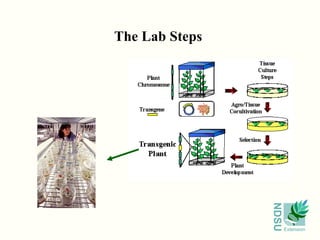
This document provides an overview of biotechnology principles and applications. It defines biotechnology as the application of technology to modify biological organisms by adding genes from other organisms. The document discusses how genes are identified, isolated, and manipulated to introduce desired traits. It describes techniques such as homology cloning, complementary genetics, and map-based cloning used to isolate genes. The document explains how genes are introduced into plants using transformation methods like Agrobacterium and biolistics. It provides examples of transgenic crops and their applications in agriculture.





















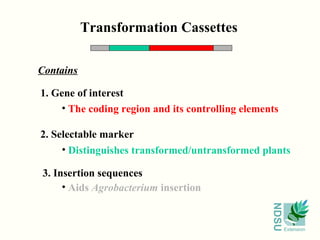

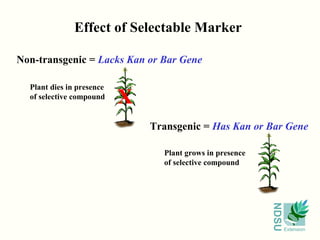
![Selectable Marker
Promoter Coding Region
Promoter Region
• Normally constitutive
ex.: CaMV35s (Cauliflower Mosaic Virus 35S RNA promoter
Coding Region
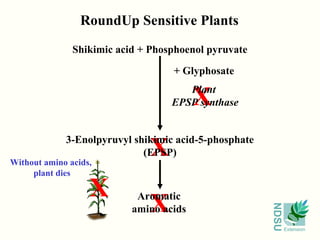
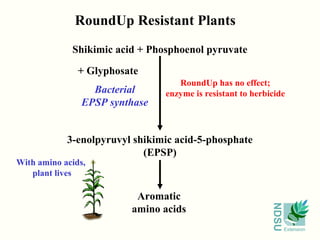
• Gene that breaks down a toxic compound;
non-transgenic plants die
ex.: nptII [kanamycin (bacterial antibiotic) resistance]
aphIV [hygromycin (bacterial antibiotic) resistance]
Bar [glufosinate (herbicide) resistance]
NDSU
Extension](https://image.slidesharecdn.com/techniques-of-biotechnology-mcclean-good-121130185332-phpapp02/85/Techniques-of-biotechnology-mcclean-good-36-320.jpg)