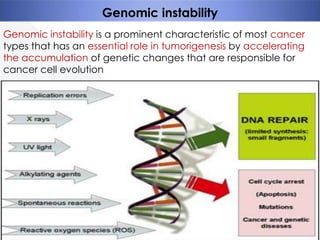
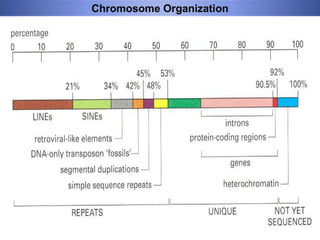
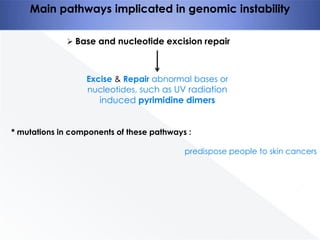
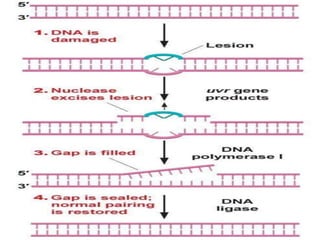
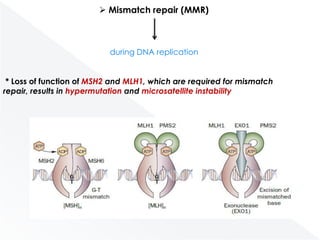
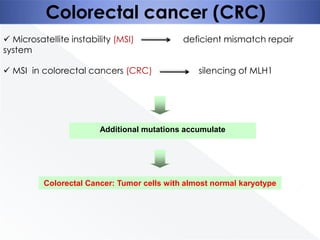
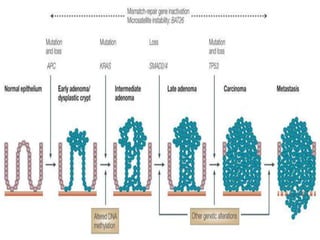
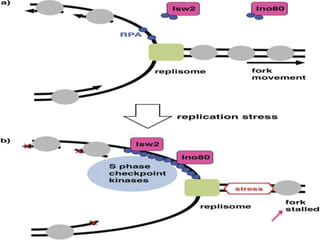
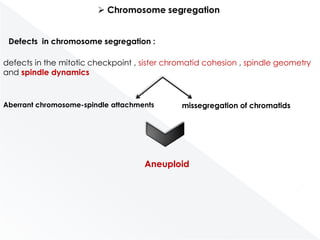
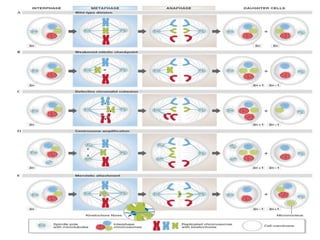
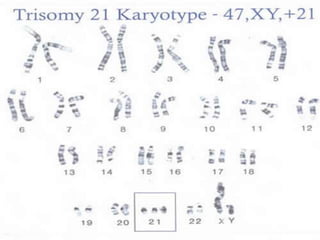
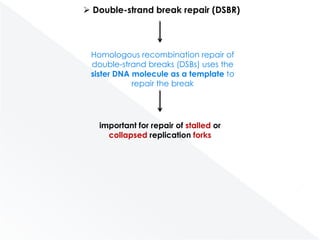
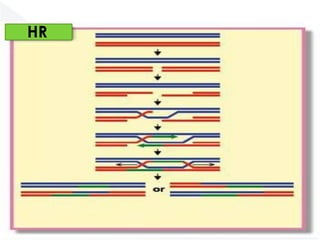
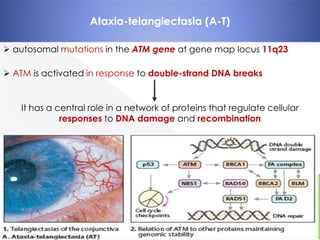
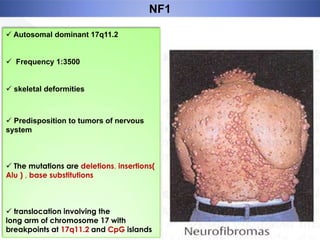
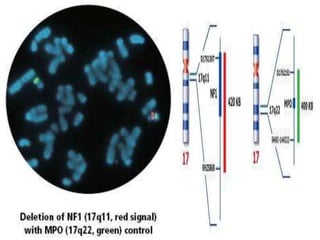
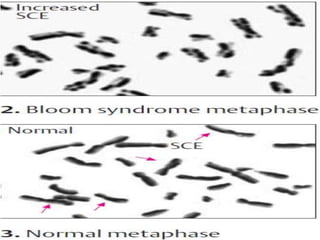
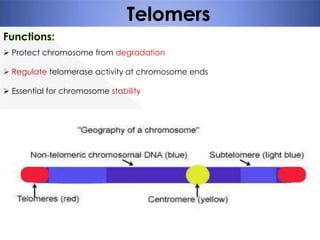
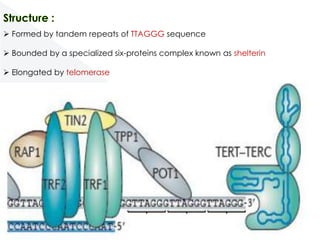
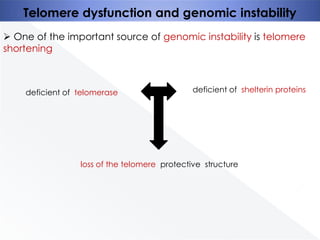
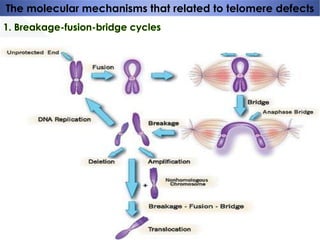
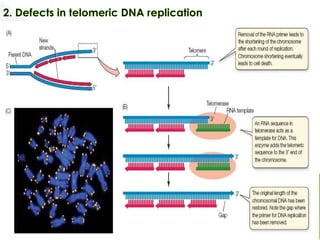
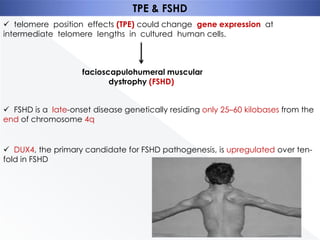
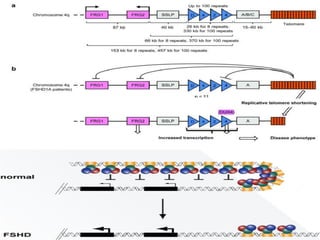
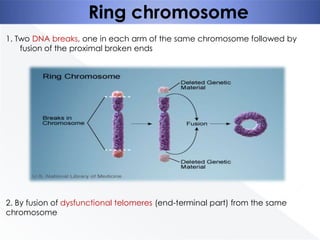
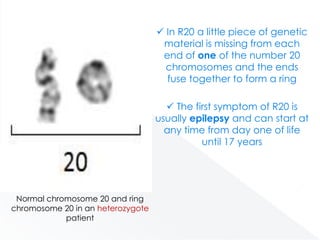
Genomic instability plays an important role in cancer development by accelerating the accumulation of genetic changes in cancer cells. Several mechanisms can cause genomic instability, including defects in DNA repair pathways like base excision repair, mismatch repair, and double-strand break repair. Loss of function in DNA repair genes like MLH1 and MSH2 can lead to hypermutation and microsatellite instability in colorectal cancer. Other causes include problems with DNA replication, chromosome segregation, and telomere dysfunction. Genetic disorders involving genomic instability include ataxia-telangiectasia, neurofibromatosis type 1, Bloom syndrome, and ring chromosomes.