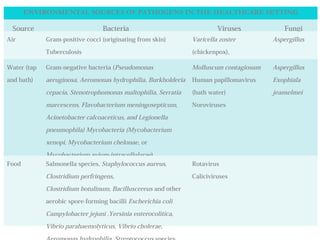
This document discusses drug resistance and nosocomial infections. It begins by describing the discovery of antibiotics by Alexander Fleming in 1928 and how antibiotics work by either killing bacteria or preventing their growth. While antibiotics were initially a "miracle cure", overuse and misuse has led to the development of drug-resistant bacteria. Resistance can arise through genetic mutations that make bacteria less susceptible to antibiotics or through horizontal gene transfer between bacteria. The document examines several antibiotic targets and mechanisms of resistance, such as beta-lactamase enzymes providing resistance to penicillins and altered cell walls conferring vancomycin resistance. It stresses the importance of properly using and prescribing antibiotics to slow the development and spread of drug-resistant bacteria.





![Sources of Antibiotics
[ v 'mekt n]ɪ ə ɪ 伊维菌素
利福霉素类
[ p li 'm ks n]ˌ ɒ ː ɪ ɪ 多粘菌素 [ bˌ æs 'tre s n]ɪ ɪ ɪ 杆菌肽素
双效菌素
[ ks z 'la d nw nz]ɒ ə ɒ ɪ ɪ ʌ 恶唑烷酮类 [la 'nez l d]ɪ ɒ ɪ 利奈唑胺](https://image.slidesharecdn.com/8-drugresistance-150727150817-lva1-app6891/85/8-drug-resistance-6-320.jpg)






![Origins of Resistance
Nearly all clinically useful antibiotics are natural products, or their
synthetic derivatives; most were isolated from other microbes
- Fungi (penicillins, cephalosporins)
- Soil bacteria of genus Streptomyces (erythromycin, streptomycin,
tetracycline, vancomycin)
In 1999, only 1 class of antibiotic was totally synthetic (Ciprofloxacin)
Antibiotics are an ancient weapon...
[d 'rev t vz]ə ə ɪ
环丙沙星](https://image.slidesharecdn.com/8-drugresistance-150727150817-lva1-app6891/85/8-drug-resistance-13-320.jpg)
![Origins of Resistance
What does the antiquity [æn't kw ti]ɪ ə 古代 of antibiotic resistance tell
us?
- There is likely to be considerable genetic variation in natural
populations for genes that can potentially confer drug resistance
(i.e., the raw genetic material is there)
- Strong selection will quickly lead to the explosive growth of
resistant individuals, especially when most cells are susceptible
Widespread antibiotic use =
- nukes their competition[nu k]ː 以核武器攻击
- the fittest survive and reproduce, passing on their resistance both
to clonal offspring and to other unrelated bacteria](https://image.slidesharecdn.com/8-drugresistance-150727150817-lva1-app6891/85/8-drug-resistance-14-320.jpg)


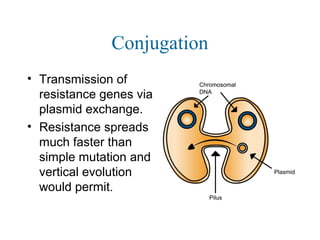
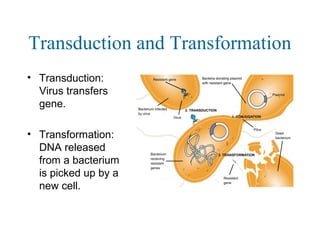

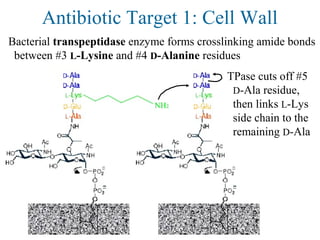
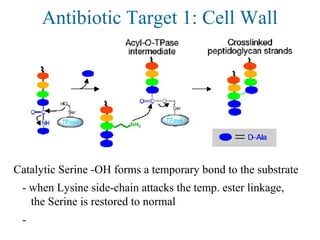
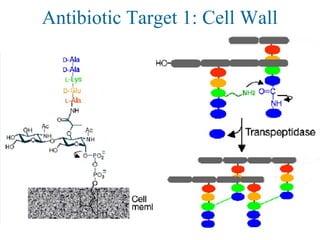
![Antibiotic Targets
The major classes of antibiotics affect 1 of 3 targets in bacteria cells:
(1) Cell wall biosynthesis
penicillins
cephalosporins
vancomycin (non-ribosomal peptide)
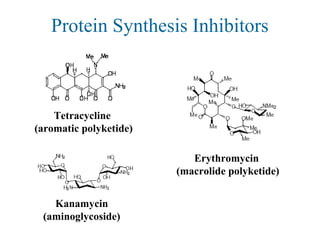
(2) Protein synthesis
erythromycin (macrolide ['mækro la d]ʊ ɪ 大环内酯物 polyketides)
tetracycline (aromatic [ˌær 'mə æt k]ɪ 芳香的 polyketides 聚酮化合物 )
streptomycin, kanamycin (aminoglycosides)
(3) DNA replication
quinolones (Cipro) 盐酸环丙沙星制剂
(β-lactams)](https://image.slidesharecdn.com/8-drugresistance-150727150817-lva1-app6891/85/8-drug-resistance-20-320.jpg)

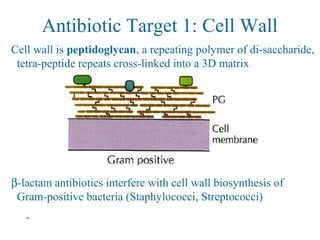
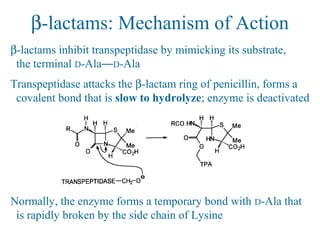
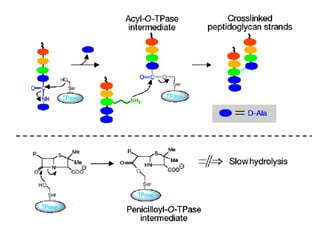
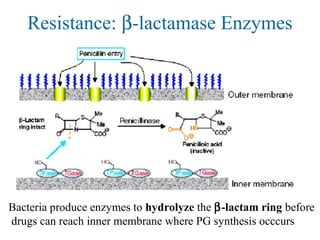
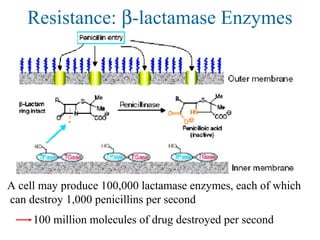
![Overcoming β-lactam Resistance
Augmentin combines β-lactam antibiotic w/ clavulanate 克拉
维酸钾 , a
“suicide” β-lactam that occupies the β-lactamase enzymes
- Allows active drug (amoxacillin, [ m ks 's l n]ə ɒ ɪ ɪ ɪ 阿莫西林 )
(resistance) slow to
hydrolyze
(cell wall enz.)](https://image.slidesharecdn.com/8-drugresistance-150727150817-lva1-app6891/85/8-drug-resistance-29-320.jpg)
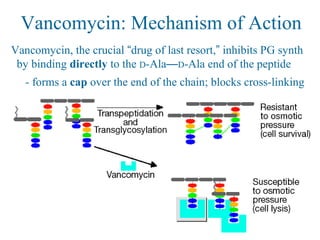
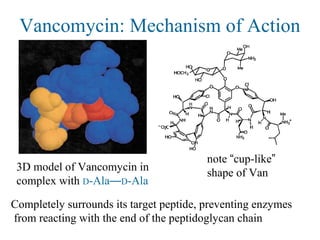
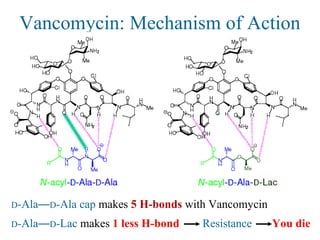

![Overcoming Van Resistance
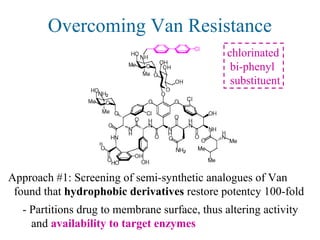
Approach #2: Screening combinatorial libraries for novel small
molecules that cleave the D-Ala—D-Lac depsipeptide
[dep'saipepta d]ɪ 缩酚酸肽
- Look for drugs that can effectively function like an enzyme
Combinatorial library of 300,000 tripeptide derivatives yielded
3 hits, all w/ an N-terminal serine & an intramolecular H-bond
Pharmacophore deduced from computer modeling studies
N
HO
NH2
O
SProC5 “resensitized” bacteria
with Van-resistance, by cleaving
their D-Ala—D-Lac depsipeptide
SProC5
Chiosis & Boneca, Science 2001](https://image.slidesharecdn.com/8-drugresistance-150727150817-lva1-app6891/85/8-drug-resistance-39-320.jpg)
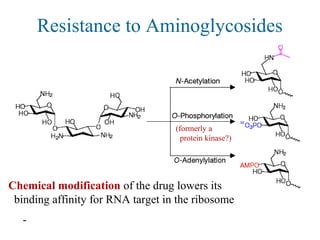

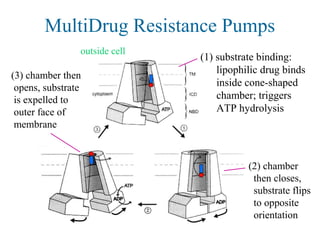
![Erythromycin Resistance
In addition to efflux pumps, erthyromycin resistance can arise
from reprogramming the target (akin[ 'k n]ə ɪ 近似的 to Van
resistance)
Methylation of a specific adenine ['æd n n]ə ɪ on the 23S rRNA
component of the ribosome
- decreases binding affinity of erythromycin-class drugs
- does not impair protein synthesis
- present as a self-immunity mechanism in erythromycin-
producing bacteria](https://image.slidesharecdn.com/8-drugresistance-150727150817-lva1-app6891/85/8-drug-resistance-44-320.jpg)
![Overcoming Erythromycin Resistance
Introduction of a 3-keto ['ki to ]ː ʊ 氧化 group into macrolide
ring of erythromycin class antibiotics alters conformation
- no induction of ribosome-methylating genes
- lower susceptibility to efflux by pumps
Erythromycin](https://image.slidesharecdn.com/8-drugresistance-150727150817-lva1-app6891/85/8-drug-resistance-45-320.jpg)
![Selection favoring Resistance
What causes the rapid occurrence of widespread resistance?
(1) Incomplete treatment: people fail to finish the full course of their
medication
- in the 1980’s, tuberculosis was almost wiped out w/ antibiotics
- in 1990’s, came back with a vengence['vend ns]ʒə 复仇 , due to
resistant strains
- 25% of previously-treated tuberculosis patients relapsed with drug
resistant strains; most had failed to complete their initial course
(2) Livestock doping['do p ]ʊ ɪŋ 服用禁药 : 50% of antibiotics used
by livestock farmers to increase yield of chicken, beef, pork
- high levels of antibiotics used in livestock result in strongly
resistant bacterial strains, which can then infect humans](https://image.slidesharecdn.com/8-drugresistance-150727150817-lva1-app6891/85/8-drug-resistance-46-320.jpg)