Bharti
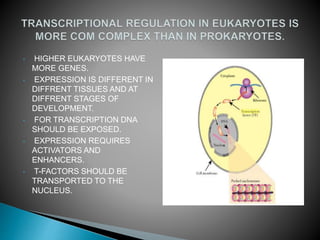
- 1. • HIGHER EUKARYOTES HAVE MORE GENES. • EXPRESSION IS DIFFERENT IN DIFFRENT TISSUES AND AT DIFFRENT STAGES OF DEVELOPMENT. • FOR TRANSCRIPTION DNA SHOULD BE EXPOSED. • EXPRESSION REQUIRES ACTIVATORS AND ENHANCERS. • T-FACTORS SHOULD BE TRANSPORTED TO THE NUCLEUS.
- 2. PROPERTIES. • RESPOND TO A SPECIFIC STIMULUS. • CAPABLE OF ENTERING THE NUCLEUS. • RECOGNISE & BIND TO SPECIFIC DNA SEQUENCE. • DIRECT/INDIRECT CONTACT WITH T-APPARATUS. • T-FACTORS HAVE TWO DOMAINS : 1ST THAT BINDS TO DNA & 2ND THAT INTERACTS WITH T-APPARATUS
- 3. Mediator: protein complex that sits on the top of RNA Pol II & provides site of contact for activators (grant RNA pol permission to proceed forward from promoter ; contact with general T-factors – TFIIB TFIID TFIIH) Mediator contains 20 protein subunits & constant core (attached to this are other subunits that vary between organisms & also between different tisues within same organism) It combines & receives signals from multiple activators/repressors & send result to RNA Pol II enzyme Some subunits of mediator act in a positive manner while others act in a negative manner. Individual mediator proteins :- ‘co-activator proteins’
- 4. Enhancers loop around DNA so that activator proteins bound at enhancer can make contact with T-apparatus via mediator complex How enhancer is prevented from activating genes further along the chromosome ?? Chromosomes are divided into regulatory neighborhoods by special sequences : ‘boundary elements’ (bcz they form boundaries to the regions of heterochromatin) or ‘INSULATORS’ Insulators are regions of DNA that bind to ‘ZINC FINGERS’ (also called insulator binding proteins ; IBP’s) & these are rich in GC regions & CG sequences may be methylated. Eg: in vertebrates, CTCF (CCCTC-binding factors) must bind to insulator to block enhancers When insulator is methylated, it dsnt bind to CTCF & no longer functions Insulators prevent enhancers from interfering with the wrong genes & also prevent the spread of heterochromatin. Insulators : fixed/mobile (gypsy element ;a retrotransposon)
- 5. During interphase,a filamentous web of proteins (nuclear matix) appears inside nuclear membrane ; DNA is attached to these proteins by sites called MARs (also called SARs i.e. scaffold attachment regions) . These protein bound MARs sites recognise the bent DNA (DNA with multiple runs of A’s is bent) MARs : 200-1000 bps long, AT-rich Next to MARs are ‘topoisomerase II recognition sites’ Chromatin remodeling also starts from MARs site Eg : in transgenic animals, the efficient expression of transgene is helped my making sure that it lies between two MAR sites bcz this region is more likely to be opened up for transcription.
- 6. Looped domains between MAR sites Alhough many loops are present but only a single loop of histone free DNA is drawn coming from a region of the nuclear scaffold. MARs contain MAR proteins that anchor DNA to the scaffold.
- 7. Negative regulators act by occupying the recognition site of activator on DNA ti prevent its binding. Eg: activation of H2B gene requires binding of activator CTF to CAAT sequence. But CDP (CAAT displacement protein) can also occupy CAAT box & prevent binding activator thereby preventing the assembly of T- apparatus but CDP dsnt block the binding site for RNA pol Eg: MyoD (T-factor needed for the formation of muscle cells & member of bHLH proteins : DNA binding proteins that bind DNA as dimers; bHLH function as heterodimers(tissue specific bHLH + widely spread E protein) By itself, MyoD dimerizes poorly. So it forms mixed dimers with E12/E47 (similar in shape & str to MyoD but expressed in all tissues unlike MyoD) . Id protein binds to MyoD & E12/E47 & inhibits differentiation.
- 8. Myod: a T-factor Interference with activators
- 9. Heterochromatin is densely packaged DNA that can’t be transcribed bcz RNA Pol can’t gain access to promoter Histones : H1,(H2A,H2B,H3 & H4: core histones) Acetylation of nulceosomes: Core histones can be acetylated (form less condensed chromatin) & degree of aceytlation affects the state of nucleosome aggregation & gene expression. Enzymes (HATs, HDACs eg: human CBP p300 proteins) bind to T-factors Chromatin remodeling complexes (CRCs): 1st they slide nucleosomes along a DNA molecule & exposing seq. for transcription and 2nd ability to rearrange histones loosening nucleosome structure allowing access to DNA 2 families of CRCs: 1) larger Swi/Snf (switch snuff) complexes 2) smaller ISWI (initiation switch) complexes
- 11. I. T-factor binds to the DNA II. HATs binds to the T-factor III. HAT acetylates the histones & association of nucleosomes is loosened IV. CRC slides & rearranges nucleosomes allowing the binding access to the DNA V. Binding of T-factors VI. Binding of RNA pol to the DNA VII. Initiation requires a positive signal to be transmitted via the mediator complex from 1 or more specific T- factors
- 12. There are 3 types of methylases: 1) Maintenance methylases: add methyl groups to newly made DNA at locations opp. Methyl groups on old parental DNA strand (ensuring that pattern of methylation is inherited during chromosome division) 2) De novo methylases: change the pattern of methylation 3) Demethylases: removes the methyl groups Methylation silences gene expression About half the genes are loacted close to CG islands. Housekeeping genes (expressed in all tissues) possess non methylated CG islands. In contrast, Cg islands of tissue specific genes are non- methylated only in those particular tissues where the genes are expressed.
- 13. Genes are silenced by methylation of DNA or covalent modifications of both DNA & histones Silencing affects a single gene/cluster of genes/substantial region of chromosome/whole chromosome Methylation of cytosine in CG/CNG sequences occurs where methyl grp project into the major groove of DNA & hinder the binding of T-factors Methylated CG seq. are recognised & bound by MeCPs recognised by other proteins that remove acetyl groups from histones condensation of DNA to form heterochromatin DNA methylation patterns are repogrammed when a new zygote is formed : just after fertilisation, methylation pattern of most of the DNA are erased leading to new patterns in a tissue specific manner Those genes whose promoter regions are methylated are silenced.
- 15. Imprinting occurs when methylation patterns from the gametes survive the formation of the zygote & affect gene expression in the new organism. A few special genes retain their methylation patterns through fertilisation. Imprinting ensures that only one of a pair of genes in a diploid cell is expressed. The second copy is silenced by methylation. (the choice as to which allele of a genes to express depends on its parental origin) 20 imprinted genes are known in mouse. Eg: IGF-II from father is expressed whereas maternal allele isn’t. Usually, it’s the paternal allele that is expressed in zygote bt nt always. Eg: of expression of paternal allele is IGF-IIR (receptor for IGF-II)
- 16. Special form of imprinting C.elegans: expression level on both X-chromosomes(f) is halved. In males, thr is no Y chromosome ie. Single unpaired X-chromosome (XO) & XX are hermaphrodites. (sdc2 & msl2) Drosophila: expression of genes on the single X-chromosome is doubled Mammals: 1of the pair of X-chromosome in each female cell is silenced ( X-inactivation by methylation of Xist gene/ non coding RNA inactivation) Worms & insects: protein complexes which dec./inc. transcription of genes located on X-chromosome