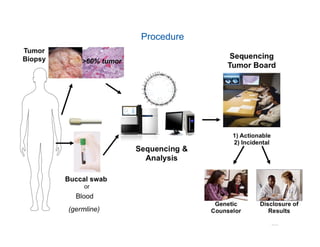
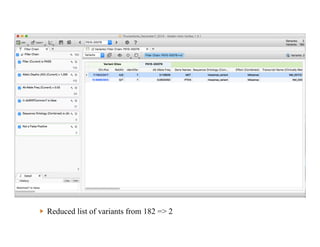
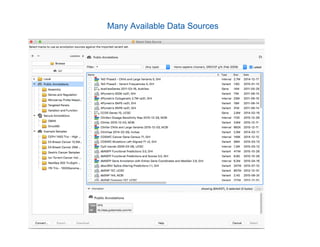
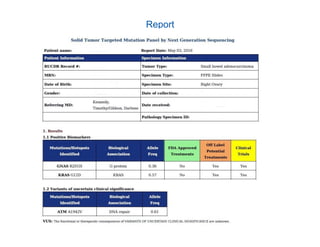
Jeffrey Rosenfeld described his work using tumor sequencing to personalize cancer treatment. His lab uses a targeted sequencing panel to identify driver mutations in tumors, then filters variants using software like VarSeq. Potentially actionable mutations are evaluated using databases like N-Of-One and discussed at molecular tumor boards to guide treatment. Integrating sequencing results into clinical decision making and research databases allows improving outcomes for patients.