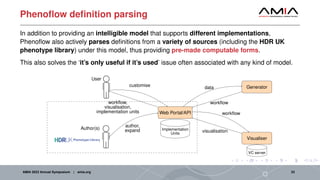
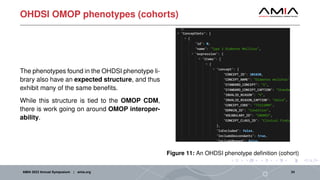
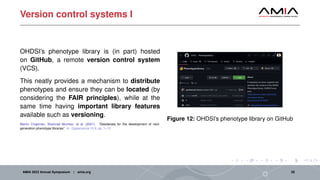
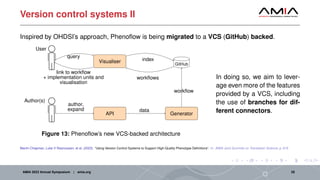
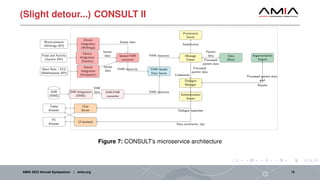
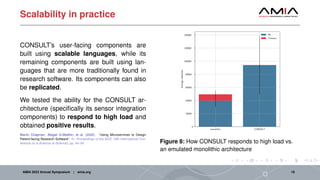
The document discusses different architectural considerations for building phenotype libraries that are accessible at a large scale. It covers software architecture, definition structure, and distribution mechanisms. For software architecture, it advocates for a microservices approach to allow components to be built with different technologies and scaled independently. For definition structure, it presents Phenoflow's model that standardizes definitions and generates computable forms. For distribution, it notes the benefits of hosting definitions in version control systems like GitHub to improve discoverability. The overall goal is to make definitions easily located, downloaded, and interpreted by many users through careful library design.

















![Phenoflow’s definition model III
2 - icd10 A case is identified in the presence of
patients associated with the stated icd10
COVID-19 codes.
logic
step
Input Output
covid19_cohort Potential covid19
cases.
covid19_cases_icd10 covid19 cases, as
identified by icd10
coding.
csv
icd10.py python -
for row in csv_reader :
newRow = row . copy ( )
for c e l l in row :
i f [ value for value in
row [ c e l l ] . s p l i t ( " , " )
i f value in codes ] :
newRow[ " covid19 " ] = "CASE"
...
Computational
Implementation
Units
icd10.js javascript -
for ( row of csvData ) {
newRow = row . s l i c e ( ) ;
for ( c e l l of row ) {
i f ( c e l l . s p l i t ( " , " )
. f i l t e r ( code=>codes .
indexOf ( code) > −1). length ) {
newRow. push ( "CASE" ) ;
...
Abstract
Functional
Figure 10: Individual step of COVID-19 phenotype definition and implementation units.
AMIA 2023 Annual Symposium | amia.org 22](https://image.slidesharecdn.com/scalablearchitecturesforphenotypelibrariesbuildinginternationalphenotypelibraries-240125190338-9c0bafcd/85/Scalable-architectures-for-phenotype-libraries-24-320.jpg)