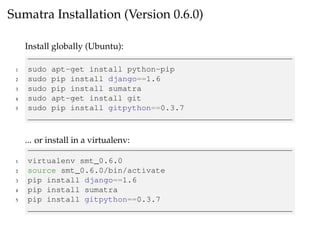
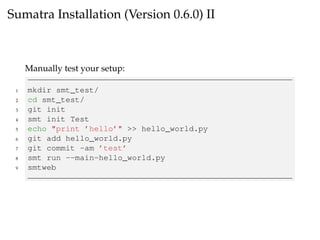
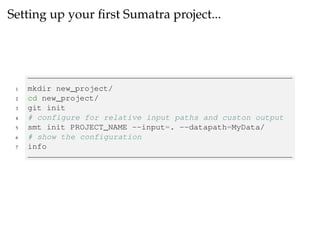
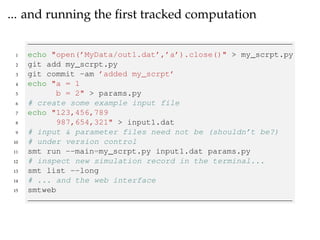
The document provides a comprehensive guide on using Git and Sumatra, including installation steps and basic commands for initialization, committing changes, and reverting to previous versions. It details how to set up and run a Sumatra project, along with useful resources and alternatives to Sumatra. Additionally, it highlights limitations of Sumatra and offers advice for tracking computations.