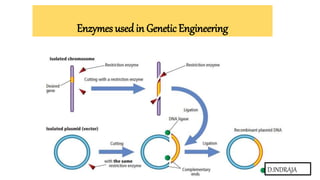
ENZYMES USED IN GENETIC ENGINEERING
- 1. Enzymes used in Genetic Engineering D.INDRAJA
- 2. • The ability to manipulate DNA in vitro depends entirely on the availability of purified enzymes that can cleave, modify and join the DNA molecule in specific ways. • At present, no chemical method can achieve the ability to manipulate the DNA in vitro in a predictable way. • Only enzymes are able to carry out the function of manipulating the DNA. • Each enzyme has a vital role to play in the process of genetic engineering. • The various enzymes used in genetic engineering are as follows: Restriction endonucleases polymerases DNA ligases S1 nuclease Alkaline phosphatase Poly nucleotide kinase
- 4. What are restriction enzymes? Molecular scissors that cuts DNA. Identifies specific Recognition sites. Found naturally in prokaryotes as a defense mechanism. Do not cut host DNA- But how? A useful tool in DNA modification and manipulation.
- 5. Source of Restriction Enzymes •The natural source of restriction endonucleases are bacterial cells. •These enzymes are called restriction enzymes because they restrict infection of bacteria by certain viruses (i.e., bacteriophages), by degrading the viral DNA without affecting the bacterial DNA. Thus, their function in the bacterial cell is to destroy foreign DNA that might enter the cell. • Restriction enzymes prevent the replication of the phage by cleaving its DNA at specific sites. • The host DNA is protected by Methylases which add methyl groups to adenine or cytosine bases within the recognition site thereby modifying the site and protecting the DNA. •The restriction enzyme recognizes the foreign DNA and cuts it at several sites along the molecule. •Each bacterium has its own unique restriction enzymes and each enzyme recognizes only one type of sequence.
- 6. History • The existence of restriction enzymes was first postulated by W. Arber. • He noticed that when the DNA of a bacteriophage entered a host bacterium it was cut up into smaller pieces and, for this, he theorized the presence of restriction enzyme. • In 1970, Hamilton Smith and his co-workers first isolated a restriction enzyme from the bacterium Haemophilus influenzae strain . • The enzyme, called Hindll, recognizes a six base-pair dsDNA sequence. After discovery of Hindll restriction enzyme, EcoRI, was isolated and characterized from Escherichia coli strain
- 7. • Restriction enzyme, also called restriction endonuclease, is a protein produced by bacteria that cleaves DNA at specific sites along the molecule. • Restriction endonucleases cut the DNA double helix in very precise ways. It cleaves DNA into fragments at or near specific recognition sites within the molecule known as restriction sites. • They have the capacity to recognize specific base sequences on DNA and then to cut each strand at a given place. Hence, they are also called as ‘molecular scissors’. Nomenclature of Restriction Enzymes • Each enzyme is named after the bacterium from which it was isolated, using a naming system based on bacterial genus, species and strain. For example, the name of the EcoRI restriction enzyme was derived as: • E – Escherichia: Genus • co- coli: specific species • R- RY13: strain • I- First identified: order of identification in the bacterium
- 8. Types of Restriction Enzymes Traditionally, four types of restriction enzymes are recognized, designated I, II, III, and IV, which differ primarily in structure, cleavage site, specificity, and cofactors. Type I restriction enzymes Type II restriction enzymes Type III restriction enzymes Type IV restriction enzymes These types are categorization based on: • Their composition. • Enzyme co-factor requirement. • The nature of their target sequence. • position of their DNA cleavage site relative to the target sequence.
- 9. Type I enzymes • These cleave at sites remote from a recognition site; require both ATP and S-adenosyl-L-methionine to function; multifunctional protein with both restriction and methylase activities. • one enzyme with different subunits for recognition ,cleavage ,and methylation ,Recognizes and methylates a single sequence but cleaves DNA up to 1000 bp away . • Type I restriction enzymes possess three subunits called HsdR, HsdM, and HsdS; • HsdR is required for restriction digestion • HsdM is necessary for adding methyl groups to host DNA (methyltransferase activity), • HsdS is important for specificity of the recognition (DNA-binding) site in addition to both restriction digestion (DNA cleavage) and modification (DNA methyltransferase) activity. TYPE III ENDONUCLEASES • Cleave DNA at immediate vicinity, about 20-30 base pairs away from recognition sequence. • Recognises two separate non-palindromic sequences that are inversely oriented. • ATP, SAM (not essential) and Mg2+ acts as co-factor. • Separate enzymes for restriction and modification, but share a common subunit. • They are components of prokaryotic DNA restriction-modification mechanisms that protect the organism against invading foreign DNA • Type III enzymes are hetero-oligomeric, multifunctional proteins composed of two subunits, Res (P08764) and Mod (P08763).
- 10. • The Mod subunit recognises the DNA sequence specific for the system and is a modification methyltransferase; as such, it is functionally equivalent to the M and S subunits of type I restriction endonuclease. • Res is required for restriction digestion, although it has no enzymatic activity on its own. • These enzymes methylate only one strand of the DNA, at the N-6 position of adenosyl residues, so newly replicated DNA will have only one strand methylated, which is sufficient to protect against restriction digestion. Type IV enzymes • These target modified DNA, e.g. methylated, hydroxy methylated and glucosyl-hydroxy methylated DNA. • Two different enzymes but recognition sequence is symmetric , cleavage occurs in one side of recognition sequence up to 20 bp away . Type V enzymes • Type V restriction enzymes (e.g., the cas9-gRNA complex from CRISPRs) utilize guide RNAs to target specific non-palindromic sequences found on invading organisms. • They can cut DNA of variable length, provided that a suitable guide RNA is provided. The flexibility and ease of use of these enzymes make them promising for future genetic engineering applications
- 11. Type II enzymes • cleave within or at short specific distances from a recognition site • They form homodimers, with recognition sites that are usually undivided and palindromic and 4–8 nucleotides in length. • They recognize and cleave DNA at the same site, and they do not use ATP or AdoMet for their activity— they usually require only Mg2+ as a cofactor. • It can either cleave at the centre of both strands to yield a blunt end, or at a staggered position leaving overhangs called sticky ends. • These are the most commonly available and used restriction enzymes. • They are different subtypes in type II they are: Type IIS, Type IIE, Type IIF, Type IIT, Type IIG, Type IIB, Type IIM
- 12. Recognition Sites •The DNA sequences recognized by restriction enzymes are called palindromes. • Palindromes are the base sequences that read the same on the two strands but in opposite directions. • For example, if the sequence on one strand is GAATTC read in 5’→3′ direction, the sequence on the opposite strand is CTTAAG read in the 3’→5′ direction, but when both strands are read in the 5’→ 3′ direction the sequence is the same. • In addition, there is a point of symmetry within the palindrome. In the example, this point is in the center between the AT/AT. •The value of restriction enzymes is that they make cuts in the DNA molecule around this point of symmetry. •Some enzymes cut straight across the molecule at the symmetrical axis producing blunt ends. •Of more value, however, are the restriction enzymes that cut between the same two bases away from the point of symmetry on two strands, thus, producing a staggering break. 5′ GAATTC 3′ 3′ CTTAAG 5′
- 13. Palindrome Sequences • The mirror like palindrome in which the same forward and backwards are on a single strand of DNA strand • The Inverted repeat palindromes is also a sequence that reads the same forward and backwards, but the forward and backward sequences are found in complementary DNA strands • Inverted repeat palindromes are more common and have greater biological importance than mirror- like palindromes Inverted repeat palindromes
- 14. Mechanism of Action • Restriction Endonuclease scan the length of the DNA , binds to the DNA molecule when it recognizes a specific sequence and makes one cut in each of the sugar phosphate backbones of the double helix – by hydrolyzing the phosphodiester bond. • Specifically, the bond between the 3’ O atom and the P atom is broken. • Direct hydrolysis by nucleophilic attack at the phosphorous atom • 3’OH and 5’ PO4 3- is produced. Mg2+ is required for the catalytic activity of the enzyme. • It holds the water molecule in a position where it can attack the phosphoryl group and also helps polarize the water molecule towards deprotonation
- 15. Sticky ends • Most restriction enzymes make staggered cuts • Staggered cuts produce single stranded “sticky-ends” • “Sticky Ends” Are Useful DNA fragments with complimentary sticky ends can be combined to create new molecules which allows the creation and manipulation of DNA sequences from different sources. • As an example of how a restriction enzyme recognizes and cuts at a DNA sequence, let's consider EcoRI, a common restriction enzyme used in labs. EcoRI cuts at the following site • If another piece of DNA has matching overhangs (for instance, because it has also been cut by EcoRI), the overhangs can stick together by complementary base pairing. For this reason, enzymes that leave single-stranded overhangs are said to produce sticky ends. Sticky ends are helpful in cloning because they hold two pieces of DNA together so they can be linked by DNA ligase “overhangs”
- 16. Blunt ends • Some restriction enzymes cut DNA at opposite base • They leave blunt ended DNA fragments • They are harder to ligate together (the ligation reaction is less efficient and more likely to fail) because there are no single-stranded overhangs to hold the DNA molecules in position. • These blunt ended fragments can be joined to any other DNA fragment with blunt ends. • Enzymes useful for certain types of DNA cloning experiments • HindII, PvuII and AluI are examples of blunt end cutters
- 18. • Blunt ends are modifies and used in R DNA technology with the help of oligonucleotides such as • Linkers • Adaptors • Homopolymer tails Linkers • Linkers are the chemically synthesized double stranded DNA oligonucleotides containing on it one or more restriction sites for cleavage by restriction enzymes, e.g. Eco RI, Hind III, Bam HI, etc. • Linkers are ligated to blunt end DNA by using DNA ligase. • Both the vector and DNA are treated with restriction enzyme to develop sticky ends. • The staggered cuts i.e. sticky ends are then ligated with T4 DNA ligase with very high efficiency to the termini of the vector and recombinant plasmid DNA molecules are produced.
- 19. Limitations It may be the case that the restriction enzyme used to generate the cohesive ends in the linker will also cut the foreign DNA at internal sites. Solution: CHOOSE ANOTHER RESTRICTION ENZYME
- 20. Adaptors • An adaptor is a short, chemically synthesized, double stranded DNA molecule which is used to link the ends of two other DNA molecules. • These are the molecules having sticky ends so they can ligate to the blunt ends of the DNA and by using DNA ligase recombinants can be constructed Problem • The sticky ends of individual adaptors could base pair with themselves to form dimers and the new DNA molecule remains blunt-ended.
- 21. Solution • Adaptor molecules alter their 5' terminus(From 5'-P to 5'-OH) by an enzymatic treatment of the enzyme Alkaline phosphatase to prevent self ligation. • Afterwards, they can be treated with Polynucleotide kinases to restore it & ligate to vectors.
- 22. Terminal transferase / homo polymer tails • It is the enzyme that converts blunt end of DNA fragments into sticky end. • If the restriction enzyme cuts DNA forming blunt ends, then efficiency of ligation is very low. So the enzyme terminal transferase converts bunt end into sticky end. • Terminal transferase enzyme synthesize short sequence of complementary nucleotide at free ends of DNA, so that blunt end is converted into sticky end. • This group of enzymes catalyses the addition of one or more deoxyribonucleotides to the 3' terminus of the DNA molecule. • The enzyme can work on both double stranded as well as single stranded DNA molecules without the need of any primers. • The enzyme is obtained from calf thymus tissue.
- 23. • The enzyme is used for labelling 3'ends of DNA. Also, it can be used for adding complementary homopolymeric tails to DNA molecules Applications of terminal transferase in creating recombinant DNA.
- 24. ISOSCHIZOMERS • Restriction enzymes that have the same recognition sequence as well as the same cleavage site are Isoschizomers NEOSCHIZOMERS • Restriction enzymes that have the same recognition sequence but cleave the DNA at a different site within that sequence are Neoshizomers Artificial restriction enzymes • Artificial restriction enzymes can be generated by fusing a natural or engineered DNA binding domain to a nuclease domain (often the cleavage domain of the type IIS restriction enzyme FokI). • Such artificial restriction enzymes can target large DNA sites (up to 36 bp) and can be engineered to bind to desired DNA sequences. • Zinc finger nucleases are the most commonly used artificial restriction enzymes and are generally used in genetic engineering applications • Artificial ribonucleases that act as restriction enzymes for RNA are also being developed.
- 25. Factors affecting the activity of restriction enzymes 1.Star activity: Under sub-optimal reaction conditions, some restriction enzymes cleave base sequences at sites different from the defined recognition sequence. In other words, they cleave at non-specific sites. This phenomenon is called star activity. • Some of the factors that induce star activity are high salt and glycerol concentration, presence of impurities, excessive enzyme compared to substrate DNA, increased incubation time, or incompatible buffer and cofactor. 2. Methylated DNA: Several DNA molecules are methylated at the recognition site, making them resistant to cleavage by certain restriction enzymes. • For example, most E. coli strains express Dam or Dcm methyltransferases that methylate specific recognition sites to form G6mATC and C5mCA/TGG, respectively. G6mATC is resistant to cleavage by Mbo I. 3. Temperature: Most endonucleases optimally digest the target DNA at 37 °C. However, there are some exceptions with lower or higher optimal temperatures. For example, Taq I optimally digests at 65 °C and Apa I digests at 25 °C
- 26. Applications • They are used in gene cloning and protein expression experiments • Restriction enzymes are used in biotechnology to cut DNA into smaller strands in order to study fragment length differences among individuals ( Restriction Fragment Length Polymorphism- RFLP). • Each of these methods depends on the use of agarose gel electrophoresis for separation of the DNA fragments.
- 27. POLYMERASES
- 28. DNA polymerases • These are enzymes that catalyse the synthesis of a new DNA strand from a pre-existing strand. The enzyme adds deoxyribonucleotides to the free 3’-OH of the chain undergoing elongation. The direction of synthesis is 5’-3’.Ithas three major requirements for its activity (1) a template strand for which the enzyme synthesizes a complementary strand (2) a primer with a free 3’-OH group that hybridizes with the template to form a double stranded region that initiates the polymerization (3) a pool of all the four dNTPs that are used to synthesize the new DNA strand. In addition, some cofactors like Mg2+ ions may be required in a buffer solution with correct pH for optimum activity
- 29. • Different types of DNA polymerases are used in recombinant DNA technology. We will study the following types in detail. 1. E. coliDNA Polymerase I 2. Klenow Fragment 3. Thermostable DNA Polymerase 4. Reverse Transciptase E. coli DNA polymerase I • E. coli DNA polymerase I (PolI) is an enzyme that has both DNA polymerase as well as DNA nuclease activity. • This enzyme binds to the ‘nick’ region (region of a double stranded DNA where one or more nucleotides of one strand are missing, making it single stranded). • The polymerase activity of the enzyme synthesizes the complementary strand for the nick and continues synthesizing the complete new strand by simultaneously degrading the pre-existing strand by its 5'- 3'exonuclease activity.
- 30. • If the E. coli Pol I holoenzyme is treated with a mild protease, it results in the formation of two fragments. A larger fragment retaining both 5'-3' polymerase and 3'-5' exonuclease activities; while the smaller one has only the 5'-3' exonuclease activity. The larger fragment is known as ‘Klenow fragment’. Klenow fragment
- 31. • This Klenow fragment can synthesize the new DNA strand complementary to the template but cannot degrade the existing strand. • Klenow fragment is predominantly used in DNA sequencing. • Other uses in recombinant DNA technology where Klenow fragment is used are • Synthesis of double stranded DNA from single stranded template. • Filling of 5’ overhangs created by restriction enzymes to create blunt ends. • Digestion of protruding 3’ overhangs to produce blunt ends. Thermostable DNA polymerases • Thermostable DNA polymerases are a class of DNA polymerases that remain functional even at high temperatures. • In other words, they are resistant to denaturation by heat treatment. • They are isolated from the bacterium Thermus aquaticus that lives in hot springs. • The enzyme isolated from this bacterium is known as‘Taq Polymerase’.
- 32. • Major application of Taq polymerase is in the polymerase chain reaction (PCR) technique which is used to amplify DNA fragments Reverse transcriptase (RT) • Reverse transcriptase (RT) is aRNA dependent DNA polymerase found in RNA viruses also called as retroviruses. • This enzyme is involved in the replication of retroviruses, where the RNA genome is first converted into DNA and then integrated into the host. • RT uses mRNA template instead of DNA for synthesizing new DNA strand. • The complementary DNA strand formed on the mRNA template is called the complementary DNA (cDNA). • RT also shows RNAseH activity that degrades the RNA molecule from a DNA-RNA hybrid. • Formation of a double stranded cDNA from the mRNA molecule using RT finds applications in genetic engineering. • The cDNA thus formed from any mRNA can be cloned in an expression vector and its respective protein can be made to express in large quantities
- 33. RNA polymerases • In all organisms, RNA synthesis is carried out by Enzymes known as RNA polymerases (RNAPs) -- that transcribe the genetic information from DNA in a highly-regulated, multi-stage process. • RNAP is the key enzyme involved in creating an equivalent complementary RNA copy of a sequence of DNA a process called transcription. • RNA polymerase (RNAP or RNApol), also known as DNA-dependent RNA polymerase, is an enzyme that produces RNA. • RNA polymerase enzymes are essential to life and are found in all organisms and many viruses. • In chemical terms, RNAP is a nucleotidyl transferase that polymerizes ribonucleotides at the 3' end of an RNA transcript. RNA polymeras Types of RNA polymerase • Prokaryotic RNA polymerase • Eukaryotic RNA polymerase
- 34. • Prokaryotic (Bacteria, viruses, archaea) organisms have a single type of RNA polymerase that synthesizes all the subtypes of RNA • The efficiency of E. coli RNA polymerase is around 40 nt/sec at 37ºC, and requires Mg2+ (RNA polymerase ofT3 andT7 are single polypeptides with a efficiency of 200 nt/sec) • The enzyme binds over a region of DNA covering around 60 bp E. coli RNA polymerase • The RNA polymerase of E.coli is a very large molecule (about 480,000 da or 480k daltons)containing six polypeptide chains:α2,β, β’,ω andσ • The core enzyme is composed of five α2,β, β’,ω chains and catalyzes RNA synthesis
- 35. • Eukaryotes have three different RNA polymerases: • RNAP I-transcribes rRNA genes (nucleoli) • RNAP II -transcribes mRNA genes (nucleoplasm) • RNAP III - transcribes tRNA, 5S RRNA, and other small RNAgenes (nucleoplasm) • All eukaryotic RNA polymerases are large proteins, appearing as aggregates of >500 kDa Each RNA pol is a multi-subunit protein (8 -12 subunits) Eukaryotic pol II consists of 12 subunits. • The two largest subunits are homologous to the bacterial and subunits. • In addition to the increased number of subunits, eukaryotic pol II differs from its prokaryotic counterpart in that it has a series of heptad repeats with the consensus sequence Tyr-Ser-Pro-Thr Ser Pro-Ser at the carboxyl terminal of the largest pol II subunit. – 52X humans , - 26X yeast • RNA Polymerase IV- si RNA biogenesis • RNA Polymerase V - si RNA detected DNA methylation
- 36. APPLICATIONS OF POLYMERASES • The polymerase chain reaction (PCR) • DNA sequencing • Single nucleotide polymorphism (SNP) detection • Whole genome amplification (WGA) • Synthetic biology • Molecular diagnostics.
- 37. DNA LIGASES
- 38. • If two DNA molecules have matching ends, they can be joined by the enzyme DNA ligase. DNA ligase seals the gap between the molecules, forming a single piece of DNA. • The ligases used in DNA cloning is to join together fragments of newly synthesized DNA to form a seamless strand(unbroken molecule) • Catalyses the formation of phosphodiester bonds between the 5' phosphate of one strand and the 3' hydroxyl group of another. • DNA ligases are mg++ dependent enzymes . • ‘Molecular glue’ – used to join DNA fragments to produce recombinant DNA • Types of DNA Ligases • 1. E.coli ligase – requires NAD as cofactor • 2. T4 DNA ligase – uses ATP as cofactor
- 39. Mechanism of action: • ATP in case of T4 DNA ligase and NAD+ in case of E. coli ligase first split and form an enzyme-AMP complex • The complex then binds to the nick that must expose a 5′ phosphate and 3’OH group. • The AMP then reacts with the phosphate group. • The 3’OH group on the moiety then generates a new phosphodiester bond which seals the nick
- 40. Example: Building a recombinant plasmid DNA LIGASE
- 41. Applications • DNA ligases are used with restriction enzymes to insert DNA fragments, often genes, into plasmids. • Helps to perform blunt end as well as sticky end ligation. • Some ligases are used as targets for the development of new antibacterial drugs. • ligation of blunt ended termini: this reaction is much slower than ligation of sticky ends and the ligation is improved by addition of monovalent cation and low concentration of PEG . • Ligation of synthetic linkers or adapter
- 42. S1 NUCLEASE
- 43. • The S1 nuclease was extracted from Aspergills oryzae. • The S1 nuclease is a specific single-stranded endonuclease. It can degrade single-stranded DNA and single-stranded RNA to produce 5'-single-stranded nucleotides or oligonucleotides. • Double-stranded DNA, and DNA-RNA hybrid molecules are more resistant to S1 nucleases, and only high concentrations of enzymes allow them to be digested. • It hydrolyzes single-stranded DNA at a rate that is 75 000 times faster than hydrolysed double-stranded DNA. • It is a heat stable enzyme that functions at high ionic strength, low pH and in the presence of Zn2+ ions. • The optimum pH for the enzymatic reaction was 4.0 to 4.3, and the enzyme activity decreased by 50% at pH 4.9. • Some chelating agents (such as EDTA and citric acid) can strongly inhibit S1 nuclease activity. • In addition, phosphate buffer and 0.6% SDS solution can inhibit its activity, but it is stable to urea and formamide • Another type of endonuclease called as DNase I that is isolated from cow’s pancreas is a non– specific enzymes. • It is able to cleave both single and double stranded DNAs. • It can cleave any of the internal phosphodiester bonds, thus prolonged digestion of DNA with DNase I results in its complete chewing leaving only a mixture of mononucleotides.
- 45. Nomenclature • Alternative names include endonuclease S1 (Aspergillus) • single-stranded-nucleate endonuclease • deoxyribonuclease S1 • ,Aspergillus nuclease S1 • Neurospora crassa single-strand specific endonuclease • S1 nuclease, single-strand endodeoxyribonuclease • single-stranded DNA specific endonuclease • single-strand-specific endodeoxyribonuclease • single strand-specific DNase • Aspergillus oryzae S1 nuclease Uses • Aspergillus nuclease S1 is used in the laboratory as a reagent in nuclease protection assays. • In molecular biology, it is used in removing single stranded tails from DNA molecules to create blunt ended molecules and opening hairpin loops generated during synthesis of double stranded cDNA
- 47. Alkaline phosphatase (AP) • This group of enzymes removes the phosphate group (PO3 2- ) from 5' terminus of the DNA molecule. • It is active at alkaline pH, hence the name ‘alkaline phosphatase’. • Commercially, it is obtained from three major sources, viz., E. coli (bacteria), calf intestine and arctic shrimp. • Treatment of vector DNA with AP is important in cloning experiments, as removal of 5'phosphate prevents self-annealing of the digested vector and increases the possibility of ligating with the insert DNA fragment in the presence of ligase. Also, radiolabeled DNA probes are prepared by initially removing the5'PO3 2- by AP treatment, followed by polynucleotide kinase treatment in the presence of radioactive phosphate.
- 48. Application of alkaline phosphatase treatment to prevent recircularization of vector plasmid with out insertion of foreign DNA
- 50. Polynucleotide kinase (PNK) • This group of enzymes perform a role completely opposite to the one performed by AP. • PNK catalyses the transfer of a phosphate group from ATP to the 5' terminus of the DNA molecule after dephosphorylation by alkaline phosphatase. • This enzyme is obtained from E. coli infected with T4 phage. • It can be performed in 3 ways they are 1. Forward reaction 2. Exchange reaction 3. Dephosphorylation reaction Forward reaction 5’OH DNA +ATP (rp32) 5’p32DNA+ADP PNK
- 51. Exchange reaction 5’P DNA +ADP 5’OH DNA +ATP ATP ATP*(P32) radio labelled 5’OH DNA + ATP (rp32) 5’p32 DNA + ADP 5’p DNA 5’OH DNA +pi 5’OH DNA + ATP*(r p32) 5’p32 DNA +ADP PNK Dephosphorylation reaction AP PNK