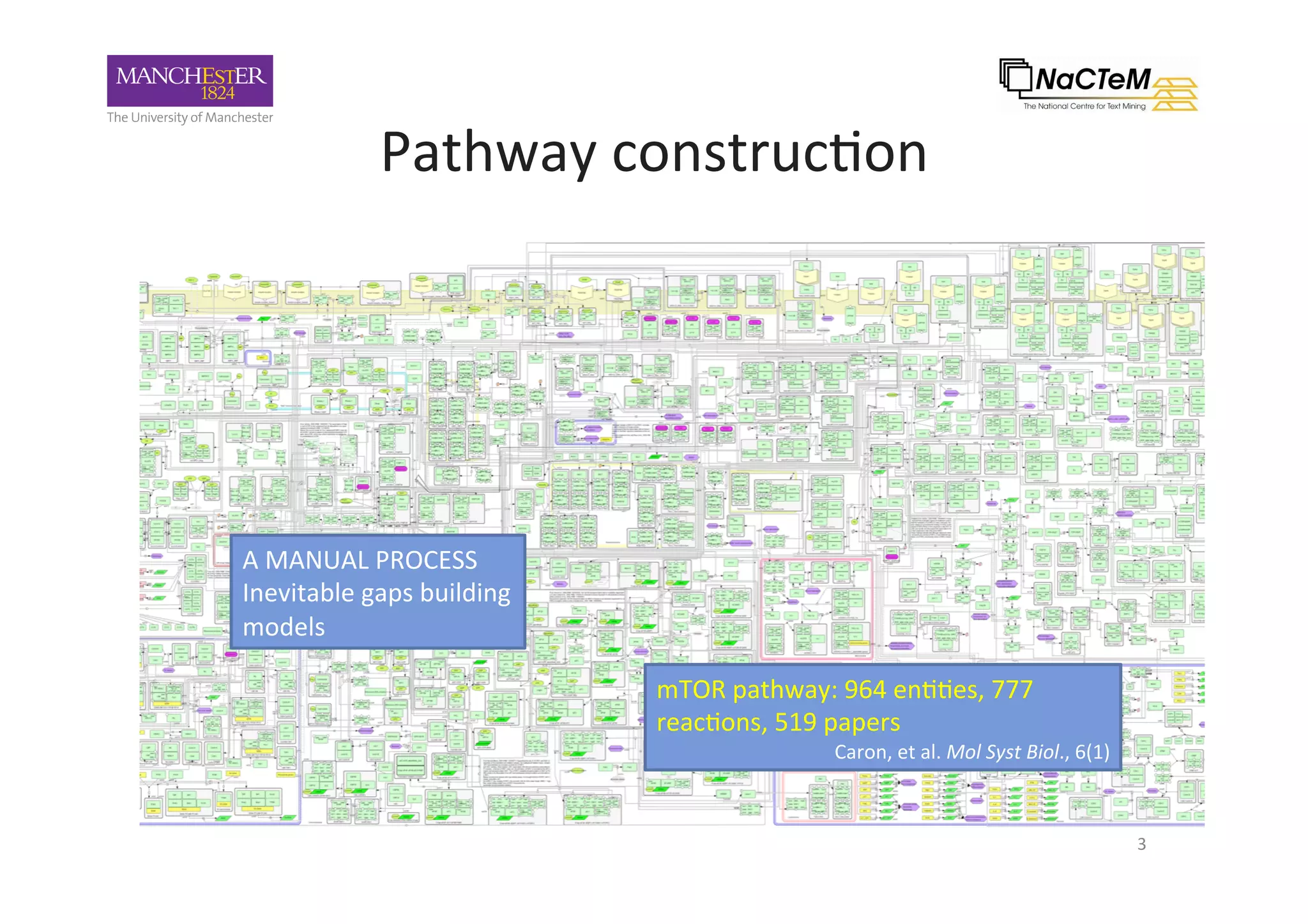
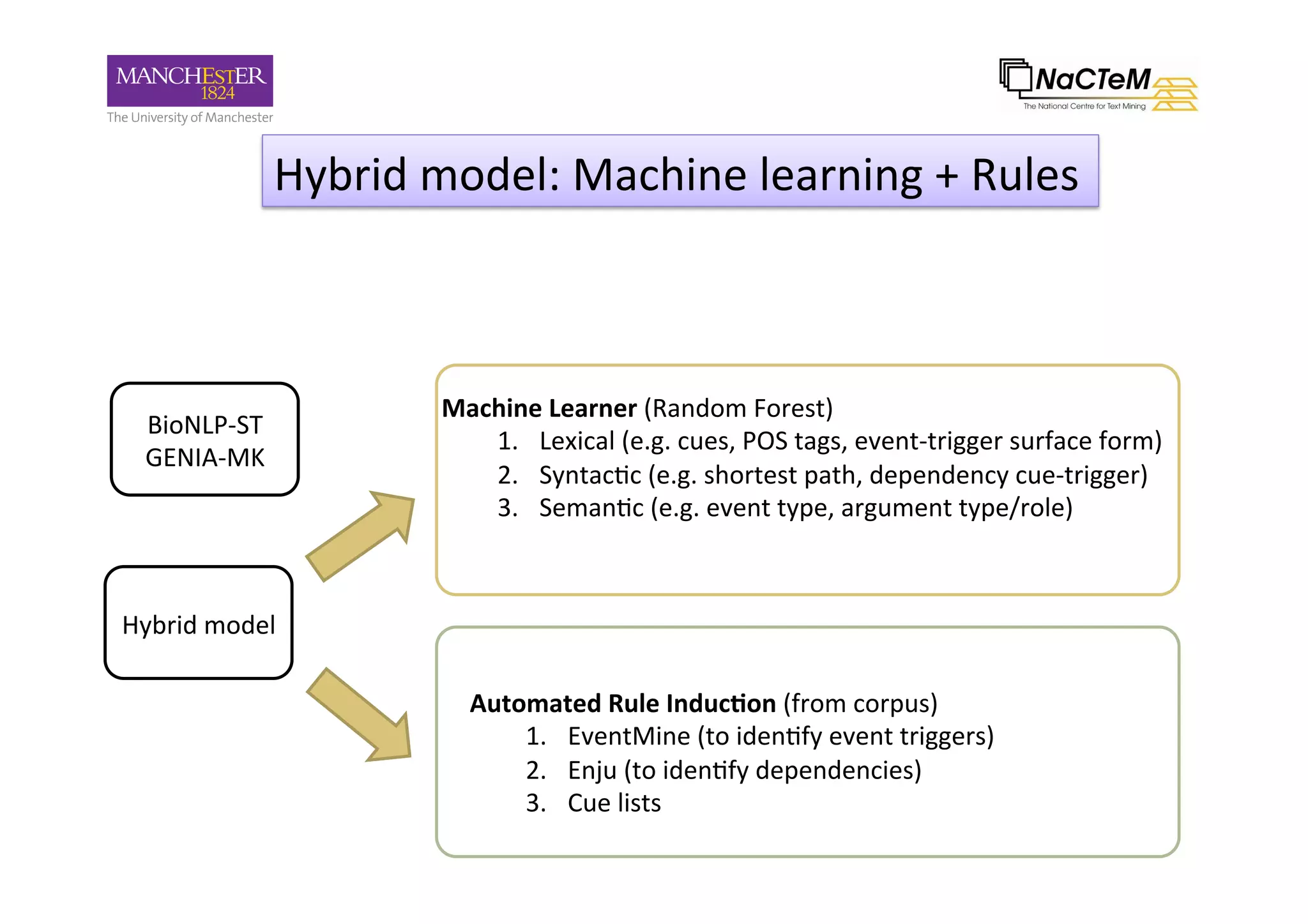
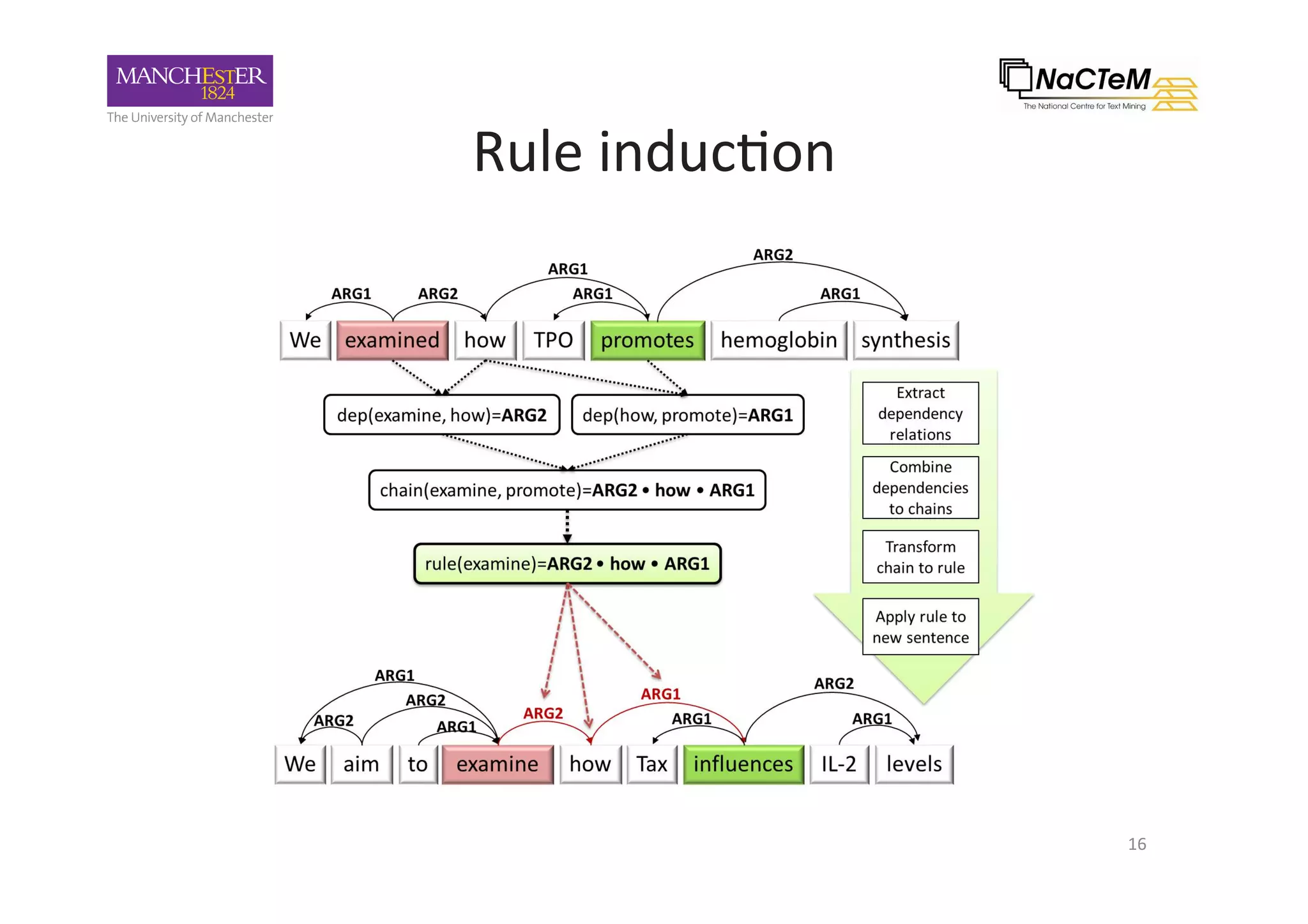
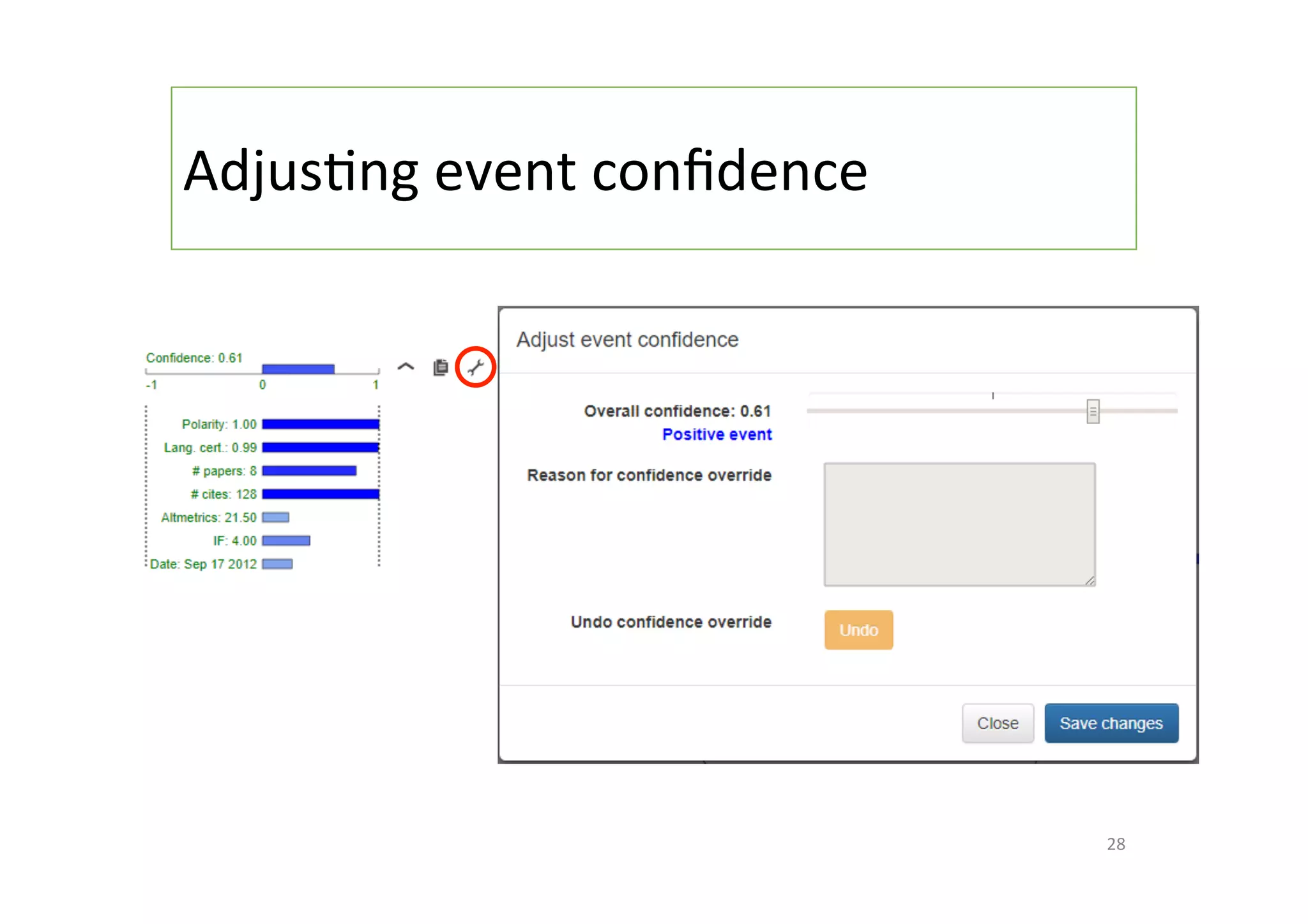
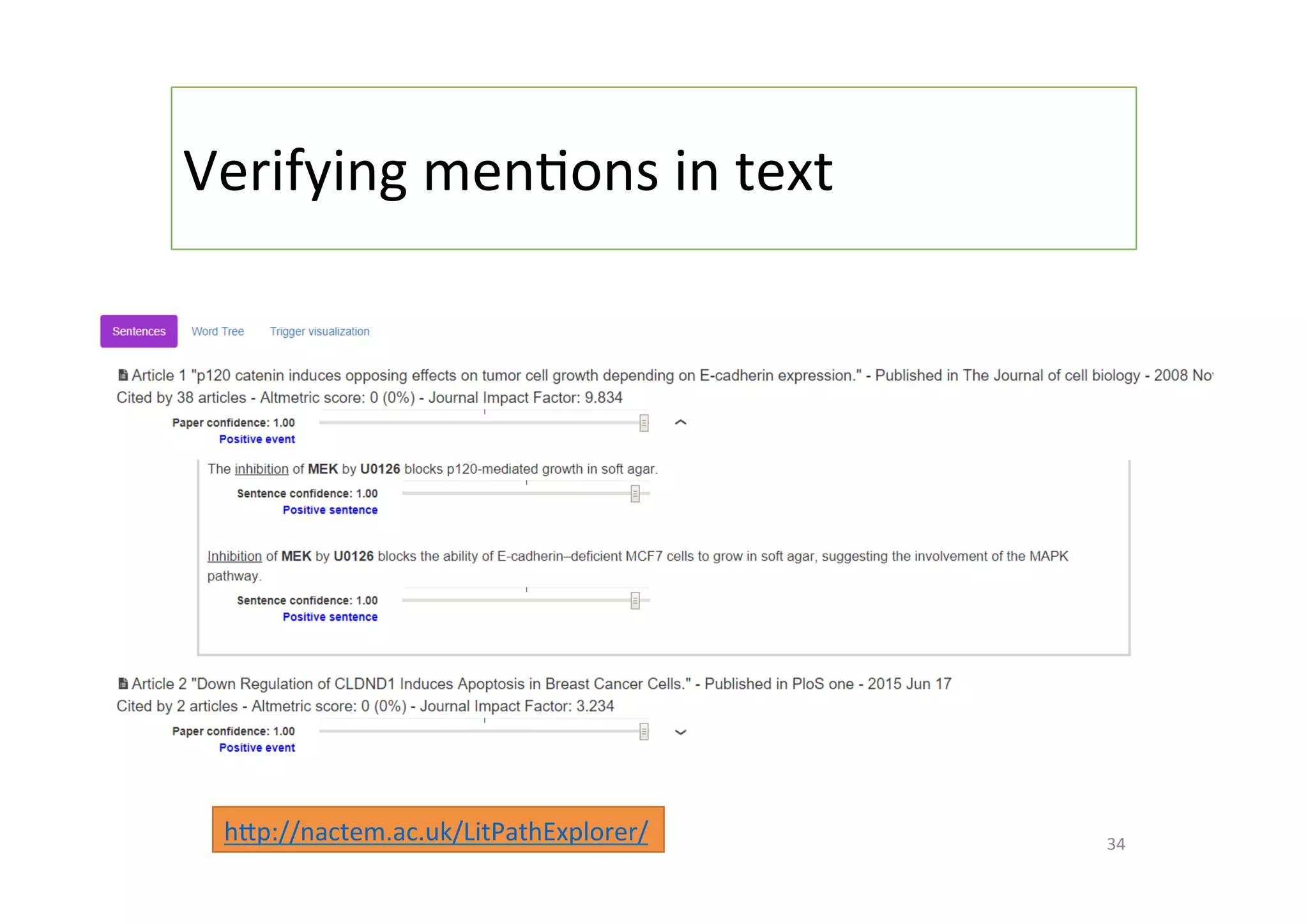
The document discusses a project named 'Big Mechanism' aimed at enhancing cancer biology understanding through deep reading and text mining. It highlights the challenges of manual annotation in literature mining, the integration of uncertainty measures into event extraction, and introduces tools like LitPathExplorer for visualizing and validating biological pathways. The initiative focuses on increasing the efficiency and accuracy of research by enabling automated processes and improving access to scientific knowledge.