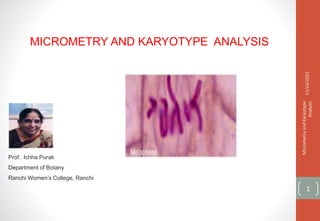
Micrometry and karyotype analysis
- 1. Prof. Ichha Purak Department of Botany Ranchi Women’s College, Ranchi 11/14/2021 Micrometry and Karyotype Analysis 1 MICROMETRY AND KARYOTYPE ANALYSIS
- 2. MICROMETRY / MICROMEASUREMENT Micrometry is measuement of Microscopic objects. For this unit of measurement used is micron (µ) m- micrometer which is =1/1000mm . In order to measure microscopic objects, micrometers are used. There are two types of micrometers:- a) Stage micrometer b) Ocular micrometer The stage micrometer is used to determine the value of one division of ocular for a certain microscope. The measurement of any object is possible with the help of ocular inserted into eyepiece of a microscope. Once the value of ocular micrometer for a microscope is determined at a given magnification (Caliberation is done), the measurement of any object (material) is possible with the help of ocular micrometer only. 11/14/2021 Micrometry and Karyotype Analysis 2
- 3. DESCRIPTION OF MICROMETERS Stage micrometer is a simple slide, in its centre a scale is present ,which has 1 mm divided into 100 divisions, therefore I division of stage micrometer is equal to 0.01mm= 10 µm Has 11/14/2021 Micrometry and Karyotype Analysis 3 10 X 40 X
- 4. The ocular micrometer is round cover slip like structure . In the center of this round glass also a scale is present, having 100 divisions. After every 10 division digits 10,20,30-100 are marked. The value of 1 division of ocular is unknown and varies from microscope to microscope and also varies in different magnifications 11/14/2021 Micrometry and Karyotype Analysis 4
- 5. PROCEDURE of Caliberation The stage micrometer is placed on the microscope, below the objective and ocular micrometer is introduced in the eye piece. By rotating the eye piece, scale of ocular micrometer is brought parallel to scale of stage micrometer. The two scales are coincided at any point. The no. of stage division between 10 ocular divisions is counted at 10x. 10 ocular division = X division of stage micrometer 1 ocular division = X/10 division of stage micrometer = X/10x .01mm=X/10 x .01 x 1000 µm (micron)= Xµm At low power 10 ocular division is equal to 15 stage divisions. So least count of the microscope at low power is 15 micron (um). In the same way least count is calculated for high power. At high power(40x) 10 ocular division coincides with only 3.5 stage divisions, so least count is 3.5 micron 11/14/2021 Micrometry and Karyotype Analysis 5
- 6. Least count calculation of provided microscope/ Caliberation 1div of stage = 0.01 mm. or 1/100 mm. 1mm= 1000 micron = .01 x 1000 =10 micron 1div of stage = 10 micron IN LOW POWER- 10 div of ocular coincides with 15 div of stage micrometer 1 div of ocular coincides with 15/10 div of stage micrometer = 15/10 x.01mm = 15/10x.01x1000µm = 15 µm IN HIGH POWER :- 10 div of ocular coincides with 3.5 div of stage micrometer 1 div of ocular coincides with 3.5/10 div of stage micrometer = 3.5/10x.01mm = 3.5/10 x .01x1000 µm = 3.5µm 11/14/2021 Micrometry and Karyotype Analysis 6
- 7. Least count calculation of provided microscope In Low Power10x,10x 15µm In high power 10x40x 3.5 µm 11/14/2021 Micrometry and Karyotype Analysis 7
- 8. For measurement of length and breadth of cells (Spirogyra,Oedogonium,Onion root tip cells),low power of microscope is used. The slide containing material is placed below objective and ocular micrometer is introduced into eyepiece. By rotating eyepiece ocular scale is brought parallel to length of cell (at least 5 readings are taken and averaged). No of ocular divisions multiplied by least account gives length of cell in Micron (µm). In the same way for measuring breadth, occular is brought parallel to breadth and number of ocular divisions are counted and multiplied with least account. 11/14/2021 Micrometry and Karyotype Analysis 8 For measurement of the chromosomes a well scattered metaphase plate is focused under the microscope. By rotating the eyepiece ocular scale is brought parallel to each chromosome and all chromosomes and their short arm and long arm are measured. To know the length in micron the number of ocular division is multiplied by least account.
- 9. Measurement of Vegetative cells of Oedogonium SN Length of Cell Breadth of Cell No of Oc divisions Average Least Count Length in micron No of Oc divisions Average LC Breadth in micron 1. 15 14 15 210 µ 3 3.4 15 51.0 µ 2 14 4 3 12 3 4 13 4 5 16 3 11/14/2021 Micrometry and Karyotype Analysis 9
- 10. B D F A-10x 10x Least account 15 µm B -10x40x Least account 3.5 µm Measurement of length and breadth of Oedogonium cells with the help of ocular micrometer introduced into eye piece of microscope C D A 11/14/2021 Micrometry and Karyotype Analysis 10
- 11. 10X 40X LC 3.5µ 11/14/2021 Micrometry and Karyotype Analysis 11 Photograph of Ocular and stage micrometer at same magnification
- 12. KARYOTYPE ANALYSIS Some features of chromosome like number, size and centromeric position of individual chromosomes are rather characteristic of a given species. The term karyotype is used for characteristic chromosome complement of an organism. Karyotype represents the chromosome constitution of a cell of an individual. It deals with Number of chromosomes, the length of chromosome, thickness of chromosome, the position of centromere, presence of secondary constriction and the size of satellite etc. of the somatic chromosome complement. 11/14/2021 Micrometry and Karyotype Analysis 12 Generally karyotype is prepared from well scattered chromosomes of mitotic metaphase plate. For this chromosomes are measured with the help of high power magnification of microscope or by camera lucida (having prism) drawing of metaphase plate made on paper & also of stage micrometer on same magnification or photo micrograph of metaphase plate and also either of stage micrometer or ocular micrometer in the same magnification are taken and then measurement is done.
- 13. Meiotic prophase and metaphase are also studied to identify homologous pairs, bivalents , chaisma formation, their termination ,recombination etc. Each organism /species has characteristic number of chromosomes Pisum sativum (Pea) 2n=14 Allium cepa (Onion ) 2n=16 Vicia faba (Baxla) 2n=12 Lens culinaris (Lentil) 2n=14 11/14/2021 Micrometry and Karyotype Analysis 13 Zygote and Somatic cells are diploid (2n) having two sets of chromosomes, which are contributed during fertilization by male and female gametes which are haploid (n) Two chromosomes in diploid set are sex chromosomes (Heterosomes) and others are autosomes Some genera and species have more than 2 sets of chromosomes showing polyploid series Triticum or wheat shows diploid count of 14,28 and 42 Triticum monococcum 2n=14 (diploid) Triticum dicoccum 2n=28 (Tetraploid) Triticum aestivum 2n=42 ( Hexaploid) Basic chromosome number X=7
- 14. The normal human karyotype contain 22 pairs of autosomal chromosomes and one pair of sex chromosomes. Normal karyotype for females contain two X chromosomes and are denoted 46, XX Normal karyotypes for males have an X and a Y chromosome and are denoted 46,XY. Any variation from the standard karyotype may lead to development of abnormalities. HUMAN KARYOTYPE Homo sapiens (human ) 2n=46 11/14/2021 Micrometry and Karyotype Analysis 14 Male XY Female XX
- 16. For karyotype analysis some of following parameters can be undertaken Total Length of individual chromosome ( TLIC ) in micron (µm ) Arm ratio(r) or index to centromeric position Centromeric Index Index to relative length of chromosome Relative Chromosome Length Thickness of chromosome Average chromosome length (ACL) Total Chromatin Length of Haploid Complement (TCL) Total Chromatin volume of haploid complement(TCV) F Percentage Total Form Percentage (TF%) Symmetry (S) percentage Karyotype category(KC)-Stebbins(1971) Karyotype formula(KF) Idiogram/ Karyogram 11/14/2021 Micrometry and Karyotype Analysis 16
- 17. Total Length of Individual Chromosome (TCIL) It is measured in micron ( µm) and includes length of both arms and centromeres (from one telomere to another telomere).The chromosomes are grouped variously on the size of the chromosome. A B C D 1)5.01-10 3.01-5.00 1.01-3.00 0.25-1.00 Large Median Small Minute 2)7.5-10.1 5.01-7.5 2.5-5.00 Less than 2.5 micron Large Median Small Minute 11/14/2021 Micrometry and Karyotype Analysis 17
- 18. Arm Ratio -It is the ratio of long arm & short arm of a chromosome. Classification of chromosome on arm ratio (r) is made as per Levan et.al(1964). Value of r(arm ratio ) can range from 1 (if SA=LA) to (limit for SA=0) Arm ratio( LA/SA) Symbol Position of centromere 1.00 M Absolute median 1.01-1.7 m Median 1.71-3.00 Sm Sub-median 3.01-7.00 St Sub-terminal More than 7.01/∞ T Terminal Arm ratio(r) or index to centromeric position 11/14/2021 Micrometry and Karyotype Analysis 18
- 19. Continued : Index to centromeric position S/L first proposed by Battaglia (1955), it is reciprocal to the arm ratio (r ) . Its values can range from 1 (if S = L) to 0 (if S = 0). It is the ratio of short arm to long arm of a chromosome, as proposed again by Sawa(1965) in Characeae Arm Ratio (S/L) Short arm/Long arm Symbol used Position of centromere Type of chromosome 1.00 m median Metacentric 0.5-0.99 sm submedian Submetacentric Less than 0.49 St Sub-terminal Acrocentric 0.00 t terminal Telocentric 11/14/2021 Micrometry and Karyotype Analysis 19
- 20. Short arm (S)= P arm Long arm (L)= Q arm Centromeric index = S/(L+S) It is the proportion of short arm in respect with the whole chromosome. Its values can range from 0.5 (if S = L) to 0 (if S = 0). Complementary to Centromeric Index=L/(L+S) it is the proportion of long arm in respect with the whole chromosome. Its values can range from 0.5 (if S = L) to 1 (if S = 0). 11/14/2021 Micrometry and Karyotype Analysis 20
- 21. Index to relative length Types of chromosomes and the symbols Less than 0.75 Small (s) 0.75-1.00 Medium small (M1) 1.01-1.25 Medium large (M2) More than 1.25 Large(L) Index to relative length of chromosome It is ratio of length of each chromosome to mean length of complement. Relative Chromosome Length (RCL) -It is ratio in percentage between total length of individual chromosome and total chromatin length of haploid complement RCL= TLIC/TCL x100 11/14/2021 Micrometry and Karyotype Analysis 21
- 22. Thickness Of Chromosome Chromosomes are classified in to 3 categories on the basis of thickness of chromosome TYPE RANGE OF THICKNESS Slender 0.1 to less than 1 micron Moderately Thick 1.00 micron Thick more than 1.0 micron 11/14/2021 Micrometry and Karyotype Analysis 22
- 23. Average Chromosome Length (ACL )- Average chromosome length is the ratio between total chromatin length of haploid complement and total number of chromosomes in haploid set. Total Chromatin Length (TCL)- TCL is sum total of length of all chromosomes of a complement (haploid set) (CL) Total Chromatin Volume (TCV)- Total chromatin volume of haploid complement is sum total of chromatin volume. (CV=πr2l) F Percentage- is the ratio in percentage of short arm length to entire chromosome length F% = Short arm length of chromosome / Length of chromosomes x100 . 11/14/2021 Micrometry and Karyotype Analysis 23
- 24. Symmetry (S) percentage – It is ratio in percentage of length of the shortest chromosome to length of longest chromosome of the complement and is calculated as follows S%=G I=Gradient Index Total Form Percentage (TF%)- It is a ratio in percentage of total sum of short arm length to total chromatin length of the complement as advocated by Huziwara (1962) TF% = Total length of short arm in a chromosome set /Total length of chromosome set x 100 11/14/2021 Micrometry and Karyotype Analysis 24
- 25. Karyotype Category (KC) - A karyotype exhibiting large differences between smallest and largest chromosome and fewer metacentric chromosomes is asymmetric, which is believed to be an advance feature over symmetric karyotype ,in which less difference between largest and smallest chromosome and more chromosomes are with metacentric centromere Karyotypes have been classified into 12 categories (1A- 4C) as per Stebbins(1971) taking into account both the degree of difference between the largest(LC) and Smallest(SC) chromosome of complement and position of centromere on chromosome He established these by recognizing three degrees of difference (A-C) between the largest and smallest chromosome of the complement and four degrees (1-4) with respect to the proportion of chromosomes which are metacentric with an arm ratio of less than 2:1 11/14/2021 Micrometry and Karyotype Analysis 25
- 26. . KARYOTYPE CATEGORIES AS PER STEBBINS(1971) LC/SC P=No of chromosomes with sm centromere /No of total chromosomes Proportion of chromosome with arm ratio< 2:1 0.00 0.01-0.50 0.51-0.99 1.00 <2:1 1A(most primitive) 2A 3A 4A 2:1-4:1 1B 2B 3B 4B >4:1 1C 2C 3C 4C(most advanced) LC= longest chromosome SC= Smallest chromosome 11/14/2021 Micrometry and Karyotype Analysis 26
- 27. The Karyotype of 1A category of Stebbins (1971) are most primitive having less difference between largest and smallest chromosome and less number of chromosomes with asymmetric centromeric position ( Sm,St,t) and more with M & m centromeric position (Levan et. al 1964) where as 4C category of Stebbins(1971) is most advanced having large difference between largest and shortest chromosome and more number of chromosomes with Sm,St and t centromeric position. Karyotype asymmetry- The concept of karyotype symmetry Vs asymmetry was developed by Levitzky(1931) The information about karyotype asymmetry has been known to be significant in plant taxa where association occur between increasing asymmetry of karyotype and other characteristics such as chromosome number, plant morphology or plant habit. 11/14/2021 Micrometry and Karyotype Analysis 27 According to Stebbins (1971) , increasing asymmetry occurs through pericentric inversion, changing median centromeric position to subterminal one A B C D Eᵒ F G H I J Median A B C D H G Fᵒ E I J Sub-terminal
- 28. Karyological features of Lens culinaris L-4149 n=7,2n=14 Chr.no Long Arm(LA) (micron) Short Arm(SA) (micron) Satelite Total Length of Chromosome In micron LA/SA Chr. Category on the basis of Centromeric position size 1 4.25 2.55 - 6.80 1.67 m B 2 4.25 1.70 - 5.95 2.50 sm B 3 3.00 1.70 - 5.10 2.0 Sm B 4 2.55 1.70 - 4.25 1.5 m C 5 2.55 1.70 - 4.25 1.5 m C 6 2.55 1.70 - 4.25 1.5 m C 7 2.55 0.85 - 3.40 3.00 sm C 11/14/2021 Micrometry and Karyotype Analysis 28 Karyotypic Formula (KF):- It illustrates types of chromosomes on size and index of centromeric position of the chromosome compliment. Karyotypic Formula = (1m2sm)B + (3m1sm)C
- 29. The chromosome complement reveals that chromosome length ranges from 3.40 µm (SC) to 6.80 µm (LC) Karyotype reveals 4 pairs with median and 3 pairs with submedian centromere Total chromatin length (TCL) of diploid set is 68 .0µm & of haploid set = 34.0 µm Average chromosome length – 4.86 µm LC/SC= 6.80/3.40= 2.0 P Ratio- No of chromosomes with Sm centromere/total no of chromosomes = 3/7= 0.42 Karyotype Category as per Stebbins (1971) - 2B 11/14/2021 Micrometry and Karyotype Analysis 29
- 30. Idiogram/ Karyogram:- It can be presented graphically by arranging the chromosome in descending order of length in a straight line. The longest chromosome is placed on right side and smallest on left side. All chromosomes in a complement (Karyotype) do not bear the centromere at the same position while in graphical representation either the position of centromere is fixed at a point or are located on different points and one telomere of chromosome is fixed . Sat chromosomes are represented by a small break on the arm which belongs it and a scale is taken ( 1cm=1µm ) so a chromosome having 4 µm length will cover 4 cm (4 squares of graph paper) and if it has median centromere then 2cm above and 2cm below ,the gap represents the centromere. 11/14/2021 Micrometry and Karyotype Analysis 30
- 31. Metaphase plate Outline drawing Idiogram (haploid set) n=7 Position of centromere is fixed 2m Karyological features of Lens culinaris L-4149,2n=14 11/14/2021 Micrometry and Karyotype Analysis 31 Ideogram (haploid set) n=9 Position of one telomere is fixed 3m
- 32. Pisum sativum (Pea) n=7,2n=14 Vicia faba ( Baxla) 2n=12 11/14/2021 Micrometry and Karyotype Analysis 32 Idiogram Haploid set n=6
- 33. A- Photomicrograph of metaphase plate(n=56) B- Outline drawing of metaphase plate A B Karyological features of Chara fibrosa var.fibrosa f. tylacantha(Nordst.) R D W (Purak and Noor ,2018) C- Karyogram showing 56 chromosomes KARYOTYPIC FORMULA ( 8M 6m) B + ( 10 M 22m 4sm 4 t) C + (2 t) D C 11/14/2021 Micrometry and Karyotype Analysis 33
- 34. Metaphase Allium cepa 2n=16 Metaphase Vicia faba 2n=12 11/14/2021 Micrometry and Karyotype Analysis 34 Some Metaphase Plates
- 35. CVCL (Paszko,2006) –is a ratio between standard deviation and mean of a sample x 100, ranges from 0 (no variation ) to 100 ( a measure of interchromosomal asymmetry) Coefficient of variation of chromosome length = CV CL = (SCL/XCL) X 100 =A2 X 100 MCA (Mean Centromeric Asymmetry), where Centromeric Asymmetry of a single chromosome is given by the formula (L-S)/(L+S).Accordingly, MCA = A × 100 = Mean (L-S)/(L+S) X100= A X 100 (a measure of intrachromosomal asymmetry ) Peruzzi and Eroglu (2013) Coefficient of variation of the Centromeric index = CVCI = ( SCI/XCI) X 100 (a measure of centromere position heterogeneity in the karyotype ) precisely assesses the relative variation in centromere position in a complement. Paszko (2006) These 3 parameters have the potential to display even minor karyotypic variations. KARYOTYPE ASYMMETRY : 3 ADDITIONAL PARAMETERS After Stebbins (1971) formulation of 12 categories of karyotype, some additional parameters were later on added related with Karyotype asymmetry 11/14/2021 Micrometry and Karyotype Analysis 35
- 36. Battaglia E (1955) Chromosome morphology and terminology. Caryologia 8: 179–187 Huziwara Y (1962) Karyotype analysis in some genera of Compositae. VIII. Further studies on the chromosome of Aster. American Journal of Botany 49: 116–119. Levan A, Fredga K, Sandberg AA (1964) Nomenclature for centromeric position on chromosomes. Hereditas 52: 201–220. Levitsky GA (1931) The karyotype in systematics . Bulletin of Applied Botany,Genetics and Plant Breeding 27: 220-240 Paszko A (2006) A critical review and a new proposal of karyotype asymmetry indices. PlantSystematics and Evolution 258: 39–48. Peruzzi L and Eroglu H E ( 2013) Karyotype asymmetry : again how to measure and what to measure ? CompCytogen 7(1) ; 1-9 Sawa T (1965) - Cytotaxonomy of the Characeae: karyotype analysis of Nitella opaca and Nitella flexilis. Amer. J. Bot., 52 (9): 962-970 Stebbins GL (1971) Chromosomal evolution in higher plants. London, UK: Edward Arnold (Publishers) Ltd. Watanabe K, Yahara T, Denda T, Kosuge K (1999) Chromosomal evolution in the genus Brachyscome (Asteraceae, Astereae): Statistical tests regarding correlation between changes in karyotype and habit using phylogenetic information. Journal of Plant Research 112:145–161. Zarco R ( 1986) A new method of estimating karyotype asymmetry . Taxon 35: 526-530 REFERENCES 11/14/2021 Micrometry and Karyotype Analysis 36
- 37. Some computer aided programs (SOFTWARES ) have been developed from time to time for measurement of chromosomes (short and long arm ) with the help of images uploaded on computer 1.Micromeasure can be downloaded from http://www.biology.colostate.edu/MicroMeasure 2.KaryoType can be downloaded from (http://mnh.scu.edu.cn/soft/blog/karyotype/) 3.IdeoKar: an ideogram constructing and karyotype analyzing software This software and related documentation is freely available on the website of the University of Kurdistan at: http://agri.uok.ac.ir/ideokar/index.html. 11/14/2021 Micrometry and Karyotype Analysis 37 REFERENCES Aaron Reeves (2001) MicroMeasure: A new computer program for the collection and analysis of cytogenetic data.Genome 44: 439–443 Altinordu F et. al. (2016) A tool for the analysis of chromosomes: KaryoType TAXON : 7 pp Ghader Mirzaghaderi & Karim Marzangi (2015) IdeoKar: an ideogram constructing and karyotype analyzing software, Caryologia, 68:1, 31-35
