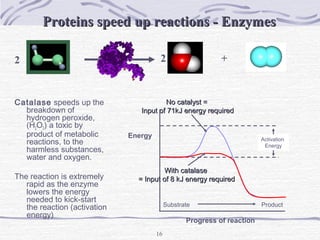
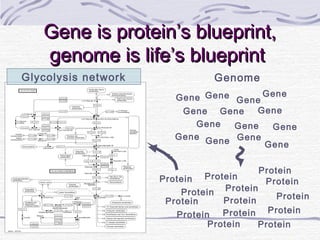
Proteins are essential biomolecules that make up 15% of cells and perform many critical functions. They are composed of amino acid monomers linked into polymers that fold into complex three-dimensional shapes defined by non-covalent interactions between residues. Protein structure determines its diverse functions, which include serving as enzymes, hormones, antibodies, structural components, transporters, and more. Understanding protein structure is necessary to elucidate proteins' roles at the molecular level.

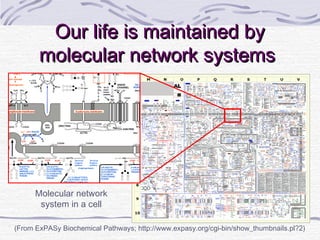








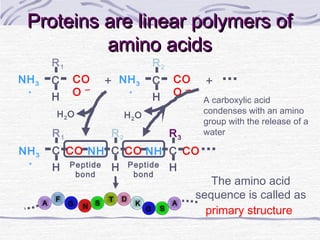
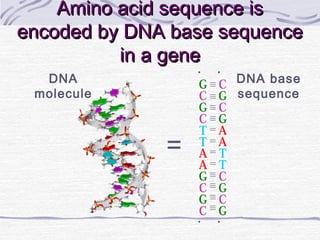
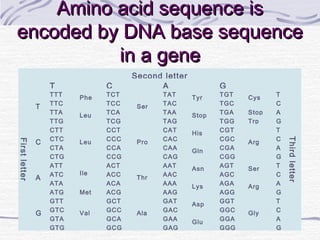
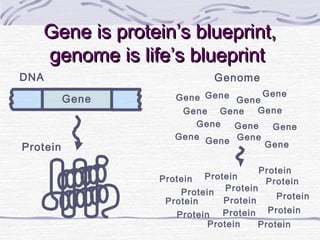
![Amino acid(s)
mg per kg body
weight
mg per
70 kg
mg per 100 kg Main food sources
H Histidine 10 700 1000
soy protein, eggs, parmesan, sesame,
peanuts[7]
I Isoleucine 20 1400 2000
eggs, soy protein & tofu, whitefish, pork
, parmesan[8]
L Leucine 39 2730 3900
eggs, soy protein, whitefish, parmesan
, sesame[9]
K Lysine 30 2100 3000
eggs, soy protein, whitefish, parmesan
, smelts[10]
M Methionine+ C Cysteine
10.4 + 4.1 (15
total)
1050 1500
eggs, whitefish, sesame, smelts,
soy protein[11]
+ eggs, soy protein,
sesame, mustard seeds,peanuts[12]
FPhenylalanine+ Y Tyrosine 25 (total) 1750 2500
eggs, soy protein, peanuts, sesame,
whitefish[13]
+ soy protein, eggs,
parmesan, sesame[14]
T Threonine 15 1050 1500
eggs, soy protein, whitefish, smelts,
sesame[15]
W Tryptophan 4 280 400
soy protein, sesame, eggs,
winged beans, chia seeds[16]
V Valine 26 1820 2600
eggs, soy protein, parmesan, sesame,
beef[17]](https://image.slidesharecdn.com/proteins-140630053132-phpapp01/85/Proteins-overview-11-320.jpg)


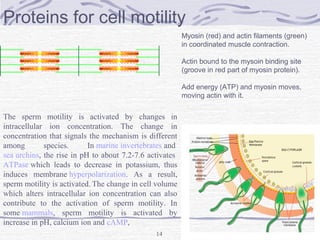







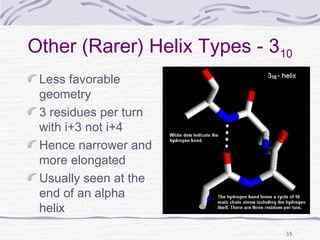

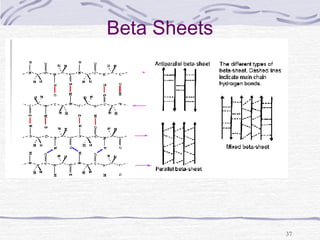





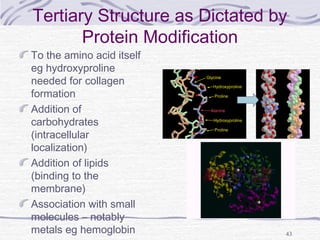
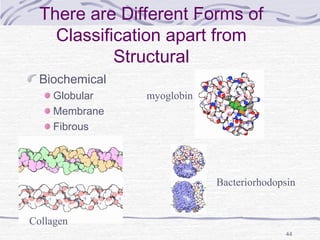

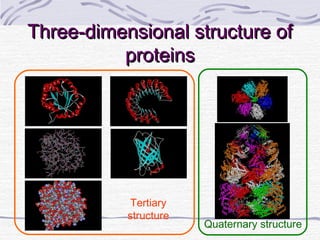









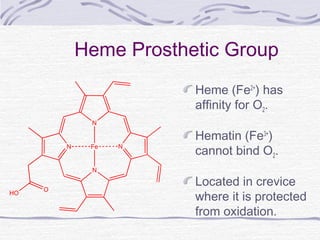



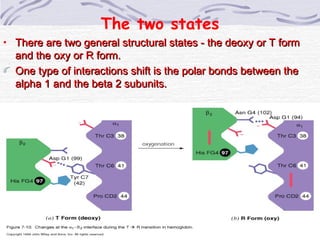
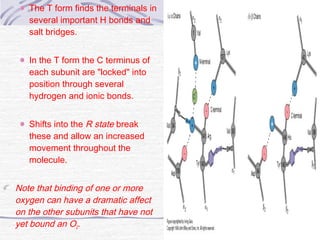
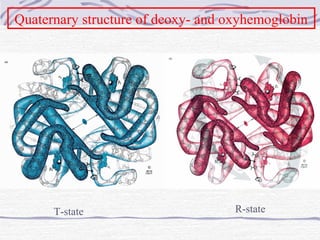


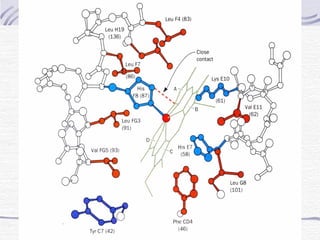



![Hemoglobin function
α2,β2 dimer which are structurally similar to myoglobin
•Transports oxygen from lungs to tissues.
•O2 diffusion alone is too poor for transport in larger
animals.
•Solubility of O2 is low in plasma i.e. 10-4
M.
•But bound to hemoglobin, [O2] = 0.01 M or that of air
•Two alternative O2 transporters are;
•Hemocyanin, a Cu containing protein.
•Hemoerythrin , a non-heme containing protein.](https://image.slidesharecdn.com/proteins-140630053132-phpapp01/85/Proteins-overview-72-320.jpg)


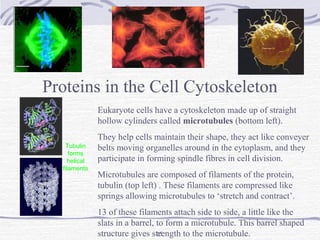
![The Bohr Effect
Higher pH i.e. lower [H+
] promotes tighter binding of
oxygen to hemoglobin
and
Lower pH i.e. higher [H+] permits the easier release of
oxygen from hemoglobin
( ) ( ) +
+
+⇔+ xHOHbOHOHb 1n22xn2
Where n = 0, 1, 2, 3 and x ≅ 0.6 A shift in the equilibrium
will influence the amount of oxygen binding. Bohr protons](https://image.slidesharecdn.com/proteins-140630053132-phpapp01/85/Proteins-overview-76-320.jpg)