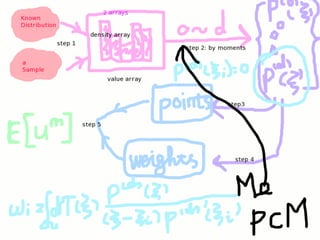
This document summarizes research on computing stochastic partial differential equations (SPDEs) using an adaptive multi-element polynomial chaos method (MEPCM) with discrete measures. Key points include:
1) MEPCM uses polynomial chaos expansions and numerical integration to compute SPDEs with parametric uncertainty.
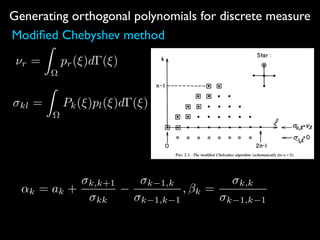
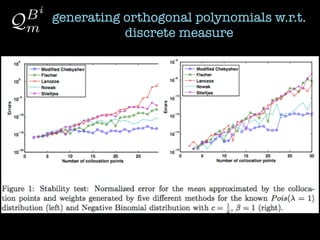
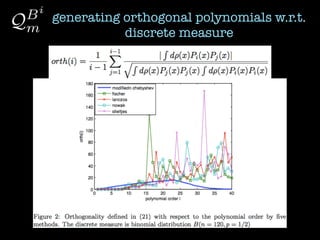
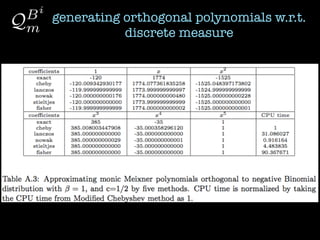
2) Orthogonal polynomials are generated for discrete measures using various methods like Vandermonde, Stieltjes, and Lanczos.
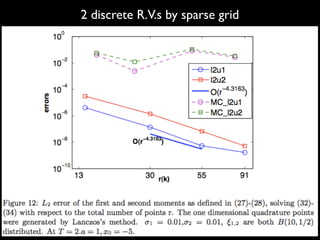
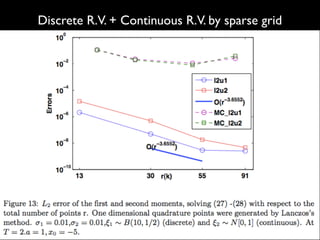
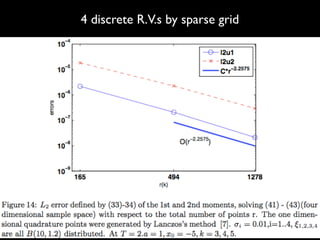
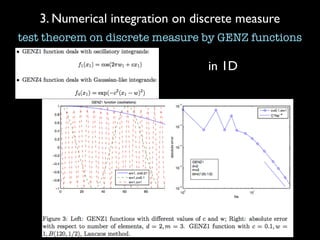
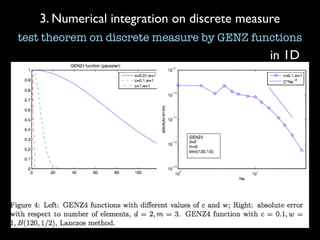
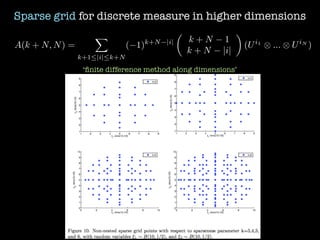
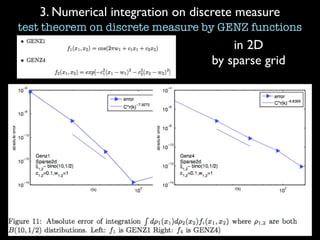
3) Numerical integration is tested on discrete measures using Genz functions in 1D and sparse grids in higher dimensions.
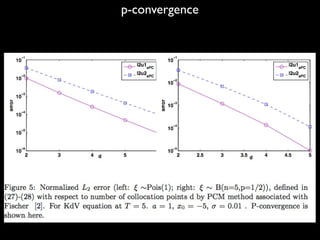
4) The method is demonstrated on the KdV equation with random initial conditions. Future work includes applying these techniques to SPDEs driven


![1.What computational SPDE is about? (MEPCM)
Xt(!) E[y(x, t; !)]
Xt(!)
Xt(⇠1, ⇠2, ...⇠n)
...
⇠n
⇠3
⇠2
⇠1
⌦
E[ym
(x, t; !)]
E[ym
(x, t; ⇠1, ⇠2, ..., ⇠n)]
fix x, t, integration
over a finite
dimensional
sample space
MEPCM=FEM on
sample space
⇠1
⇠2
⌦
⇡ ⇡
Gauss quadratures](https://image.slidesharecdn.com/spdepresentation2012-140626235328-phpapp01/85/SPDE-presentation-2012-3-320.jpg)






![Generating orthogonal polynomials for discrete measure
Fischer’s method
=
NX
i=1
i ⌧i ⌫ = + ⌧
↵⌫
i = ↵i +
2
i Pi(⌧)Pi+1(⌧)
1 +
Pi
j=0
2
j P2
j (⌧)
2
i 1Pi(⌧)Pi 1(⌧)
1 +
Pi 1
j=0
2
j P2
j (⌧)
⌫
i = i
[1 +
Pi 2
j=0
2
j P2
j (⌧)][1 +
Pi
j=0
2
j P2
j (⌧)]
[1 +
Pi 1
j=0
2
j P2
j (⌧)]2](https://image.slidesharecdn.com/spdepresentation2012-140626235328-phpapp01/85/SPDE-presentation-2012-12-320.jpg)

![Numerical example on KdV equation
ut + 6uux + uxxx = ⇠, x 2 R
u(x, 0) =
a
2
sech2
(
p
a
2
(x x0))
< um
(x, T; !) >=
Z
R
d⇢(⇠)[
a
2
sech2
(
p
a
2
(x 3 ⇠T2
x0 aT)) + ⇠T]m
L2u1 =
qR
dx(E[unum(x, T; !)] E[uex(x, T; !)])2
qR
dx(E[uex(x, T; !)])2
L2u2 =
qR
dx(E[u2
num(x, T; !)] E[u2
ex(x, T; !)])2
qR
dx(E[u2
ex(x, T; !)])2](https://image.slidesharecdn.com/spdepresentation2012-140626235328-phpapp01/85/SPDE-presentation-2012-22-320.jpg)