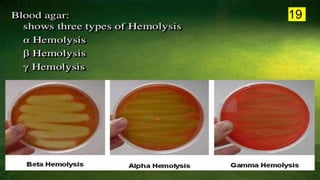
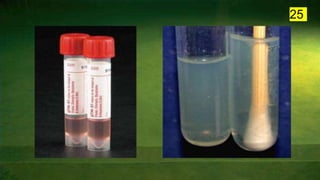
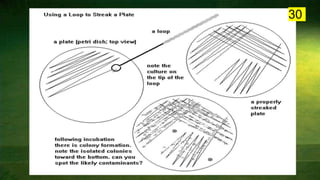
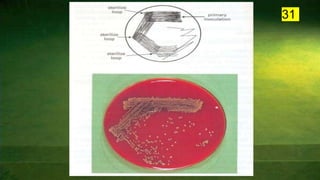
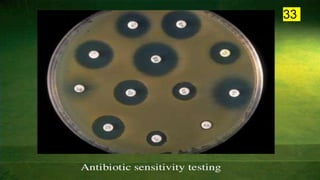
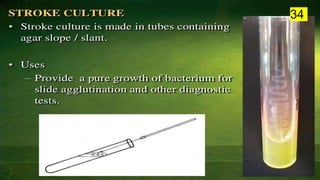
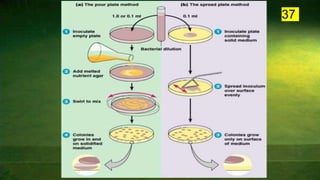
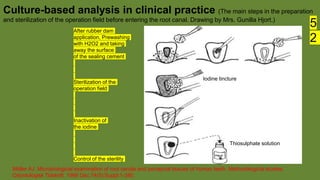
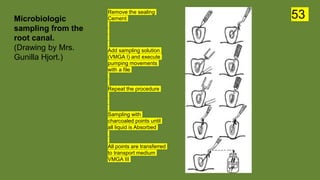
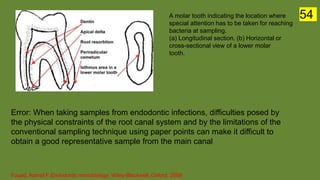
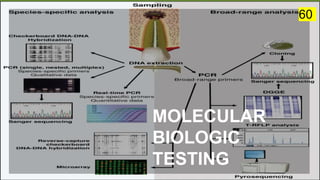
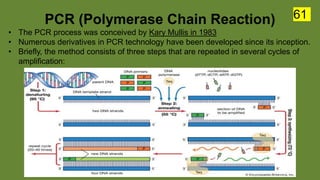
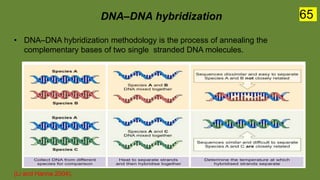
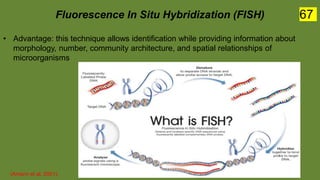
The document discusses the historical and contemporary perspectives on the microbiological aspects of endodontics. It describes some of the early culture-based techniques used to sample and identify microbes in root canals, including Grossman's culture technique from 1940. It also discusses limitations of culture-based analysis and the development of molecular biological testing methods like PCR, DNA-DNA hybridization, and fluorescence in situ hybridization that have improved identification of endodontic pathogens. Finally, it reviews recent studies on chairside culture tests and ATP bioluminescence assays that allow for rapid detection of microbes in root canals.