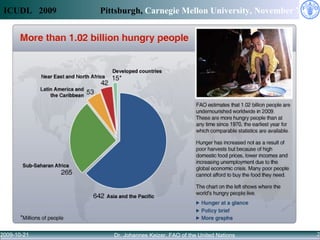
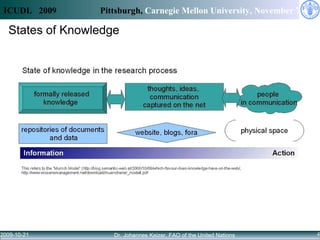
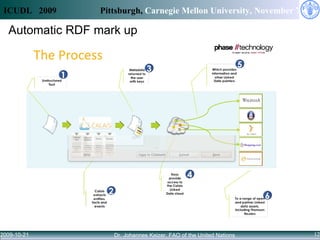
Access to knowledge is key to combating hunger and poverty, but complex information needs for agricultural development cannot be met by availability alone. Linked data principles and vocabularies can help integrate disconnected repositories and allow users to discover related information across databases. The CIARD RING aims to apply these approaches to provide value-added services that help users find information on technologies, countries, crops, and projects.