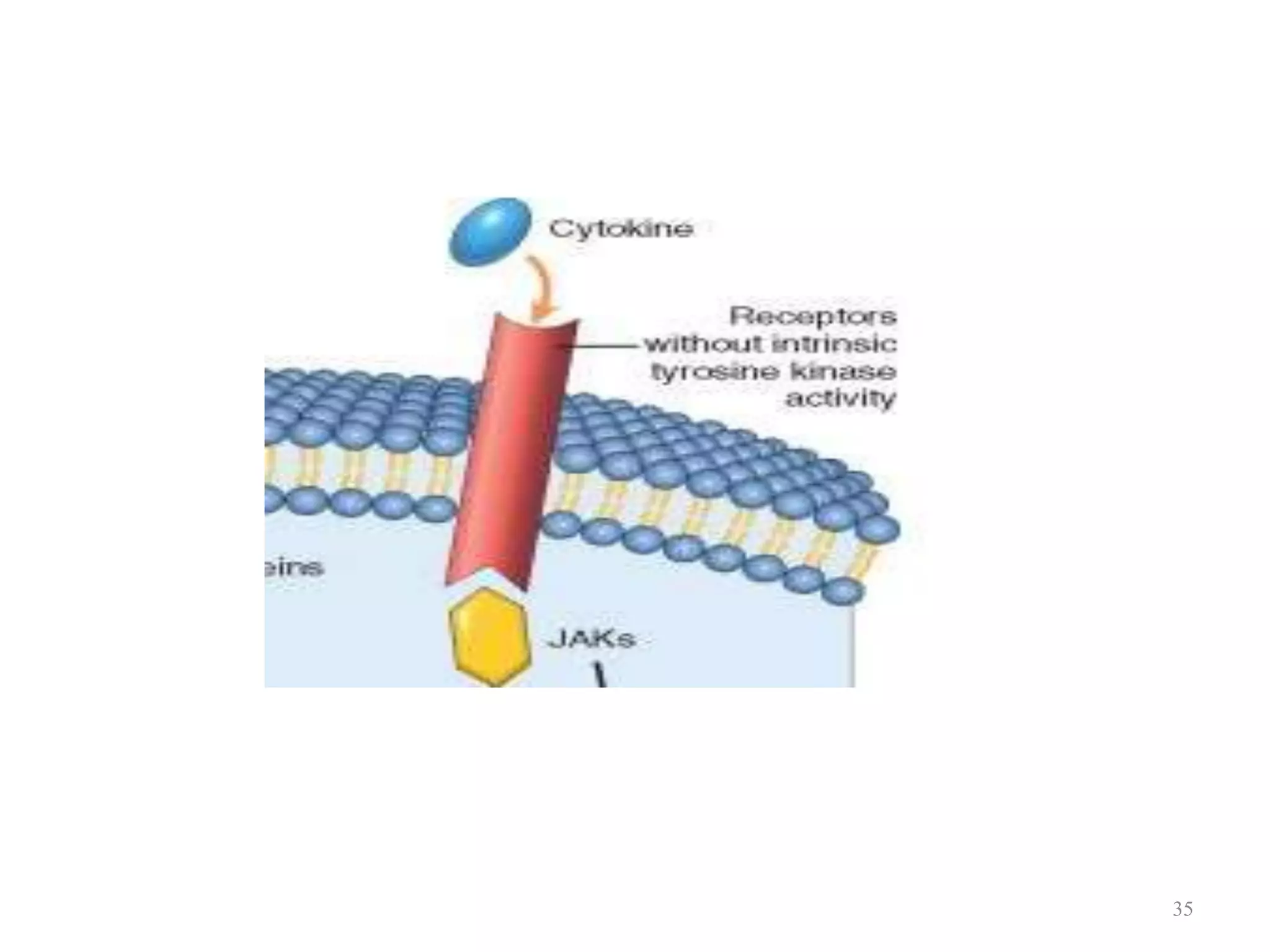
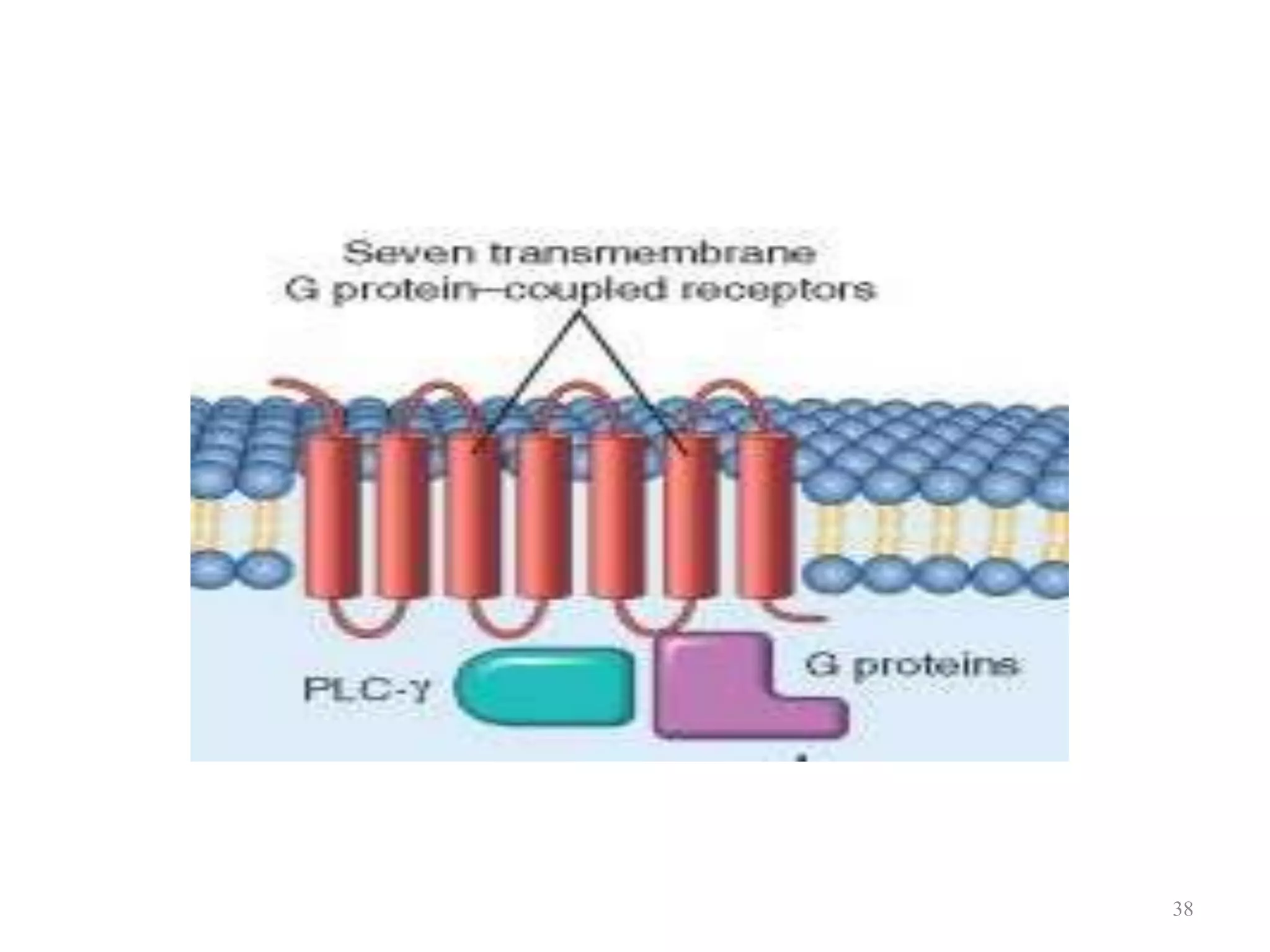
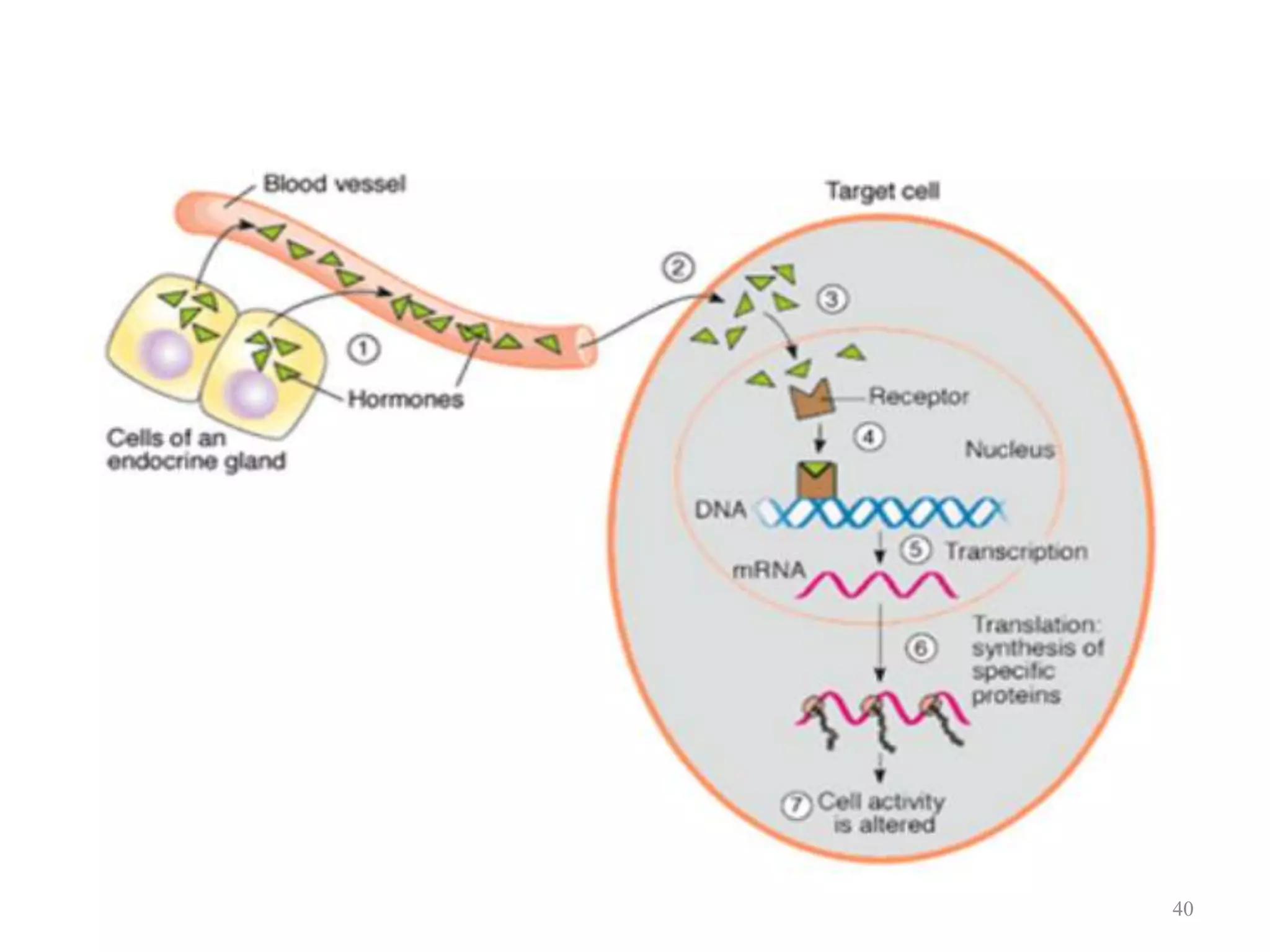
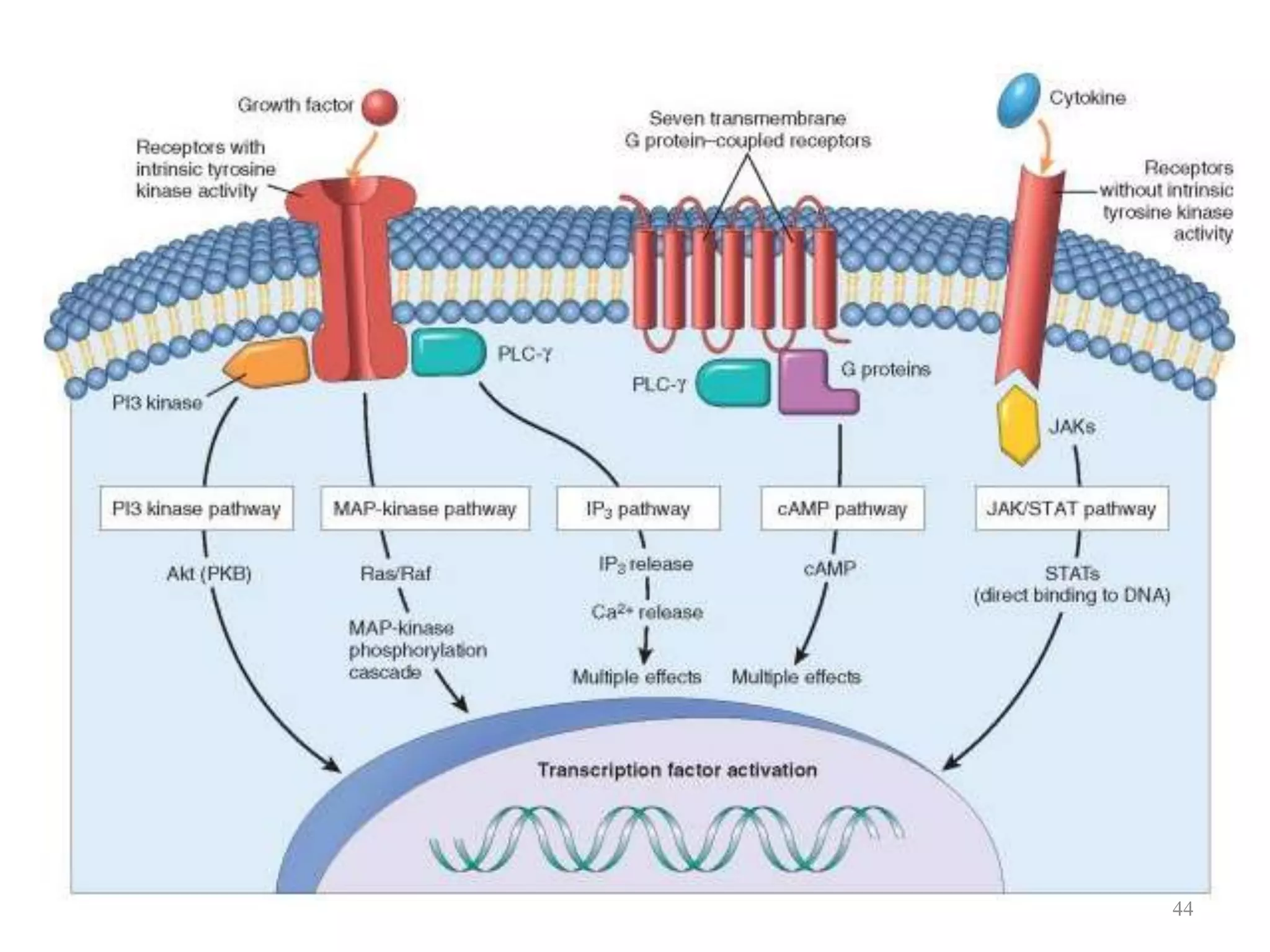
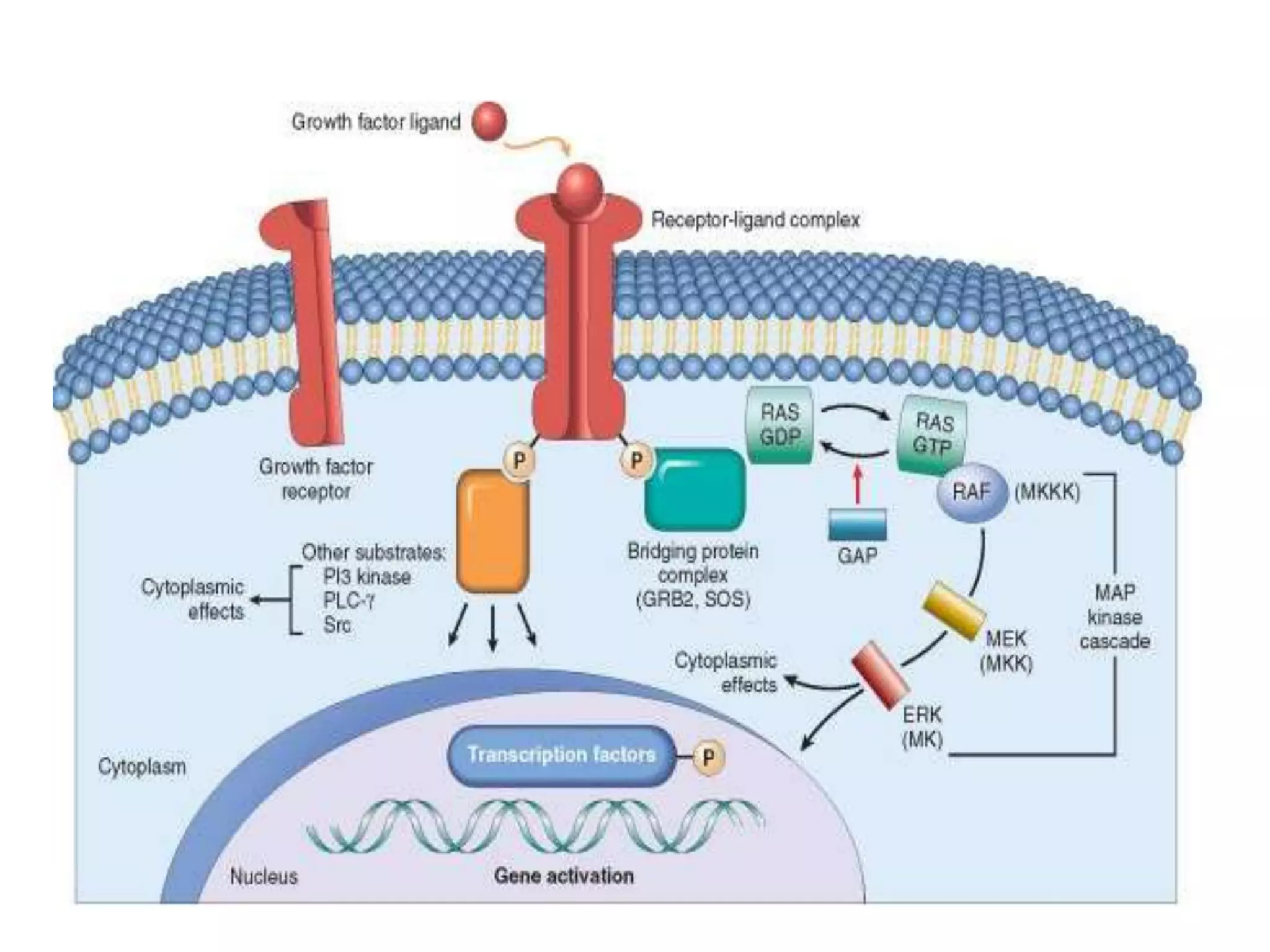
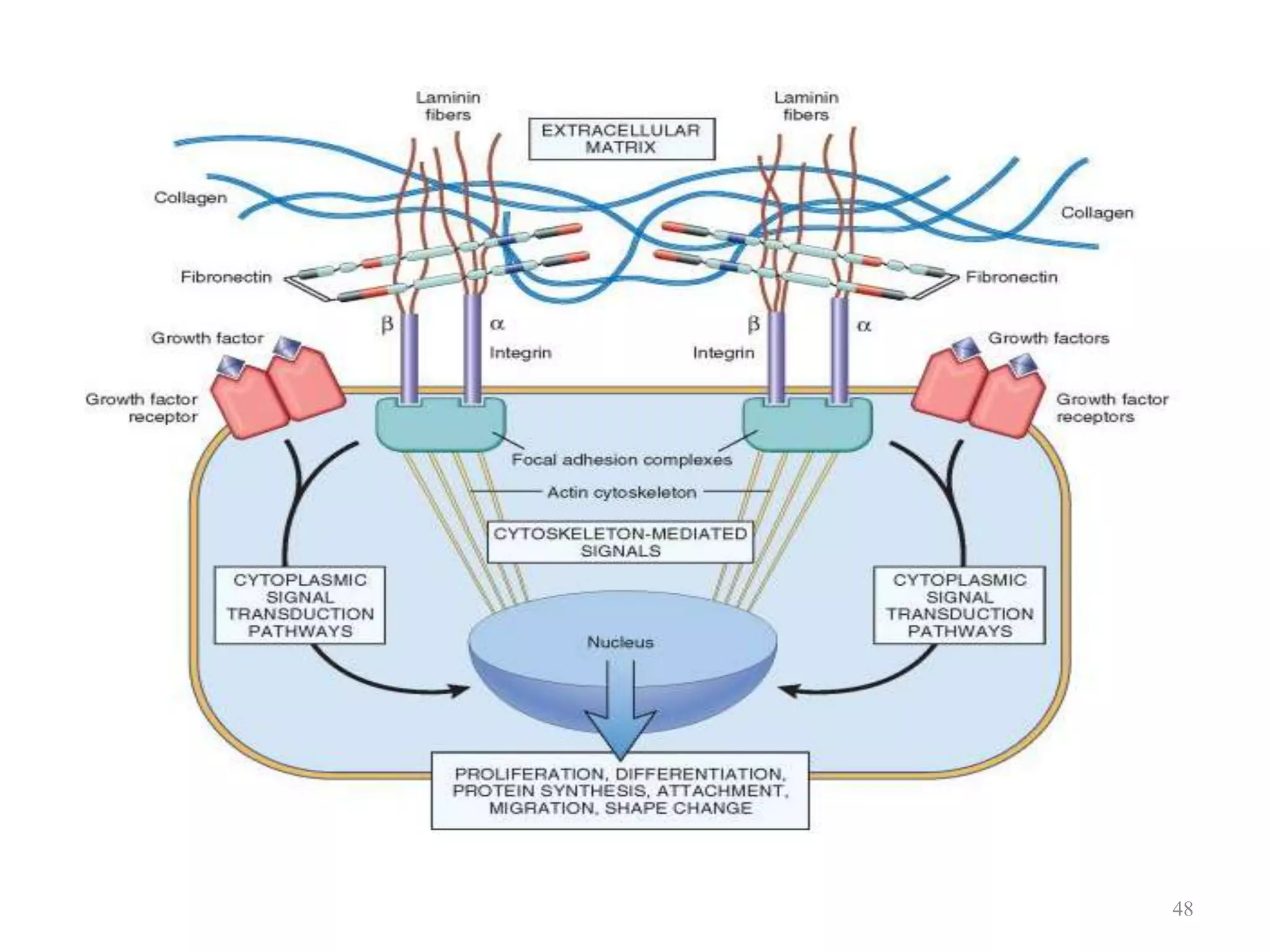
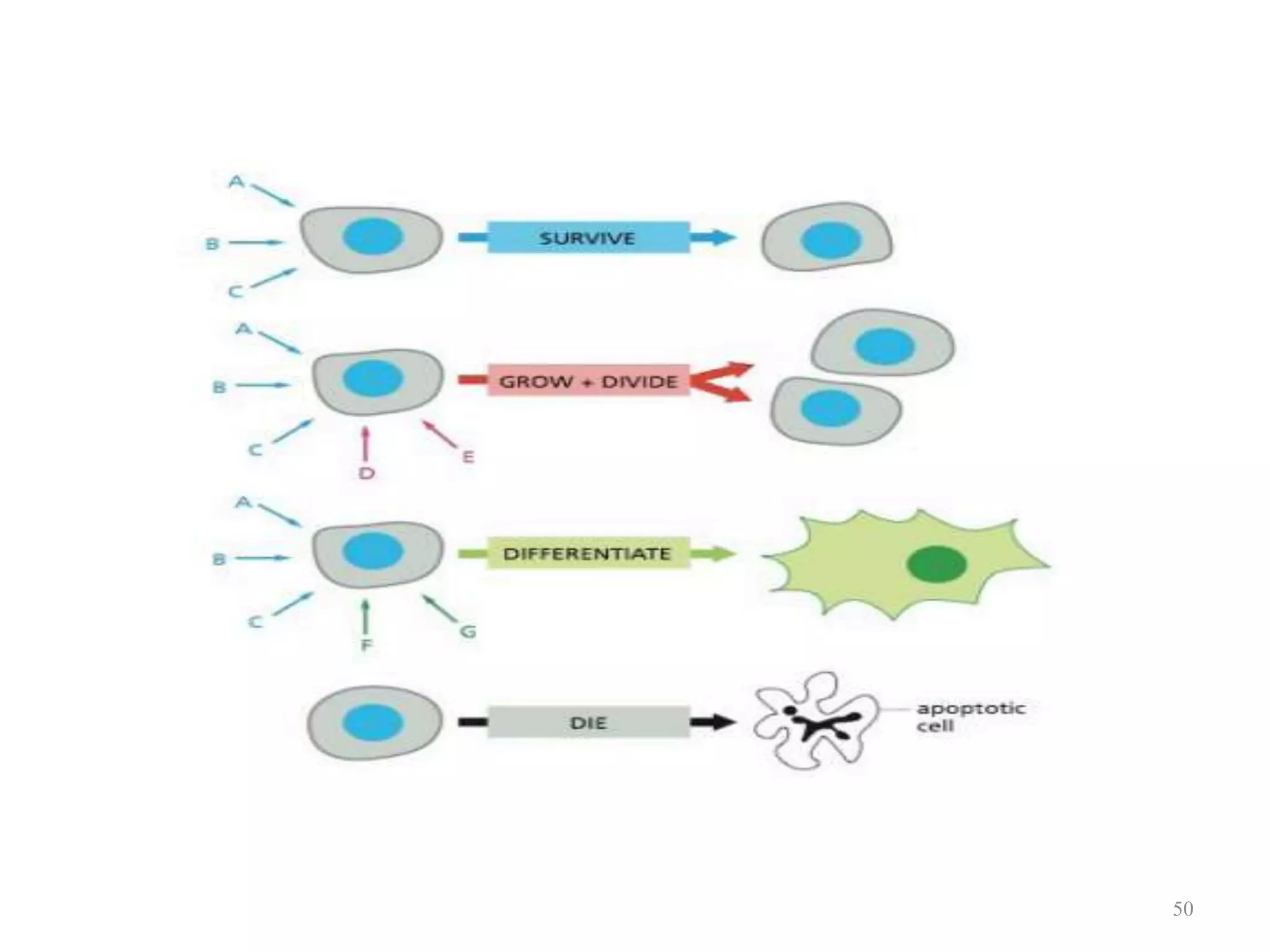
Cell growth can refer to the increase in cell size and organelles during interphase (G1 and G2 phases) or the growth of a cell population through cell division (M phase). The cell cycle consists of G1, S, G2, and M phases, separated by gap phases G1 and G2. Growth factors such as EGF, TGF-α, HGF, and PDGF act as ligands that bind to cell surface receptors and transmit signals to regulate the cell cycle. These signals activate intracellular pathways using second messengers to induce transcription factors that drive expression of genes controlling cell cycle progression and cell growth.