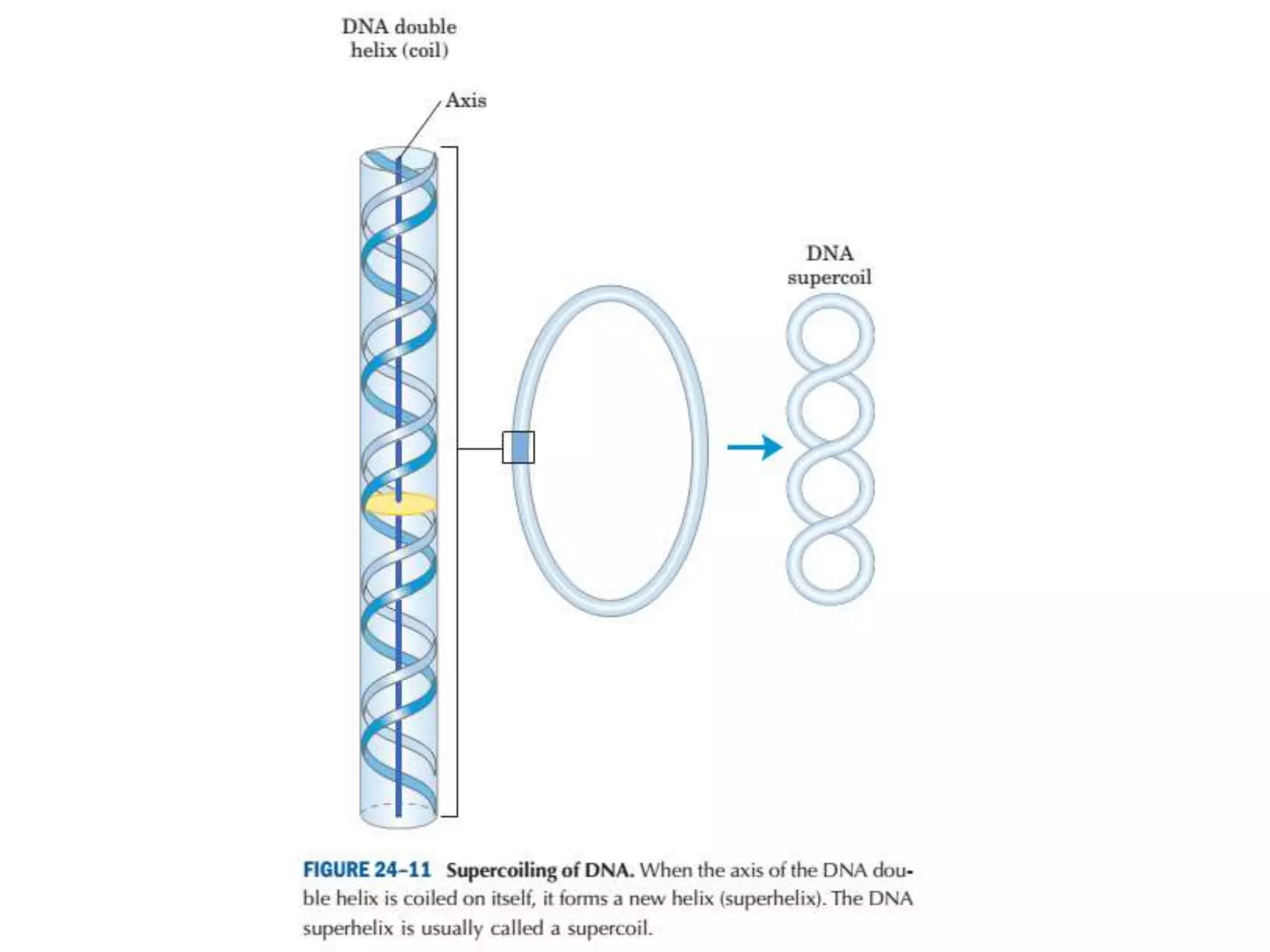
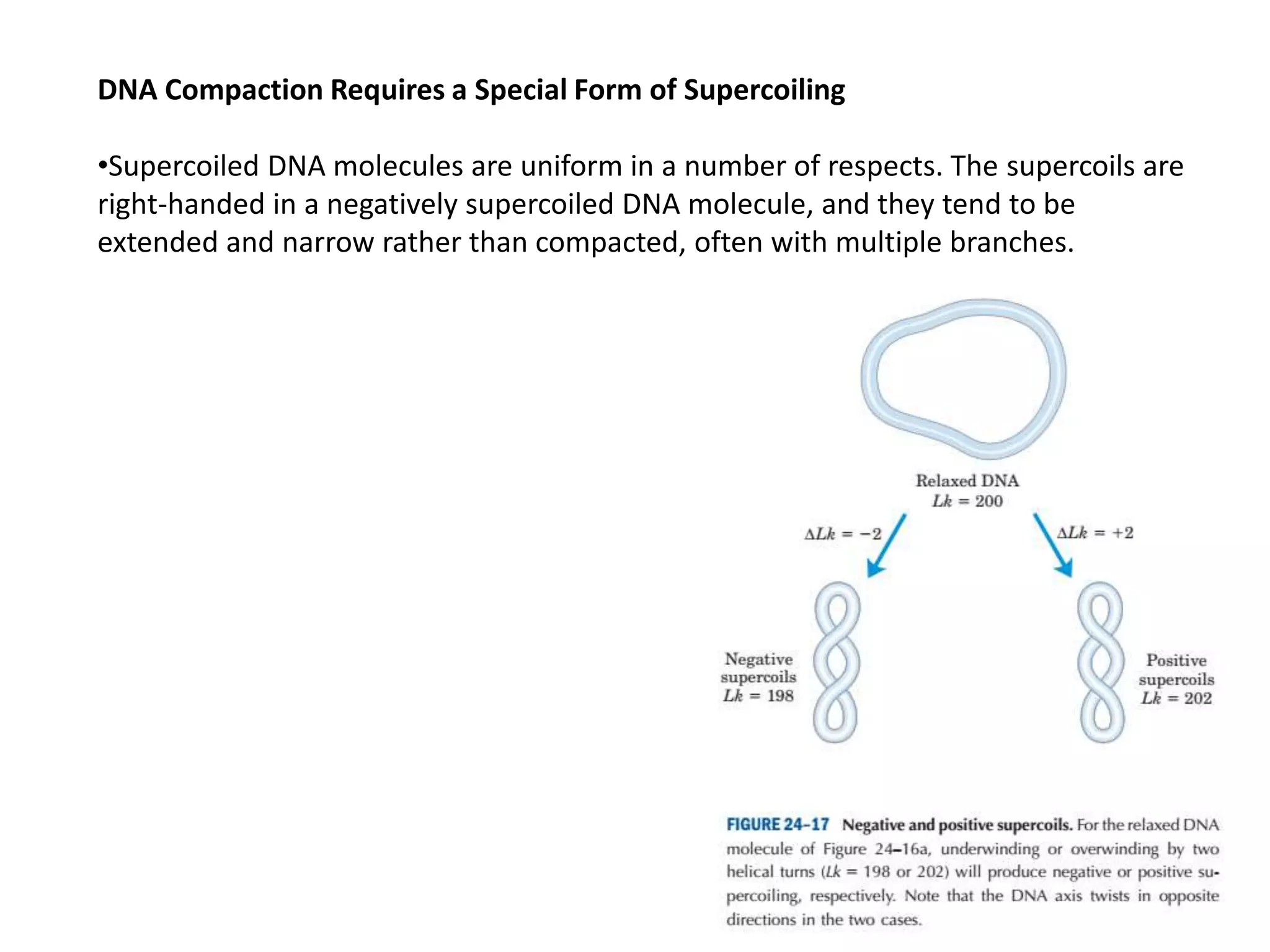
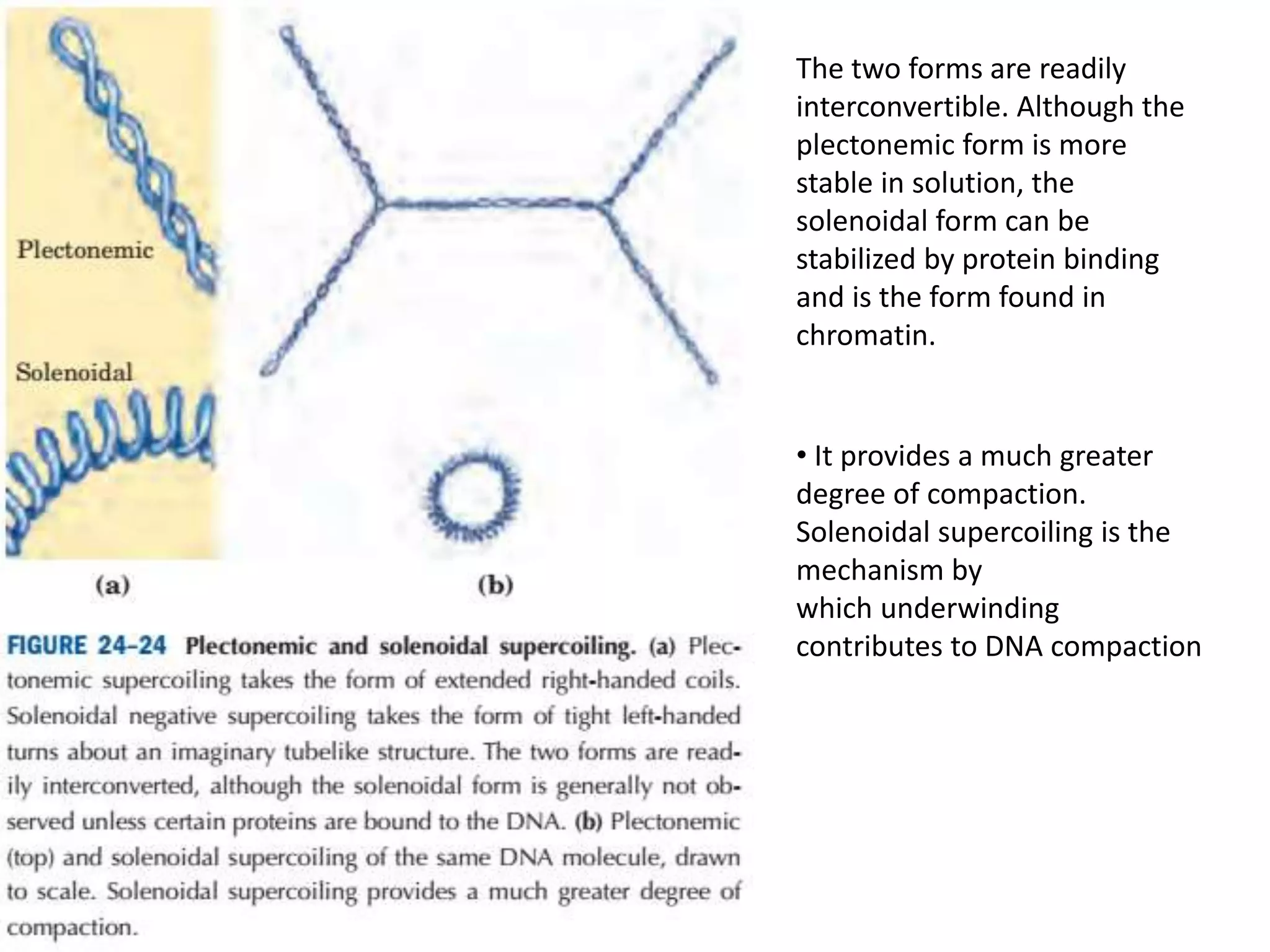
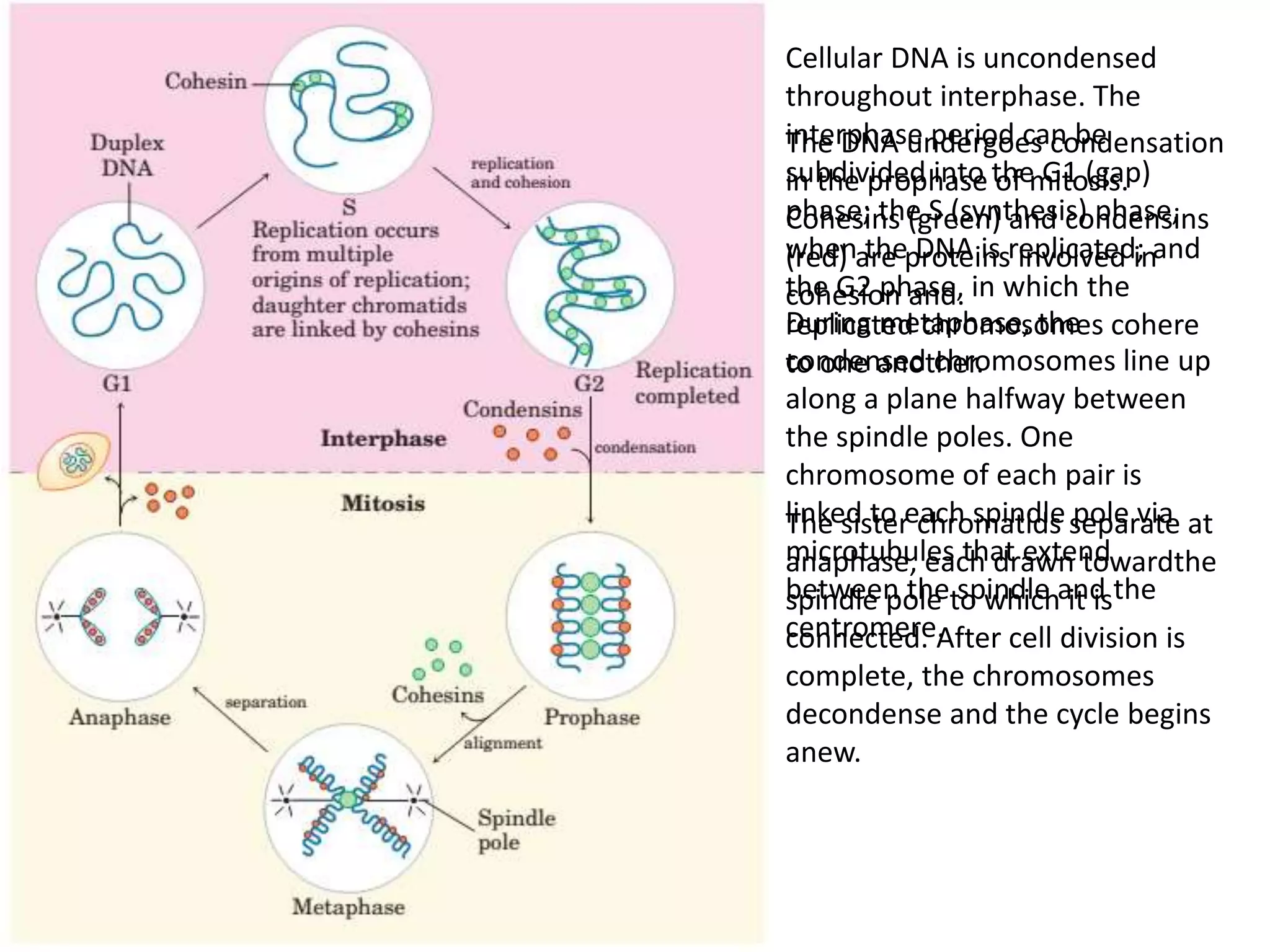
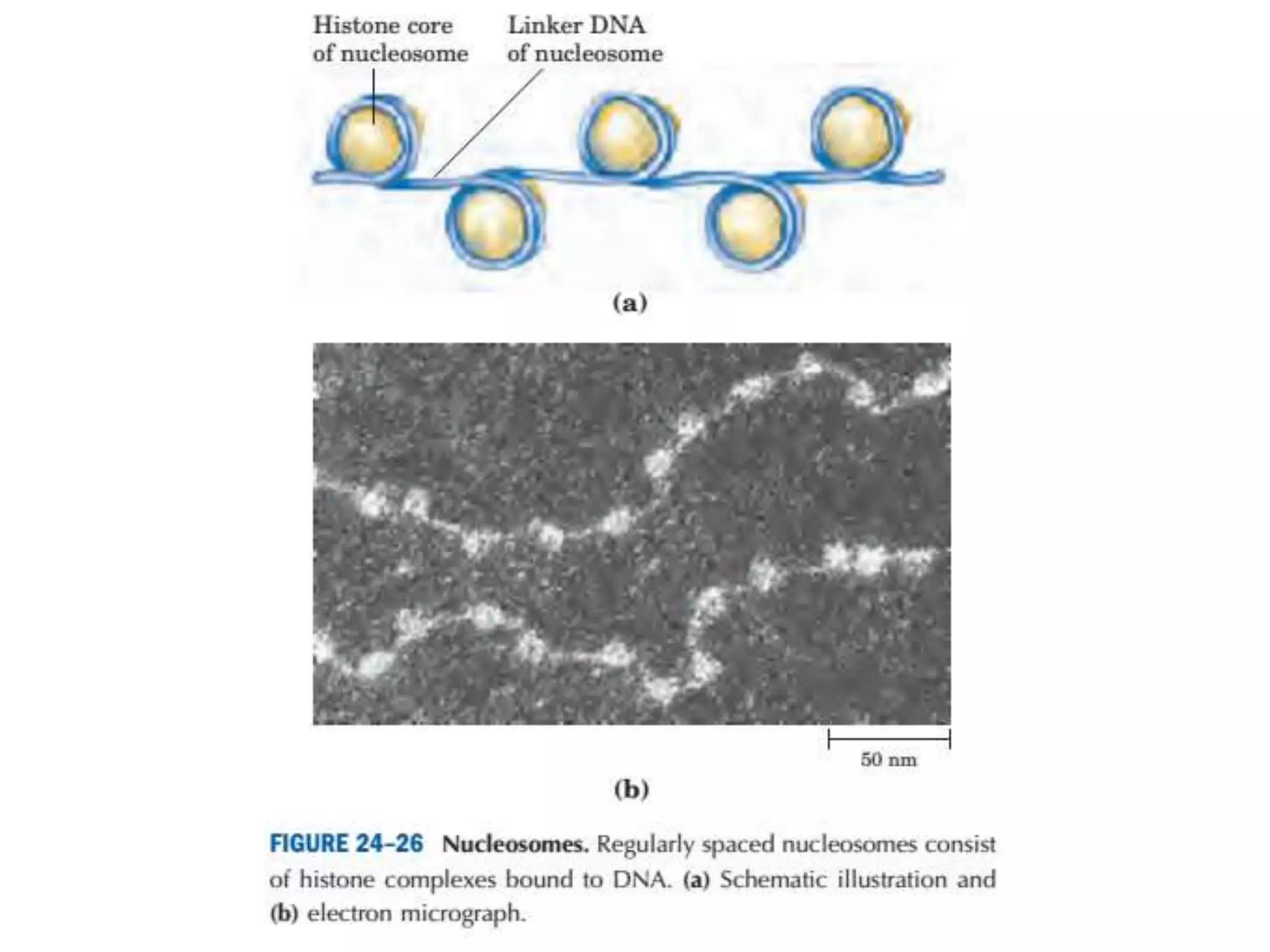
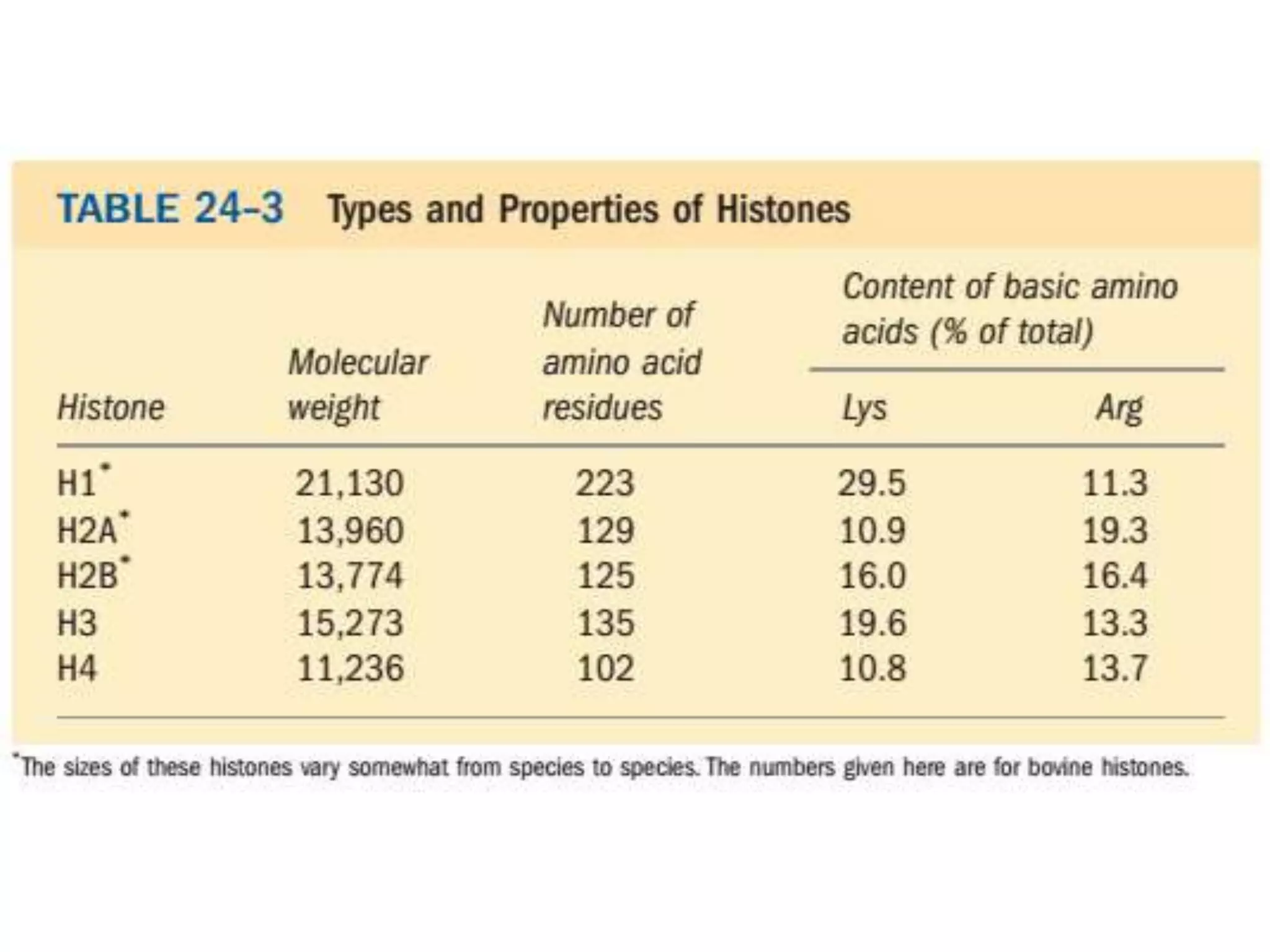
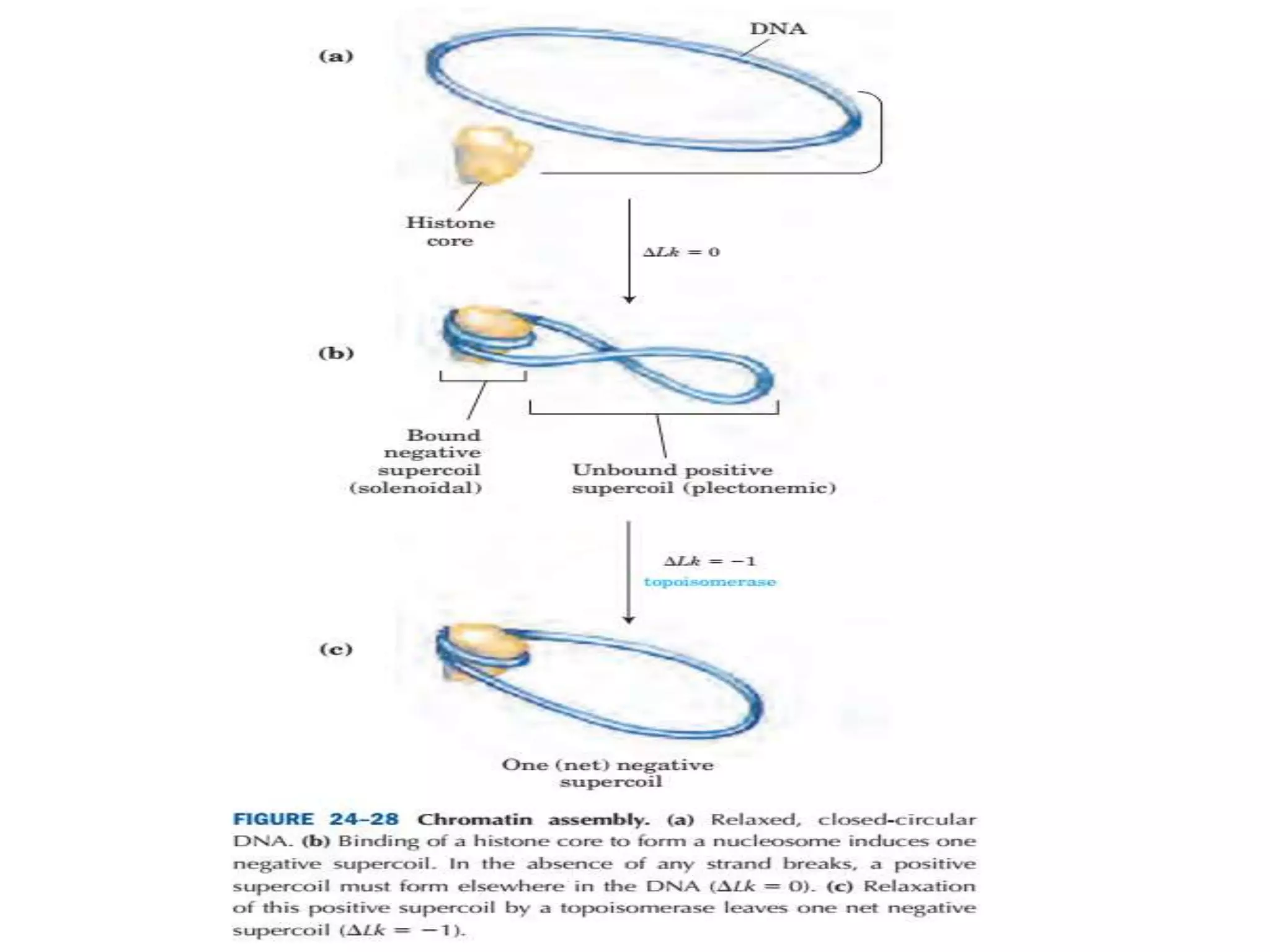
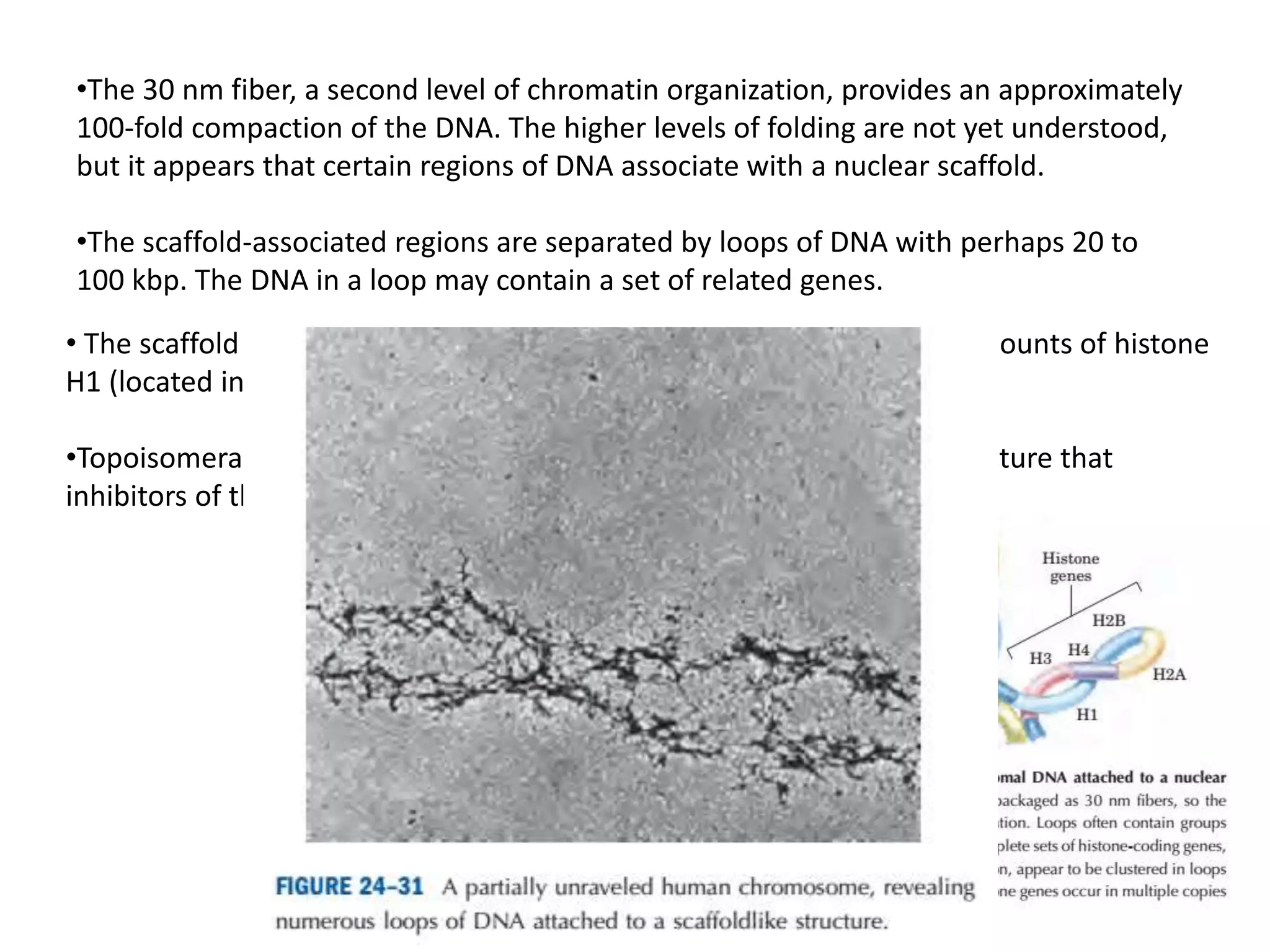
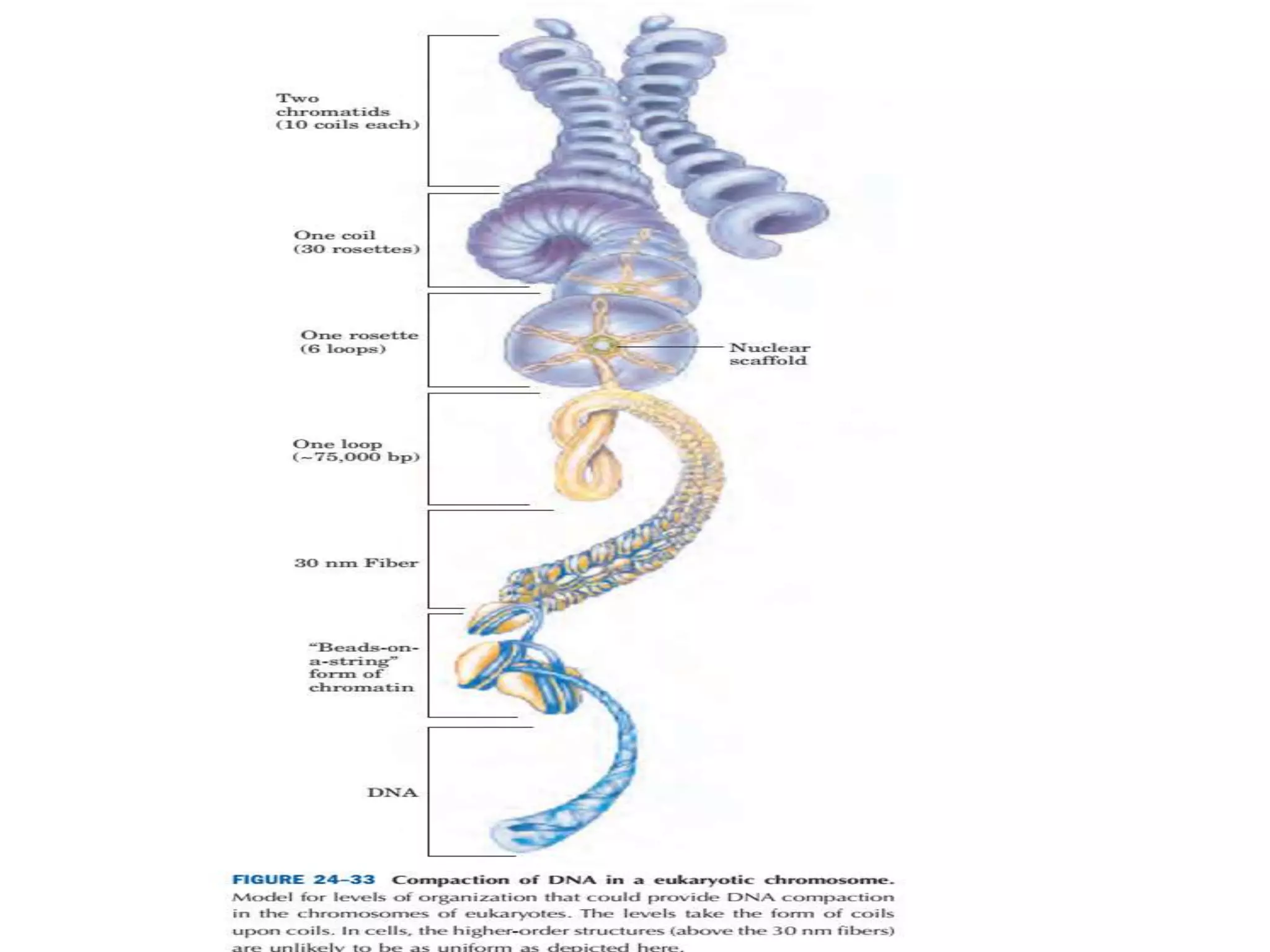
DNA is highly compacted in cells through various levels of supercoiling and chromatin structure. At the most basic level, DNA wraps around histone proteins to form nucleosomes, introducing negative supercoiling. Nucleosomes are then packed into a beads-on-a-string structure and further compacted into the 30nm fiber. Additional folding compacts the DNA over 10,000-fold into the final chromosomes. The two main types of supercoiling that facilitate compaction are plectonemic and solenoidal, with solenoidal supercoiling allowing the greatest degree of compaction and found in chromatin.