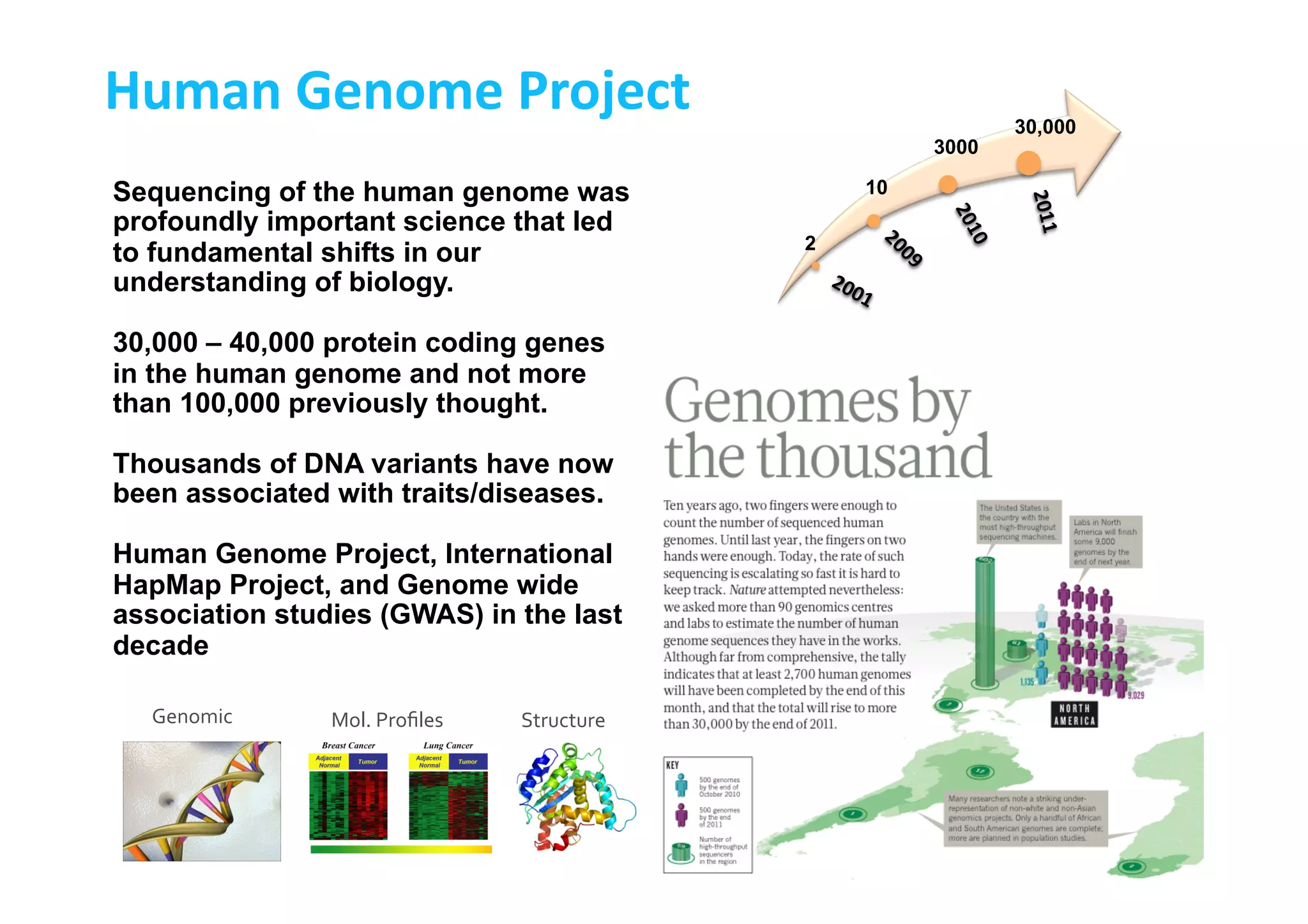
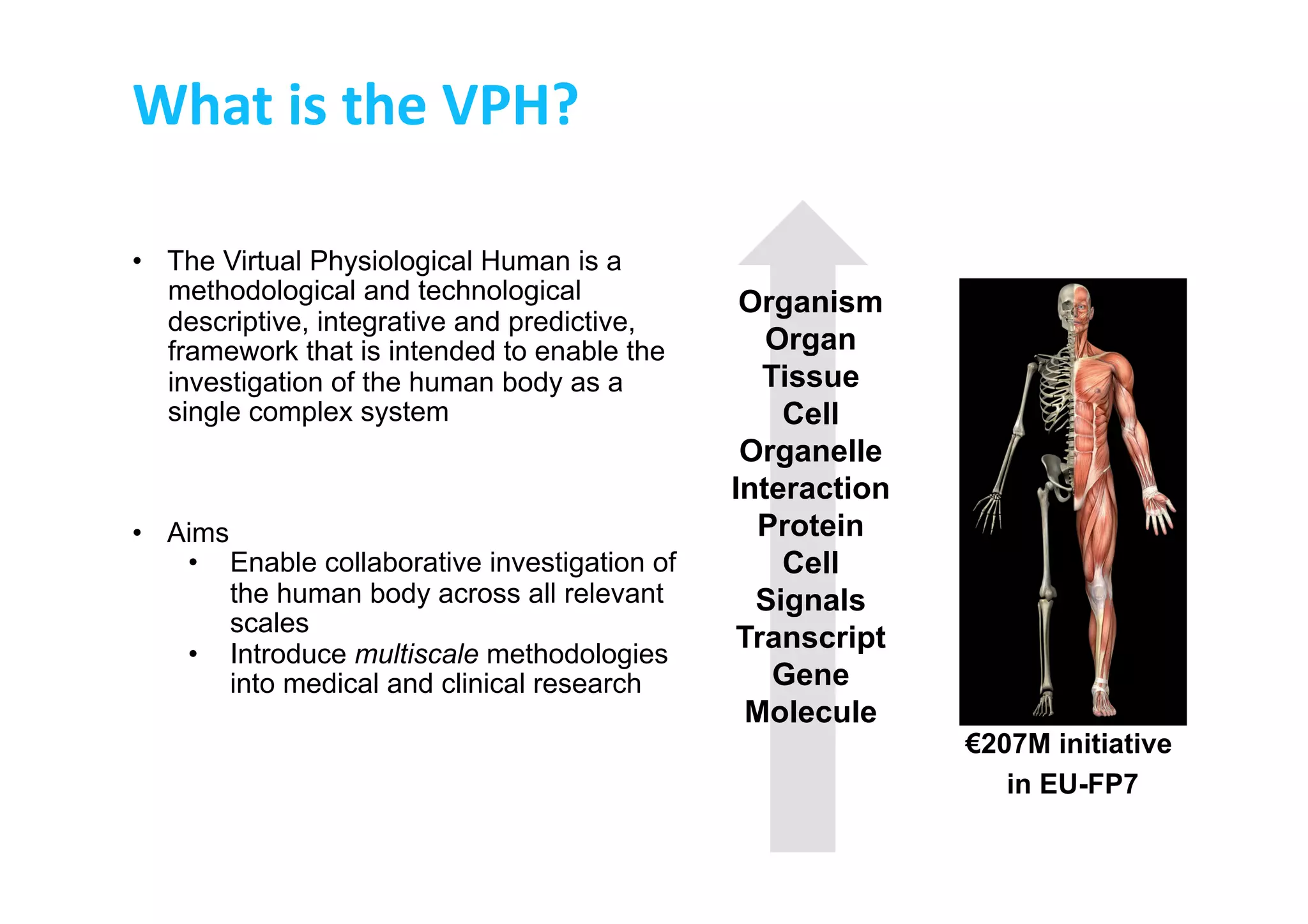
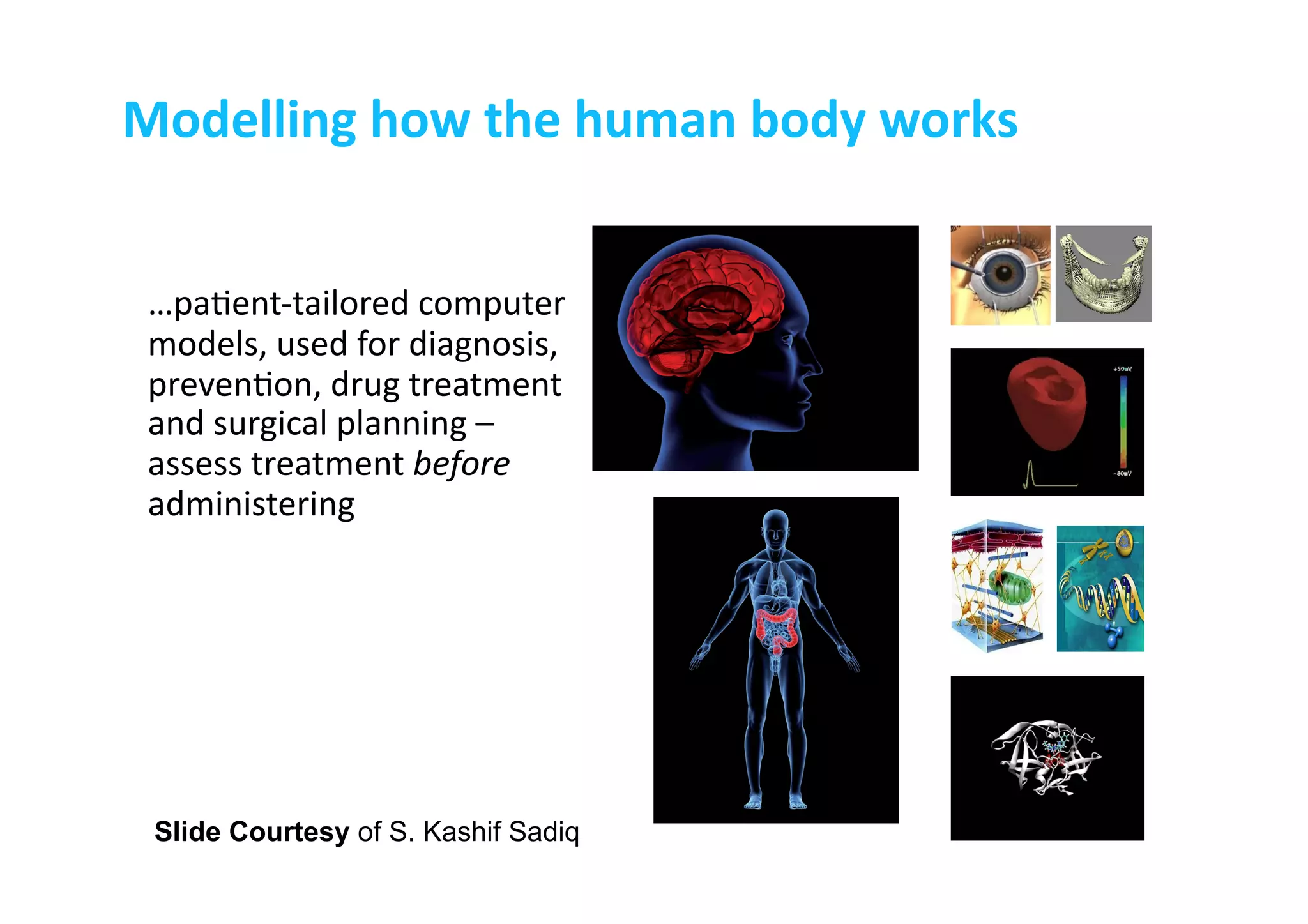
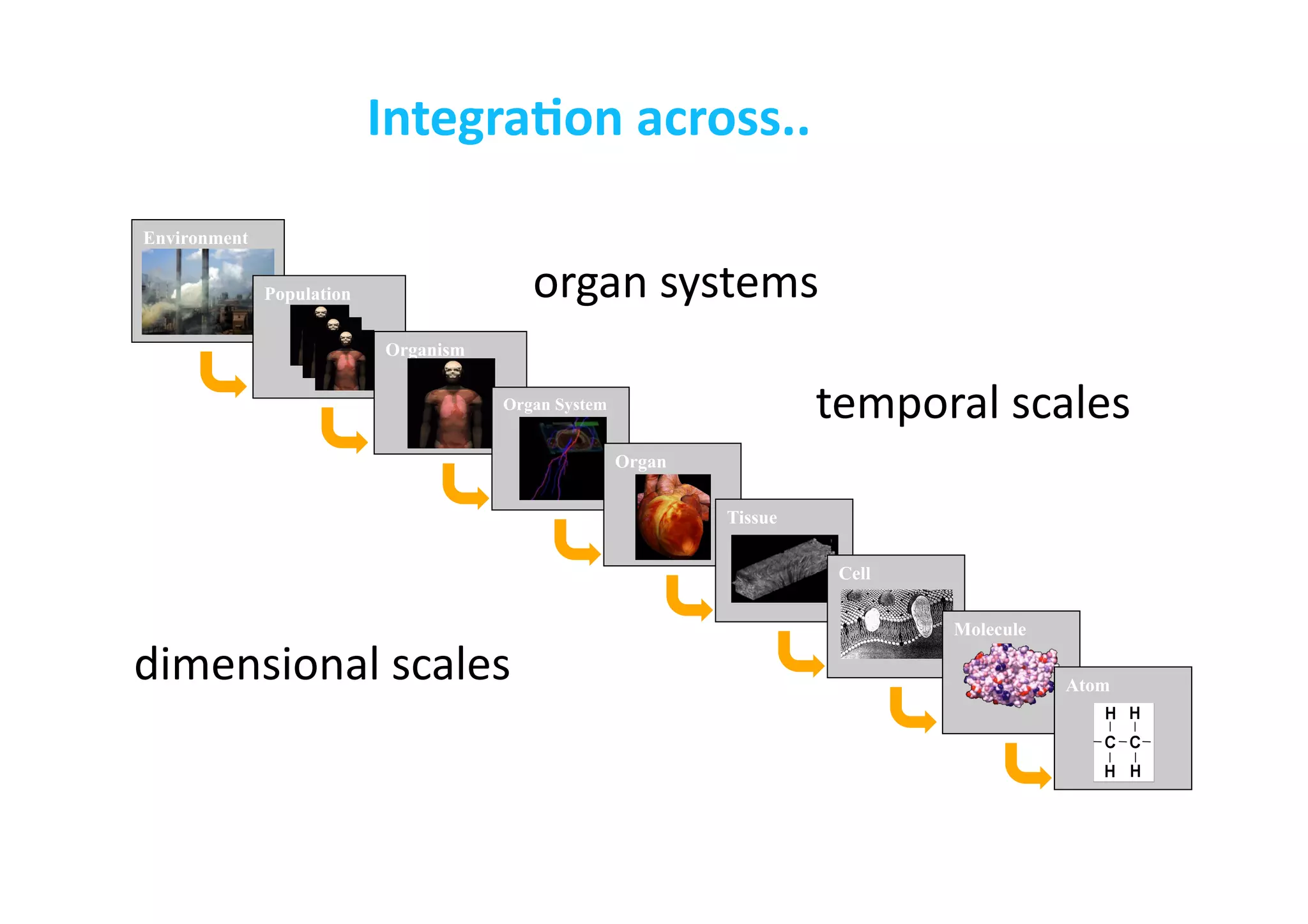
This document discusses the promises and challenges of personalized medicine. It begins by reviewing past initiatives like the Human Genome Project and the Virtual Physiological Human project. It then presents two case studies using physiological simulations for clinical decision support in surgery and for personalized drug design. The challenges of integrating data and models across multiple scales are discussed. The talk concludes that while significant progress has been made, fully realizing the promises of personalized medicine will require overcoming remaining technological and data integration challenges.