Embed presentation
Download to read offline



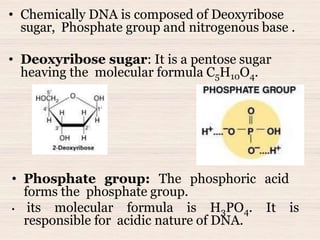
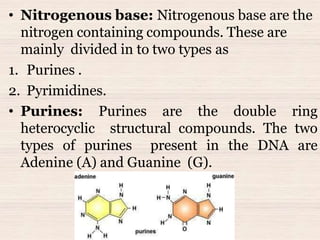
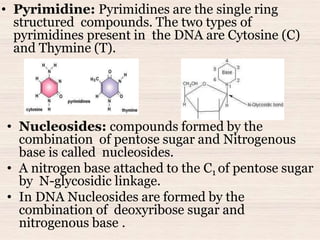
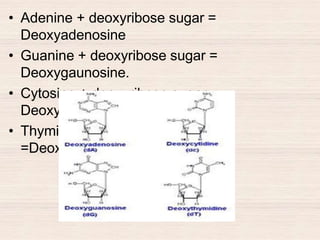
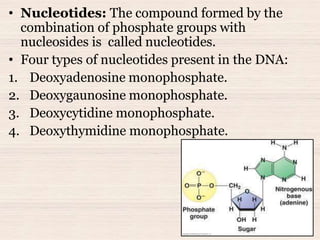
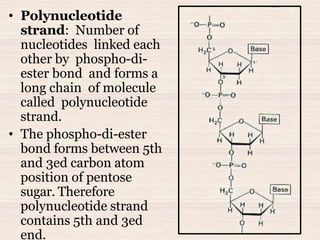





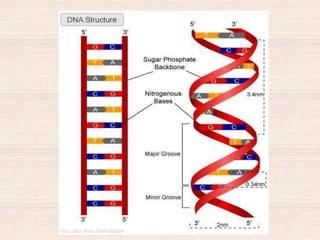
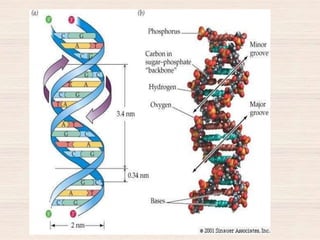


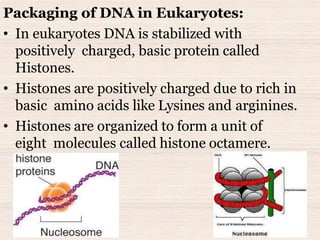
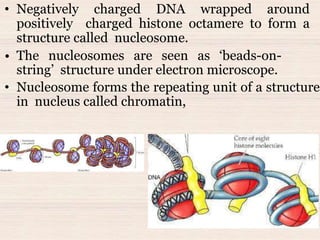


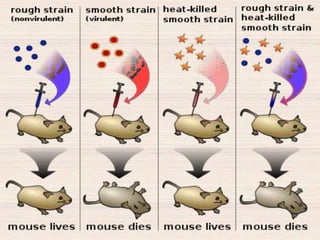



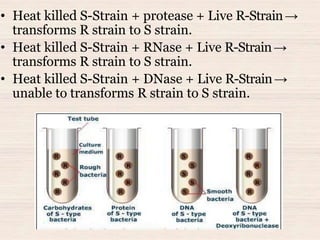

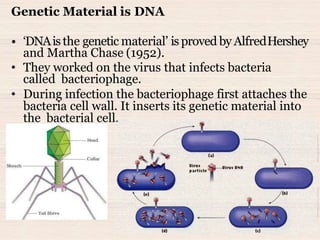

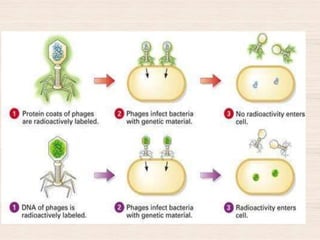







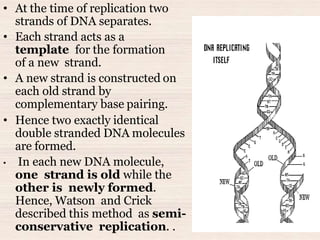
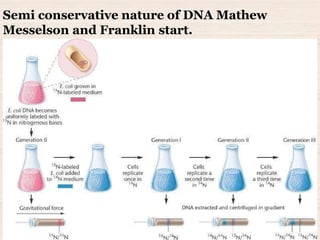
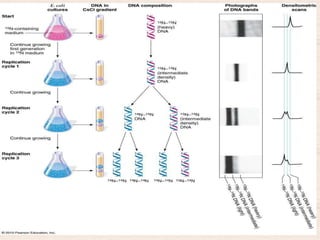
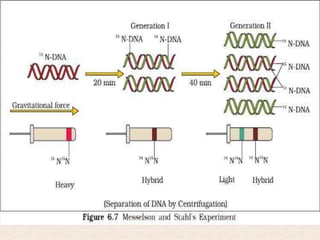
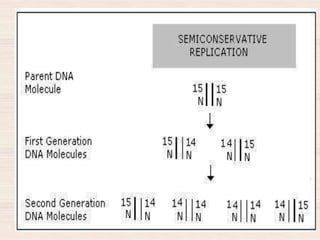


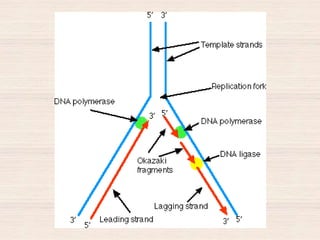
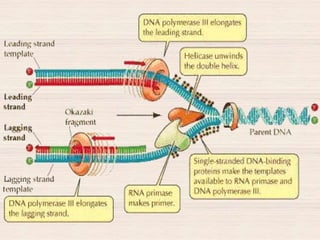



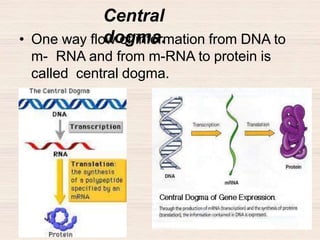

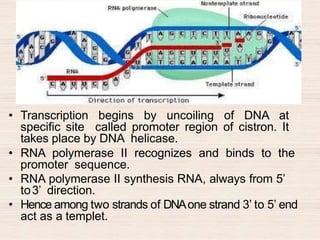
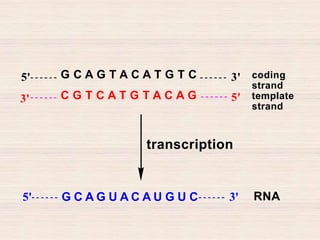
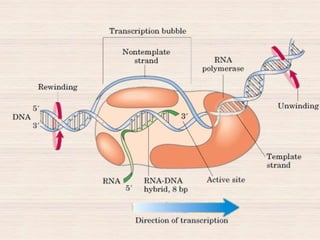





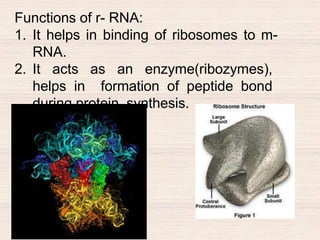








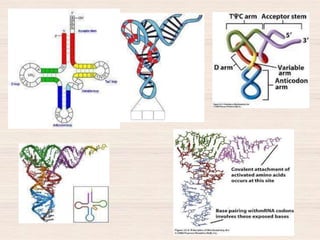

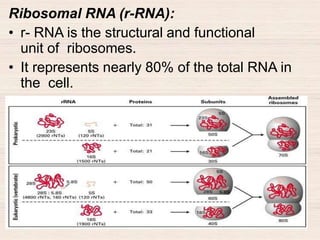



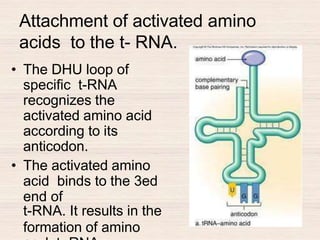
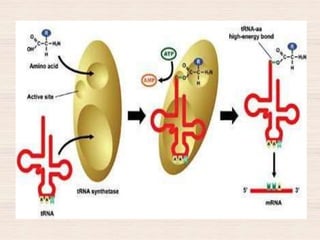
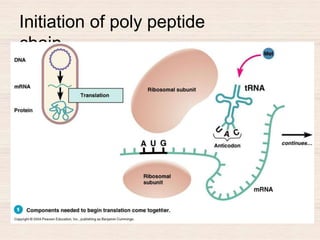

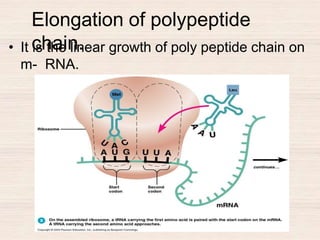

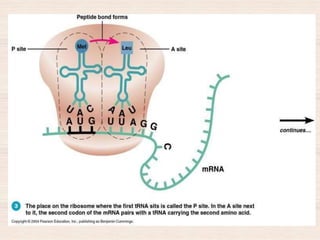
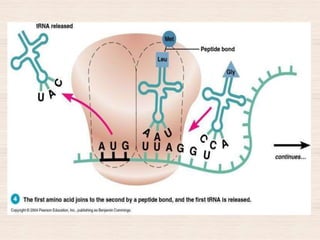
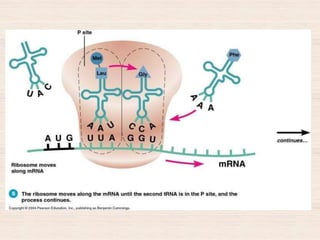
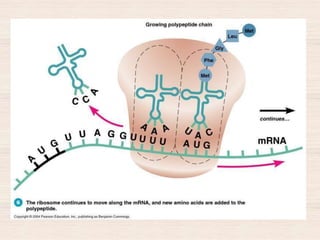

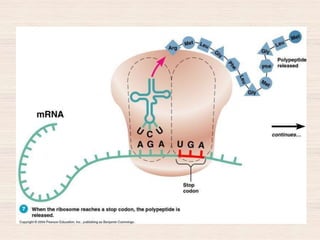
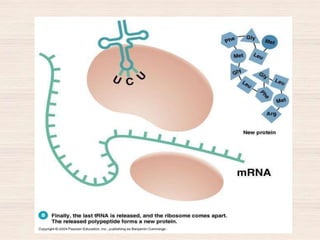



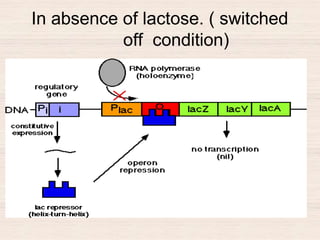
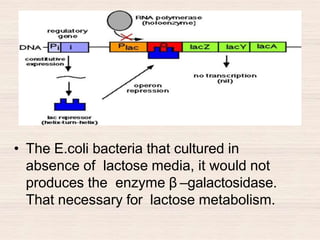
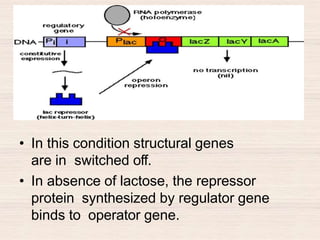
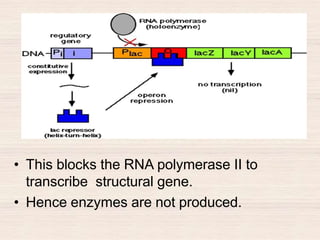
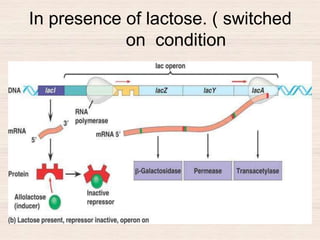
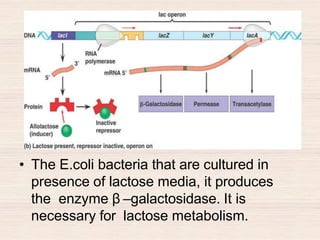
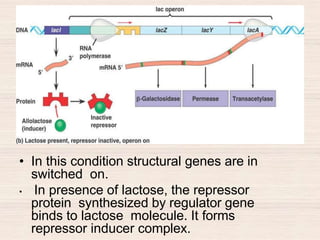
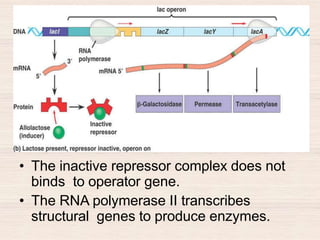
















This document discusses the structure and replication of DNA. It provides details on: 1. The key components of DNA including deoxyribose sugar, phosphate groups, and nitrogenous bases. It also describes how nucleotides and polynucleotide strands are formed. 2. The double helix structure of DNA proposed by Watson and Crick, including how the strands are antiparallel and connected through complementary base pairing. 3. Semiconservative replication of DNA, where each old DNA strand acts as a template for a new complementary strand, resulting in two identical double-stranded DNA molecules after replication. 4. Experiments by Meselson and Stahl using isotopes that provided evidence supporting the































































































































