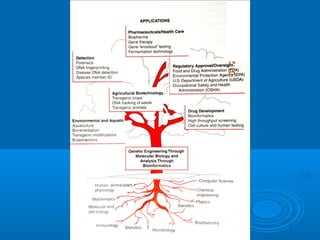
Tecnologías con DNA
- 2. • Artificial selectionArtificial selection
- 6. Recombinant DNA TechnologyRecombinant DNA Technology • Isolates and amplifiesIsolates and amplifies • specific sequences of DNAspecific sequences of DNA • incorporates them intoincorporates them into vectorvector DNA moleculesDNA molecules • ResultingResulting recombinant DNArecombinant DNA • is propagated and amplifiedis propagated and amplified (cloned)(cloned) • in organisms such asin organisms such as E. coliE. coli
- 7. Recombinant DNA VectorsRecombinant DNA Vectors • Naturally occurring circular bacteria DNANaturally occurring circular bacteria DNA moleculesmolecules (plasmids)(plasmids) • Bacterial virusesBacterial viruses (bacteriophages)(bacteriophages)
- 8. Genetically Altered BacteriaGenetically Altered Bacteria • Produce important human proteinProduce important human protein productsproducts • insulininsulin • growth hormonegrowth hormone • tissue plasminogen activator (TPA)tissue plasminogen activator (TPA) • tissue growth factor-beta (TGF-tissue growth factor-beta (TGF- ββ)) • clotting factor VIIIclotting factor VIII • Dornase Alpha (DNase)Dornase Alpha (DNase)
- 11. Fig. 15-2, p. 325 Plasmid from a bacterium DNA of interest from another organism 1 Plasmid and DNA from another organism are cut by the same restriction enzyme (in this example, Hin dIII). This produces molecules with complementary single- stranded ends. Clonable DNA fragment 2 Mix two types of molecules so their sticky ends pair. DNA ligase then forms covalent bonds at junctions, linking fragments. Recombinant DNA 3 Transfer recombinant DNA molecule to host cell, where it is copied and turned on to produce gene product.
- 12. PlasmidsPlasmids
- 19. Gen reportero Gen de interés cDNA
- 24. Fig. 15-15, p. 340
- 25. Fig. 15-16, p. 341
- 26. Fig. 15-17b, p. 342
- 27. Genetically Engineered PlantGenetically Engineered Plant
- 28. Libraries (1)Libraries (1) • Genomic DNA libraryGenomic DNA library • thousands of DNA fragmentsthousands of DNA fragments • all DNA of an organismall DNA of an organism • Chromosome libraryChromosome library • all DNA fragments of a specific chromosomeall DNA fragments of a specific chromosome
- 30. cDNA LibrarycDNA Library • Complementary DNA (cDNA)Complementary DNA (cDNA) • produced usingproduced using reverse transcriptasereverse transcriptase • makes DNA copies of eukaryotic mRNAmakes DNA copies of eukaryotic mRNA • Copies are incorporated into recombinantCopies are incorporated into recombinant DNADNA vectorsvectors
- 31. cDNAcDNA
- 32. Genetic ProbeGenetic Probe • Radioactive DNA or RNA sequenceRadioactive DNA or RNA sequence • used to screen recombinant DNA moleculesused to screen recombinant DNA molecules in bacterial cellsin bacterial cells • to find specific colony with DNA of interestto find specific colony with DNA of interest
- 34. Animation: Use of a RadioactiveAnimation: Use of a Radioactive ProbeProbe CLICK TO PLAY
- 35. Polymerase Chain Reaction (PCR)Polymerase Chain Reaction (PCR) • AutomatedAutomated in vitroin vitro techniquetechnique • targets a particular DNA sequence by specifictargets a particular DNA sequence by specific primersprimers • clones it using heat-resistant DNAclones it using heat-resistant DNA polymerasepolymerase • Used to analyze tiny DNA samplesUsed to analyze tiny DNA samples • from crime scenes, archaeological remainsfrom crime scenes, archaeological remains
- 37. Fig. 15-7, p. 329
- 39. Fig. 15-8a, p. 330
- 40. Fig. 15-8a, p. 330 DNA Cut with restriction enzyme 100 base pairs 200 base pairs 300 base pairs Mixture placed in well Standards of known size + – Origin Direction of movement 200 base pairs 300 base pairs 100 base pairs Gel
- 41. Fig. 15-8b, p. 330
- 43. RNA and ProteinsRNA and Proteins • Northern BlotNorthern Blot • RNA moleculesRNA molecules separated by electrophoresisseparated by electrophoresis • transferred to a membranetransferred to a membrane • Western BlotWestern Blot • Proteins or polypeptidesProteins or polypeptides previously separatedpreviously separated by gel electrophoresisby gel electrophoresis
- 44. DNA Microarrays (2)DNA Microarrays (2) • Cancer and other diseases exhibit alteredCancer and other diseases exhibit altered patterns of gene expressionpatterns of gene expression • DNA microarraysDNA microarrays identify disease-causingidentify disease-causing genes (or the proteins they code for)genes (or the proteins they code for)
- 46. Fig. 15-13, p. 336 1 Prepare microarray. Each microdot contains multiple copies of a specific single-stranded cDNA. Treated cell Untreated (control) cell 2 Prepare cDNA from two cell populations (treated and control). Mature mRNA Mature mRNAReverse transcriptase Reverse transcriptase cDNA copy of mRNA cDNA copy of mRNA 3 Tag each cDNA with different fluorescent dye. cDNA mRNA (discard) cDNA mRNA (discard)
- 47. Fig. 15-13, p. 336 4 Hybridize two cDNA populations to array. Laser 1 Scan array to identify fluorescence where hybridization has occurred. 5 Laser 2 Gene that was inactive in both treated and untreated cells Emissions 6 Computer analysis produces color-coded readout. Gene in treated cell that increased activity, compared to control Gene in treated cell that decreased activity, compared to control Gene that was active in both treated and untreated cells
- 48. DNA SequencingDNA Sequencing • Yields information about gene structureYields information about gene structure • and amino acid sequence of encoded proteinsand amino acid sequence of encoded proteins • Geneticists compare DNA sequencesGeneticists compare DNA sequences • with other sequences stored in databaseswith other sequences stored in databases
- 50. Chain Termination MethodChain Termination Method
- 51. Fig. 15-11a, p. 334 Single-strand DNA fragment to be sequenced +ddATP +ddCTP +ddGTP +ddTTP
- 52. Fig. 15-11b, p. 334 Radioactive primer Direction of synthesis+ddATP Reaction products from mixture containing dideoxyATP
- 53. Fig. 15-11c, p. 334 Larger fragments Smaller fragments
- 54. Fig. 15-11d, p. 334 CA TG
- 55. Automated DNA SequenceAutomated DNA Sequence CLICK TO PLAY CLICK TO PLAY
- 56. DNA FingerprintingDNA Fingerprinting • Analysis of individual’s DNAAnalysis of individual’s DNA • based onbased on short tandem repeats (STRs)short tandem repeats (STRs) (molecular markers, highly(molecular markers, highly polymorphicpolymorphic)) • Applications inApplications in • law enforcementlaw enforcement • disputed parentagedisputed parentage • tracking tainted foodstracking tainted foods
- 57. Animation: DNA FingerprintingAnimation: DNA Fingerprinting CLICK TO PLAY
- 58. Fig. 15-14, p. 339 1 2 3 4 5 6 7From blood at crime scene
- 59. Transgenic OrganismsTransgenic Organisms • Foreign DNAForeign DNA • incorporated into genetic materialincorporated into genetic material • Transgenic livestockTransgenic livestock • produce foreign proteins in milkproduce foreign proteins in milk • Transgenic plantsTransgenic plants • have great potential in agriculturehave great potential in agriculture
Editor's Notes
- Figure 23.1 Basic steps in the generation of a recombinant DNA molecule.
- Figure 15.2: Splicing foreign DNA into a vector. In this example, a bacterial plasmid is the vector.
- Figure 23.3 Plasmid vector pBR322. This plasmid is a small circular DNA of 4361 base pairs. It contains an origin of replication (ori) and genes conferring resistance to the antibiotics ampicillin (ampR) and tetracycline (tetR). The rop gene (repressor of primer) regulates DNA replication so that there are about 20 copies of the plasmid per bacterial cell. The plasmid also contains several unique restriction sites (only a few common ones are shown).
- Figure 23.2 Use of restriction enzymes to generate recombinant DNA. The vector DNA and the target DNA are cleaved by restriction endonucleases to generate ends that can be joined together. In cases where sticky ends are produced, the two molecules join by annealing (base pairing) of the complementary ends. The molecules are then covalently attached to one another in a reaction catalyzed by DNA ligase.
- Figure 23.5 Yeast artificial chromosome (YAC). This shuttle vector contains yeast centromeric DNA (CEN4) and a yeast origin of replication (ARS1). The plasmid also contains two marker genes for selection in yeast cells (TRP1 and URA3) and a marker gene for selection in E. coli (ampR). Large fragments of DNA (400 to 500 kb) can be inserted at the unique EcoRI site. Telomeric sequences are arranged in the plasmid so that cleavage with the restriction enzyme BamHI removes a fragment of DNA, leaving a linear vector with telomeric ends. Following cleavage of the recombinant DNA molecule by the action of BamHI, the linear artificial chromosome is used to transform yeast cells.
- Figure 23.1 Basic steps in the generation of a recombinant DNA molecule.
- Figure 23.1 Basic steps in the generation of a recombinant DNA molecule.
- Figure 23.6 Yeast shuttle vector that can be propagated and selected in both Escherichia coli and Saccharomyces cerevisiae. Recombinants are selected in E. coli by the ability to grow in the presence of antibiotic, and in LEU2-deficient yeast strains by the ability to grow in the absence of exogenous leucine.
- Figure 23.11 Expression of a eukaryotic protein in E. coli.
- Figure 23.4 Preparation and use of phage vector. Genomic DNA is partially digested with EcoRI (a partial digest leaves some EcoRI sites intact). The resulting fragments are separated by electrophoresis in an agarose gel. (The fragments, which are in the right-hand lane, appear as a smear due to the range of their sizes.) DNA molecules of known size (size markers) are used to locate appropriately sized fragments, which are recovered from the gel and ligated to the arms of vector DNA. The vector DNA is prepared by digesting it with EcoRI, yielding the arms, each of which contains a cos site at one end. Since arms are assembled into phages only if ligated to insert DNA of an appropriate size, recombinant phages are selected.
- Figure 23.12 Technique for creating a transgenic mouse.
- Figure 15.15: A transgenic mouse. The mouse on the right is normal, whereas the mouse on the left is a transgenic animal that expresses rat growth hormone.
- Figure 15.16: Transgenic “pharm” cows. These cows contain a human gene that codes for lactoferrin, a protein found in human mothers’ milk and in secretions such as tears, saliva, bile, and pancreatic fluids. Lactoferrin is one of the immune system’s lines of defense against disease-causing organisms. The cows secrete human lactoferrin in their milk.
- Figure 15.17: Uses of transgenic plants.
- Figure 23.17 Three cycles of the polymerase chain reaction. The sequence to be amplified is shown in blue. (1) The duplex DNA is melted by heating and cooled in the presence of a large excess of two primers (red and yellow) that flank the region of interest. (2) A heat-stable DNA polymerase catalyzes extension of these primers, copying each DNA strand. Successive cycles of heating and cooling in the presence of the primers allow the desired sequence to be repeatedly copied until, after 20 to 30 cycles, it represents most of the DNA in the reaction mixture.
- Figure 15.7: Amplification of DNA by PCR. The initial reaction mixture includes a very small amount of double-stranded DNA, DNA precursors (deoxyribonucleotides), specific nucleic acid primers, and heat-resistant Taq DNA polymerase.
- Figure 15.8: Gel electrophoresis.
- Figure 15.8: Gel electrophoresis.
- Figure 15.8: Gel electrophoresis.
- Figure 15.13: A DNA microarray.
- Figure 15.13: A DNA microarray.
- Figure 15.11: The chain termination method of DNA sequencing.
- Figure 15.11: The chain termination method of DNA sequencing.
- Figure 15.11: The chain termination method of DNA sequencing.
- Figure 15.11: The chain termination method of DNA sequencing.
- Figure 15.14: DNA fingerprinting. These DNA fingerprints show DNA from a crime scene (in middle), along with the DNA profiles of seven suspects. Can you pick the suspect whose DNA profile matches blood from the crime scene?