Applications of transcriptomice s in modern biotechnology 2
- 1. APPLICATIONS OFTRANSCRIPTOMICS IN MODERN BIOTECHNOLOGY
- 2. Presented to Dr.M.naveed Presented by Javeria sami Pakeeza rubab Rashiqa nosheen Amara hameed Laiba shoaib • L1f16mcbt0006 • L1f16mcbt0016 • L1f16mcbt0011 • L1f16mcbt0010 • l1f16mcbt0020
- 3. CONTENTS 1. INTRODUCTION 2. HISTORY 3. SCOPE AND AIMS OF TRANSCRIPTOMICS 4. TYPES OF RNA 5. FUNCTIONS OF RNA 6. APPLICATIONS OF TRANSCRIPTOMICS 7. TECHNIQUIES OF TRANSCRIPTOMICS 8. ETHICAL ISSUES
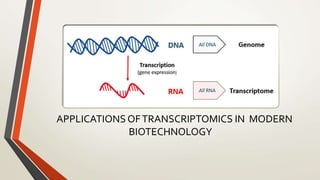
- 4. INTRODUCTION • TRANSCRIPTION It is the process where from DNA,the single stranded RNA is formed. • TRANSCRIPTS The readouts of genes are called transcripts • TRANSCRIPTOME A complete set of transcript in a cell and their quantitiy in a specific developmental stages or physical condition. TRANSCRIPTOMICS The set of all RNA molecules including mRNA, tRNA and rRNA.& non coding and coding RNA produced in one population of cell.
- 5. History • 1970 libraries of of silkmoth Mrna transcripts were collected and converted to complementary DNA (Cdna) for storage using reverse transcriptase) • 1980 ( sanger method was used to sequence random transcripts,producing expressed sequence tags (ESTs) • 1990 ( RNA microarray red expressed sequence tag blue and serial/cap analysis of gene expression 1990s ( ESTs came to prominence as an efficient method to determine the gene content of an organism without sequencing the entire genome) • 1991 ( The first attempt at capturing a partial human transcriptome was published • 2008 ( two human transcriptomes , composed of millions of transcript-derived sequences covering 16000 genes
- 6. Aims of transcriptomics • To determine the transcriptional structure of gene • To catalogue all species of transcript,including mRNAs, non-coding RNAs and small RNAs • In terms of their start sites, 5’ and 3’ ends , splicing patterns and other post transcriptional modifications • To quantify the changing expression levels of each transcript during different conditions
- 7. RNA It stands for RIBONUCLEIC ACID it is single stranded nucleic Acid and plays in a role in transferring information from DNA to protein forming system of the cell structure.
- 9. TYPES OF RNA TYPES Abbr functions 1 Messenger RNA mRNA Codes for protein 2 Ribosomal RNA rRNA translation 3 Transfer RNA tRNA translation 4 Small nuclear RNA snRNA Splicing and other functions 5 Small nucleolar RNA Sno RNA Nucleotide modification of RNAs 6 Micro RNA miRNA Gene regulation 7 8 Small interfering RNA Spliced Leader RNA RNase P Gene regulation TRNA maturation
- 10. APPLICATIONS IN TRANSCRIPTOMICES • 1. Agriculture and plant breeding • 2. Stem cells • 3. Use in risk assessment of chemicals • 4. Health n disease • 5. Expression profiling for disease • 6. Responses to environment
- 11. TRANSCRIPTOMICES IN RELATIONTO PLANT BREEDING & AGRICULTURE • Microsatellite markers • Detection • RNAi
- 12. • In agricultural crops • RNA interference technology • RNAi is an ancident evolutionary mechanism adopted by plants as a defence strategy • It is applicable to plants to acquire new traits • Small RNA characterization • it is non-protein coding small RNA molecules ranging from 20-30 nt that have a role in development,genome maintaince and plant responses to environmental stresses
- 13. • Two major groups • Micro RNAs : these are about 21 nt and usually have a post transcriptional regulatory role by directing cleavage of a specific transcript • Short interfering RNAs : these are usually 24nt long and influence de novo methylation or other modifications to silence genes • eQTLs: metabolite,protein and transcript profile can also be directly mapped onto a segregating population to provide information on loci that control gene expression level, protein modification • Association with traits eQTL, pQTL, or mQTL
- 14. Stem cell relation to transcriptomics
- 15. EXOSOMES FOR CARDIAC REPAIR • Transfering stem cells • Approaches to enhance the efficacy of stem cell therapies • Cell free components • Promote cardiac function in the pathological heart • Variety of RNA closely associated with gene expression • Variety of cellular processes • Non –coding RNA
- 16. Transcriptomics biomarker in safetly and risk assessment of chemicals • Markers for exposure to chemicals and therapeutic agents • Transcriptomic marker • Mechanism of cell injury,cell death or carcinogenic transformation • Predictor of heath • Drug safety assessments are more limited
- 17. APPLICATIONS OF TRANSCRIPTOMICS IN HEALTH & DISEASE • Application of transcriptome in autoimmune disease • Transcriptome analysis of adrenocortical cells in health and disease • Single cell transcriptomics to explore the immune system in health and disease
- 18. Expression profiling for disease Study of transcriptomics also known as expression profiling. Gene expression profiling The identification & characterization of mixture of RNA that is present in specific sample .
- 19. Response to environment • Transcriptome allow identification of genes • Biotic and abiotic stress • Novel transcriptional networks
- 20. APPLICATIONS OFTRANSCRIPTOMICES IN AUTOIMMUNE DISEASES • To identify molecular pathways involved in the inflammatory processes • To investigate the transcriptional changes • Differential expression of miRNA , a class of small noncoding RNAs that regulate gene expression • Deregulated in autoimmune diseases • Role of miRNA • Blood transcriptome • Examples of successful blood transcriptome
- 21. Techniques in transcriptomics • These are the following techniques in trasncriptomics: • Real-time PCR • Microarray • Next generation sequencing
- 22. Real-time PCR • RealTime PCR is based on the detection of the fluorescence produced by a reporter molecule which increases, as the reaction proceeds. a technology used for quantification of DNA sequences amplified in PCR
- 23. History of real-time PCR • 1993: First DNA detection method by using of EtBr. • 1996: useTaqMan detection method instead of EtBR. • 1996-1997: ABI first introduced the real-time PCR. • (Since many more instrument manufacture and many more detection method have been developed
- 24. Role of real-time PCR • Detection • Quantification • analysis
- 25. How does real-time PCR work • To amplify a segment of DNA using PCR, the sample is first heated so the DNA denatures, or separates into two pieces of single-stranded DNA. Next, an enzyme called "Taq polymerase" synthesizes - builds - two new strands of DNA, using the original strands as templates.
- 26. • On x-axis number of PCR cycle, y-axis show the fluorescent of amplification. • During exponential phase the amount of PCR product approximately double in each cycle. • As a reaction proceed reaction component are consumed and ultimately one or more of the components become limiting.in this point we show that the reaction become slow and enter the plateau phase. • Fluorescence at background levels and increase of fluorescence are not detectable then amplified product accumulate to yield a detectable fluorescence signal the cycle number at which occurred is called quantification cycle (Cq). • Large amount of template the reaction will have low or early. • Small amount of template the reaction will high or late Cq.
- 27. Microarray • A microarray is a laboratory tool used to detect the expression of thousand of gene at the same time
- 28. DNA microarray procedure: • There are following step of microarray: • Collection sample • Isolate mRNA • Hybridization • Detection the relative intensities of fluorescent under microarray scanner • Analyzed data
- 30. NEXT GENERATION SEQUENCING It is also known as high throughput approaches to DNA sequencing. It is also called massive parallel sequencingNext-generation sequencing (NGS), also known as high-throughput sequencing, is the catch-all term used to describe a number of different modern sequencing technologies including: Illumina (Solexa) sequencing.
- 31. Types of next generation sequencing • ILLUMINA OR SOLEXA SEQUENCING • DNA NANOBALL SEQUENCING • SOLID SEQUENCING
- 32. ILLUMINA OR SOLEXA SEQUENCING • The Illumina Solexa sequencing technology uses sequencing-by-synthesis on an eight- channel flowcell to produce more than 10 million reads per channel with read lengths up to 100bp. Individual fragments of a genomic DNA library are amplified on a flowcell via bridge-PCR to generate clusters of identical fragments.
- 33. DNA NANOBALL SEQUENCING • It is used to determine the entire genomic sequence of an organism.This technology has been used for multiple genome sequencing project and is scheduled to be used for more.
- 34. Solid sequencing • Solid (Sequencing by Oligonucleotide Ligation and Detection) is a next generation DNA sequencing technology developed by Life Technologies and has been commercially available since 2006. This next generation technology generates hundreds of millions to billions of small sequence reads at one time
- 35. ETHICAL ISSUES • Issues in ownership of an individual’s DNA • Security and sharing of genomic data • Increasing use of genetic variation screening both in newborn and adult • Screening for genetic variations can be harmful increasing in anxiety in individuals.
- 36. THE END