Towards diagnosis of rotator cuff tears in 3-D MRI using 3-D convolutional neural networks
•
0 likes•141 views
Towards diagnosis of rotator cuff tears in 3-D MRI using 3-D convolutional neural networks. Paper presented at the Workshop on Computational Biology at the International Conference on Machine Learning, Long Beach, USA, 2019.
Report
Share
Report
Share
Download to read offline
Recommended
FST Conference 2017Performance of a Novel Knee Implant to Treat Osteoarthritis Without Violatin...

Performance of a Novel Knee Implant to Treat Osteoarthritis Without Violatin...Rajshree Hillstrom, PhD MBA
This study demonstrates that 3D-MRI is able to evaluate the anterolateral ligament fully in all normal knees. The classification system for injury to the ALL described shows high inter- and intra-observer reliability3D-MRI Evaluation of the Anterolateral Ligament: An Evaluation of ACL Deficie...

3D-MRI Evaluation of the Anterolateral Ligament: An Evaluation of ACL Deficie...Adnan Saithna - Orthopedic Surgeon, Scottsdale, Arizona
Recommended
FST Conference 2017Performance of a Novel Knee Implant to Treat Osteoarthritis Without Violatin...

Performance of a Novel Knee Implant to Treat Osteoarthritis Without Violatin...Rajshree Hillstrom, PhD MBA
This study demonstrates that 3D-MRI is able to evaluate the anterolateral ligament fully in all normal knees. The classification system for injury to the ALL described shows high inter- and intra-observer reliability3D-MRI Evaluation of the Anterolateral Ligament: An Evaluation of ACL Deficie...

3D-MRI Evaluation of the Anterolateral Ligament: An Evaluation of ACL Deficie...Adnan Saithna - Orthopedic Surgeon, Scottsdale, Arizona
To maximize outcome in nerve transfers:
1- The recipient nerve reinnervated close to the target muscle.
2- Direct repair without intervening grafts.
3- Similarly behaving neuromuscular units (agonistic donors and recipients)
Predictors of Patients’ Functional Outcome after Motor Nerve Transfers in Man...

Predictors of Patients’ Functional Outcome after Motor Nerve Transfers in Man...Professor M. A. Imam
The Indian Dental Academy is the Leader in continuing dental education , training dentists in all aspects of dentistry and offering a wide range of dental certified courses in different formats.
Indian dental academy provides dental crown & Bridge,rotary endodontics,fixed orthodontics,
Dental implants courses.for details pls visit www.indiandentalacademy.com ,or call
0091-9248678078
Advances in diagnostic aids/certified fixed orthodontic courses by Indian den...

Advances in diagnostic aids/certified fixed orthodontic courses by Indian den...Indian dental academy
Computer Navigated Medial Opening Wedge High Tibial Osteotomy- Review of Literature by Kunal Dhurve* in Crimson Publishers: Orthopedic Research and Reviews JournalComputer Navigated Medial Opening Wedge High Tibial Osteotomy- Review of Lite...

Computer Navigated Medial Opening Wedge High Tibial Osteotomy- Review of Lite...CrimsonPublishersOPROJ
More Related Content
What's hot
To maximize outcome in nerve transfers:
1- The recipient nerve reinnervated close to the target muscle.
2- Direct repair without intervening grafts.
3- Similarly behaving neuromuscular units (agonistic donors and recipients)
Predictors of Patients’ Functional Outcome after Motor Nerve Transfers in Man...

Predictors of Patients’ Functional Outcome after Motor Nerve Transfers in Man...Professor M. A. Imam
The Indian Dental Academy is the Leader in continuing dental education , training dentists in all aspects of dentistry and offering a wide range of dental certified courses in different formats.
Indian dental academy provides dental crown & Bridge,rotary endodontics,fixed orthodontics,
Dental implants courses.for details pls visit www.indiandentalacademy.com ,or call
0091-9248678078
Advances in diagnostic aids/certified fixed orthodontic courses by Indian den...

Advances in diagnostic aids/certified fixed orthodontic courses by Indian den...Indian dental academy
Computer Navigated Medial Opening Wedge High Tibial Osteotomy- Review of Literature by Kunal Dhurve* in Crimson Publishers: Orthopedic Research and Reviews JournalComputer Navigated Medial Opening Wedge High Tibial Osteotomy- Review of Lite...

Computer Navigated Medial Opening Wedge High Tibial Osteotomy- Review of Lite...CrimsonPublishersOPROJ
What's hot (20)
Is Medial Ridge Sign a Reliable Indicator Glenoid Bone Loss-Dr. Dhanasekarapr...

Is Medial Ridge Sign a Reliable Indicator Glenoid Bone Loss-Dr. Dhanasekarapr...
Predictors of Patients’ Functional Outcome after Motor Nerve Transfers in Man...

Predictors of Patients’ Functional Outcome after Motor Nerve Transfers in Man...
Introduction to Navigation - Robotic Total Knee Replacement 

Introduction to Navigation - Robotic Total Knee Replacement
Grip Strength Rehabilitation Using a Monocular Camera(AsianCHI2020)

Grip Strength Rehabilitation Using a Monocular Camera(AsianCHI2020)
Addressing bone loss in shoulder instability lennard funk

Addressing bone loss in shoulder instability lennard funk
Correlation of Antero Inferior Glenoid Bone Loss with Number of Dislocations ...

Correlation of Antero Inferior Glenoid Bone Loss with Number of Dislocations ...
13. design of ergonomic stool shabila et al 299-305

13. design of ergonomic stool shabila et al 299-305
An Automated Pelvic Bone Geometrical Feature Measurement Utilities on Ct Scan...

An Automated Pelvic Bone Geometrical Feature Measurement Utilities on Ct Scan...
Advances in diagnostic aids/certified fixed orthodontic courses by Indian den...

Advances in diagnostic aids/certified fixed orthodontic courses by Indian den...
Computer Navigated Medial Opening Wedge High Tibial Osteotomy- Review of Lite...

Computer Navigated Medial Opening Wedge High Tibial Osteotomy- Review of Lite...
Similar to Towards diagnosis of rotator cuff tears in 3-D MRI using 3-D convolutional neural networks
ANALYTICAL AND QUANTITATIVE CYTOPATHOLOGY AND HISTOPATHOLOGY3D Reconstruction of Peripheral Nerves Based on Calcium Chloride Enhanced Mic...

3D Reconstruction of Peripheral Nerves Based on Calcium Chloride Enhanced Mic...ANALYTICAL AND QUANTITATIVE CYTOPATHOLOGY AND HISTOPATHOLOGY
Similar to Towards diagnosis of rotator cuff tears in 3-D MRI using 3-D convolutional neural networks (20)
Reliability of Three-dimensional Photonic Scanner Anthropometry Performed by ...

Reliability of Three-dimensional Photonic Scanner Anthropometry Performed by ...
EOS in MEDIC presented in Thailand, Dr NGUYEN VAN CONG, MEDIC MEDICAL CENTER

EOS in MEDIC presented in Thailand, Dr NGUYEN VAN CONG, MEDIC MEDICAL CENTER
MICROCALCIFICATION IDENTIFICATION IN DIGITAL MAMMOGRAM FOR EARLY DETECTION OF...

MICROCALCIFICATION IDENTIFICATION IN DIGITAL MAMMOGRAM FOR EARLY DETECTION OF...
3D Position Tracking System for Flexible Cystoscopy

3D Position Tracking System for Flexible Cystoscopy
Design of Modified Bio-Inspired Algorithm for Identification and Segmentation...

Design of Modified Bio-Inspired Algorithm for Identification and Segmentation...
3D Reconstruction of Peripheral Nerves Based on Calcium Chloride Enhanced Mic...

3D Reconstruction of Peripheral Nerves Based on Calcium Chloride Enhanced Mic...
Difference in Early Results Between Sub-Acute and Delayed ACL reconstruction:...

Difference in Early Results Between Sub-Acute and Delayed ACL reconstruction:...
Biomechanical Mapping in Pelvic Organ Prolapse Management

Biomechanical Mapping in Pelvic Organ Prolapse Management
MURA Dataset: Towards Radiologist-Level Abnormality Detection in Musculoskele...

MURA Dataset: Towards Radiologist-Level Abnormality Detection in Musculoskele...
Knee surg sports traumatol arthrosc 2016 24 (11) 3599

Knee surg sports traumatol arthrosc 2016 24 (11) 3599
Olga Senyukova - Machine Learning Applications in Medicine

Olga Senyukova - Machine Learning Applications in Medicine
More from Wesley De Neve
More from Wesley De Neve (20)
Investigating the biological relevance in trained embedding representations o...

Investigating the biological relevance in trained embedding representations o...
Impact of adversarial examples on deep learning models for biomedical image s...

Impact of adversarial examples on deep learning models for biomedical image s...
Learning Biologically Relevant Features Using Convolutional Neural Networks f...

Learning Biologically Relevant Features Using Convolutional Neural Networks f...
Center for Biotech Data Science at Ghent University Global Campus

Center for Biotech Data Science at Ghent University Global Campus
Center for Biotech Data Science at Ghent University Global Campus

Center for Biotech Data Science at Ghent University Global Campus
Learning biologically relevant features using convolutional neural networks f...

Learning biologically relevant features using convolutional neural networks f...
Towards reading genomic data using deep learning-driven NLP techniques

Towards reading genomic data using deep learning-driven NLP techniques
Deep Machine Learning for Making Sense of Biotech Data - From Clean Energy to...

Deep Machine Learning for Making Sense of Biotech Data - From Clean Energy to...
GUGC Info Session - Informatics and Bioinformatics

GUGC Info Session - Informatics and Bioinformatics
Ghent University Global Campus - Sungkyunkwan University: Workshop on Researc...

Ghent University Global Campus - Sungkyunkwan University: Workshop on Researc...
Ghent University and GUGC-K: Overview of Teaching and Research Activities

Ghent University and GUGC-K: Overview of Teaching and Research Activities
Biotech Data Science @ GUGC in Korea: Deep Learning for Prediction of Drug-Ta...

Biotech Data Science @ GUGC in Korea: Deep Learning for Prediction of Drug-Ta...
Exploring Deep Machine Learning for Automatic Right Whale Recognition and No...

Exploring Deep Machine Learning for Automatic Right Whale Recognition and No...
Deep Machine Learning for Automating Biotech Tasks Through Self-Learning Expe...

Deep Machine Learning for Automating Biotech Tasks Through Self-Learning Expe...
Towards using multimedia technology for biological data processing

Towards using multimedia technology for biological data processing
Multimedia Lab @ Ghent University - iMinds - Organizational Overview & Outlin...

Multimedia Lab @ Ghent University - iMinds - Organizational Overview & Outlin...
Towards Twitter hashtag recommendation using distributed word representations...

Towards Twitter hashtag recommendation using distributed word representations...
Recently uploaded
Ultrasound color Doppler imaging has been routinely used for the diagnosis of cardiovascular diseases, enabling real-time flow visualization through the Doppler effect. Yet, its inability to provide true flow velocity vectors due to its one-dimensional detection limits its efficacy. To overcome this limitation, various VFI schemes, including multi-angle beams, speckle tracking, and transverse oscillation, have been explored, with some already available commercially. However, many of these methods still rely on autocorrelation, which poses inherent issues such as underestimation, aliasing, and the need for large ensemble sizes. Conversely, speckle-tracking-based VFI enables lateral velocity estimation but suffers from significantly lower accuracy compared to axial velocity measurements.
To address these challenges, we have presented a speckle-tracking-based VFI approach utilizing multi-angle ultrafast plane wave imaging. Our approach involves estimating axial velocity components projected onto individual steered plane waves, which are then combined to derive the velocity vector. Additionally, we've introduced a VFI visualization technique with high spatial and temporal resolutions capable of tracking flow particle trajectories.
Simulation and flow phantom experiments demonstrate that the proposed VFI method outperforms both speckle-tracking-based VFI and autocorrelation VFI counterparts by at least a factor of three. Furthermore, in vivo measurements on carotid arteries using the Prodigy ultrasound scanner demonstrate the effectiveness of our approach compared to existing methods, providing a more robust imaging tool for hemodynamic studies.
Learning objectives:
- Understand fundamental limitations of color Doppler imaging.
- Understand principles behind advanced vector flow imaging techniques.
- Familiarize with the ultrasound speckle tracking technique and its implications in flow imaging.
- Explore experiments conducted using multi-angle plane wave ultrafast imaging, specifically utilizing the pulse-sequence mode on a 128-channel ultrasound research platform. (May 9, 2024) Enhanced Ultrafast Vector Flow Imaging (VFI) Using Multi-Angle ...

(May 9, 2024) Enhanced Ultrafast Vector Flow Imaging (VFI) Using Multi-Angle ...Scintica Instrumentation
Recently uploaded (20)
Bhiwandi Bhiwandi ❤CALL GIRL 7870993772 ❤CALL GIRLS ESCORT SERVICE In Bhiwan...

Bhiwandi Bhiwandi ❤CALL GIRL 7870993772 ❤CALL GIRLS ESCORT SERVICE In Bhiwan...
POGONATUM : morphology, anatomy, reproduction etc.

POGONATUM : morphology, anatomy, reproduction etc.
Site specific recombination and transposition.........pdf

Site specific recombination and transposition.........pdf
Asymmetry in the atmosphere of the ultra-hot Jupiter WASP-76 b

Asymmetry in the atmosphere of the ultra-hot Jupiter WASP-76 b
Cyathodium bryophyte: morphology, anatomy, reproduction etc.

Cyathodium bryophyte: morphology, anatomy, reproduction etc.
Biogenic Sulfur Gases as Biosignatures on Temperate Sub-Neptune Waterworlds

Biogenic Sulfur Gases as Biosignatures on Temperate Sub-Neptune Waterworlds
Porella : features, morphology, anatomy, reproduction etc.

Porella : features, morphology, anatomy, reproduction etc.
Selaginella: features, morphology ,anatomy and reproduction.

Selaginella: features, morphology ,anatomy and reproduction.
(May 9, 2024) Enhanced Ultrafast Vector Flow Imaging (VFI) Using Multi-Angle ...

(May 9, 2024) Enhanced Ultrafast Vector Flow Imaging (VFI) Using Multi-Angle ...
TransientOffsetin14CAftertheCarringtonEventRecordedbyPolarTreeRings

TransientOffsetin14CAftertheCarringtonEventRecordedbyPolarTreeRings
Role of AI in seed science Predictive modelling and Beyond.pptx

Role of AI in seed science Predictive modelling and Beyond.pptx
Thyroid Physiology_Dr.E. Muralinath_ Associate Professor

Thyroid Physiology_Dr.E. Muralinath_ Associate Professor
Towards diagnosis of rotator cuff tears in 3-D MRI using 3-D convolutional neural networks
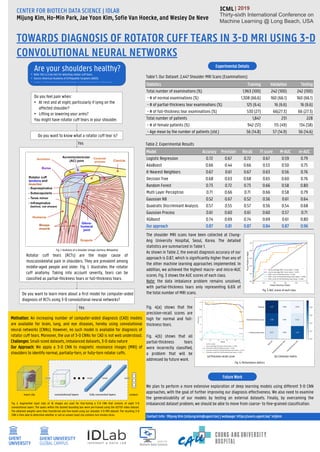
- 1. TOWARDS DIAGNOSIS OF ROTATOR CUFF TEARS IN 3-D MRI USING 3-D CONVOLUTIONAL NEURAL NETWORKS CENTER FOR BIOTECH DATA SCIENCE | IDLAB Mijung Kim, Ho-Min Park, Jae Yoon Kim, Sofie Van Hoecke, and Wesley De Neve Contact Info : Mijung Kim (mijung.kim@ugent.be) | webpage: https://users.ugent.be/~mijkim | 2019 Thirty-sixth International Conference on Machine Learning @ Long Beach, USA Do you feel pain when: • At rest and at night, particularly if lying on the affected shoulder? • Lifting or lowering your arms? You might have rotator cuff tears in your shoulder. Do you want to know what a rotator cuff tear is? Yes • Note: this is a toy test for detecting rotator cuff tears. • Source: American Academy of Orthopaedic Surgeons (AAOS) • https://orthoinfo.aaos.org/en/diseases--conditions/rotator-cuff-tears/ Fig. 1. Anatomy of a shoulder (image courtesy: Wikipedia) Rotator cuff tears (RCTs) are the major cause of musculoskeletal pain in shoulders. They are prevalent among middle-aged people and older. Fig. 1. illustrates the rotator cuff anatomy. Taking into account severity, tears can be classified as partial-thickness tears or full-thickness tears. Do you want to learn more about a first model for computer-aided diagnosis of RCTs using 3-D convolutional neural networks? Yes Motivation: An increasing number of computer-aided diagnosis (CAD) models are available for brain, lung, and eye diseases, hereby using convolutional neural networks (CNNs). However, no such model is available for diagnosis of rotator cuff tears. Moreover, the use of 3-D CNNs for CAD is not well understood. Challenges: Small-sized datasets, imbalanced datasets, 3-D data nature Our Approach: We apply a 3-D CNN to magnetic resonance images (MRI) of shoulders to identify normal, partially-torn, or fully-torn rotator cuffs. Fig. 2. Augmented input clips of 16 images are used for fine-tuning a 3-D CNN that consists of eight 3-D convolutional layers. The layers within the dashed bounding box were pre-trained using the UCF101 video dataset. The obtained weights were then transferred and fine-tuned using our shoulder 3-D MRI dataset. The resulting 3-D CNN is then able to determine whether or not an unseen input clip contains torn tendon slices. Table `1. Our Dataset: 2,447 Shoulder MRI Scans (Examinations) Statistics Training Validation Testing Total number of examinations (%) 1,963 (100) 242 (100) 242 (100) - # of normal examinations (%) 1,308 (66.6) 160 (66.1) 160 (66.1) - # of partial-thickness tear examinations (%) 125 (6.4) 16 (6.6) 16 (6.6) - # of full-thickness tear examinations (%) 530 (27) 66(27.3) 66 (27.3) Total number of patients 1,847 231 228 - # of female patients (%) 942 (51) 115 (49) 134 (58) - Age mean by the number of patients (std.) 56 (14.8) 57 (14.9) 56 (14.6) Table 2. Experimental Results Experimental Details Model Accuracy Precision Recall F1 score M-AUC m-AUC Logistic Regression 0.72 0.67 0.72 0.67 0.59 0.79 AdaBoost 0.66 0.44 0.66 0.53 0.50 0.75 K-Nearest Neighbors 0.67 0.61 0.67 0.63 0.56 0.76 Decision Tree 0.68 0.63 0.68 0.65 0.60 0.76 Random Forest 0.73 0.72 0.73 0.66 0.58 0.80 Multi Layer Perceptron 0.71 0.66 0.71 0.66 0.58 0.79 Gaussian NB 0.52 0.67 0.52 0.56 0.61 0.64 Quadratic Discriminant Analysis 0.57 0.55 0.57 0.56 0.54 0.68 Gaussian Process 0.61 0.60 0.61 0.60 0.57 0.71 XGBoost 0.74 0.69 0.74 0.69 0.61 0.80 Our approach 0.87 0.81 0.87 0.84 0.87 0.96 Are your shoulders healthy? The shoulder MRI scans have been collected at Chung- Ang University Hospital, Seoul, Korea. The detailed statistics are summarized in Table 1. As shown in Table 2, the overall diagnosis accuracy of our approach is 0.87, which is significantly higher than any of the other machine learning approaches implemented. In addition, we achieved the highest macro- and micro-AUC scores. Fig. 3 shows the AUC scores of each class. Note: the data imbalance problem remains unsolved, with partial-thickness tears only representing 6.6% of the total number of MRI scans. Fig. 3. AUC scores of each class Fig. 4(a) shows that the precision-recall scores are high for normal and full- thickness tears. Fig. 4(b) shows that all partial-thickness tears were incorrectly classified, a problem that will be addressed by future work. (a) Precision-recall curve (b) Confusion matrix Fig. 4. Performance metrics Future Work We plan to perform a more extensive exploration of deep learning models using different 3-D CNN approaches, with the goal of further improving our diagnosis effectiveness. We also need to examine the generalizability of our models by testing on external datasets. Finally, by overcoming the imbalanced dataset problem, we should be able to move from coarse- to fine-grained classification.
