Genomic evaluation of low-heritability traits: dairy cattle health as a model
•Download as PPTX, PDF•
1 like•524 views
Genetic selection has been very successful when applied to traits of moderate to high heritability, but progress has been slow for traits with low heritabilities. The problem is further compounded when novel traits are considered because data needed to calculate high-reliability PTA generally are not available. A combination of producer-recorded health event data and SNP genotypes may permit the routine calculation of PTA with reasonable reliabilities for health traits. Poster presented at the 5th International Symposium on Animal Functional Genetics in Guaruja, Brazil, in 2014.
Report
Share
Report
Share
Recommended
5-MethylcytosineDNA Methylation Patterns among Gut Predominate Commensal Escherichia coli and Lactobacilli from the Balbas and Mazekh Domestic Sheep Breeds by Pepoyan AZ* in Research in Medical &Engineering SciencesCrimson publishers-5-MethylcytosineDNA Methylation Patterns among Gut Predomi...

Crimson publishers-5-MethylcytosineDNA Methylation Patterns among Gut Predomi...CrimsonpublishersMedical
Recommended
5-MethylcytosineDNA Methylation Patterns among Gut Predominate Commensal Escherichia coli and Lactobacilli from the Balbas and Mazekh Domestic Sheep Breeds by Pepoyan AZ* in Research in Medical &Engineering SciencesCrimson publishers-5-MethylcytosineDNA Methylation Patterns among Gut Predomi...

Crimson publishers-5-MethylcytosineDNA Methylation Patterns among Gut Predomi...CrimsonpublishersMedical
High-grade (WHO grade III and IV) astrocytomas are aggressive malignant brain tumors affecting humans with a high risk of recurrence in both children and adults. To date, limited information is available on the genetic and molecular alterations important in the onset and progression of pediatric high-grade astrocytomas and, even less, on the prognostic factors that influence long-term outcome in children with recurrence. A-to-I RNA editing is an essential post-transcriptional mechanism that can alter the nucleotide sequence of several RNAs and is
mediated by the ADAR enzymes. ADAR2 editing activity is particularly important in mammalian brain and is impaired in both adult and pediatric high-grade astrocytomas.
Moreover, we have recently shown that the recovered ADAR2 activity in high-grade astrocytomas inhibits in vivo tumor growth. The aim of the present study is to investigate whether changes may occur in ADAR2-mediated RNA editing profiles of relapsed highgrade astrocytomas compared to their respective specimens collected at diagnosis, in four pediatric patients.ADAR2 editing activity in newly diagnosed versus relapsed pediatric high-grad...

ADAR2 editing activity in newly diagnosed versus relapsed pediatric high-grad...Enrique Moreno Gonzalez
Glucose-6-phosphate dehydrogenase (G6PD), elevated in tumor cells, catalyzes the first reaction in the pentose-phosphate pathway. The regulation mechanism of G6PD and pathological change in human melanoma growth remains unknown. Variant G6PD levels promote tumor cell proliferation or apoptosis via the STA...

Variant G6PD levels promote tumor cell proliferation or apoptosis via the STA...Enrique Moreno Gonzalez
Lapatinib is characterized as an ErbB1/ErbB2 dual inhibitor and has recently been approved for the treatment of metastatic breast cancer. In this study, we examined mechanisms
associated with enhancing the activity of lapatinib via combination with other therapies. OSU-03012 sensitizes breast cancers to lapatinib-induced cell killing: a role...

OSU-03012 sensitizes breast cancers to lapatinib-induced cell killing: a role...Enrique Moreno Gonzalez
More Related Content
What's hot
High-grade (WHO grade III and IV) astrocytomas are aggressive malignant brain tumors affecting humans with a high risk of recurrence in both children and adults. To date, limited information is available on the genetic and molecular alterations important in the onset and progression of pediatric high-grade astrocytomas and, even less, on the prognostic factors that influence long-term outcome in children with recurrence. A-to-I RNA editing is an essential post-transcriptional mechanism that can alter the nucleotide sequence of several RNAs and is
mediated by the ADAR enzymes. ADAR2 editing activity is particularly important in mammalian brain and is impaired in both adult and pediatric high-grade astrocytomas.
Moreover, we have recently shown that the recovered ADAR2 activity in high-grade astrocytomas inhibits in vivo tumor growth. The aim of the present study is to investigate whether changes may occur in ADAR2-mediated RNA editing profiles of relapsed highgrade astrocytomas compared to their respective specimens collected at diagnosis, in four pediatric patients.ADAR2 editing activity in newly diagnosed versus relapsed pediatric high-grad...

ADAR2 editing activity in newly diagnosed versus relapsed pediatric high-grad...Enrique Moreno Gonzalez
Glucose-6-phosphate dehydrogenase (G6PD), elevated in tumor cells, catalyzes the first reaction in the pentose-phosphate pathway. The regulation mechanism of G6PD and pathological change in human melanoma growth remains unknown. Variant G6PD levels promote tumor cell proliferation or apoptosis via the STA...

Variant G6PD levels promote tumor cell proliferation or apoptosis via the STA...Enrique Moreno Gonzalez
Lapatinib is characterized as an ErbB1/ErbB2 dual inhibitor and has recently been approved for the treatment of metastatic breast cancer. In this study, we examined mechanisms
associated with enhancing the activity of lapatinib via combination with other therapies. OSU-03012 sensitizes breast cancers to lapatinib-induced cell killing: a role...

OSU-03012 sensitizes breast cancers to lapatinib-induced cell killing: a role...Enrique Moreno Gonzalez
What's hot (20)
Beyond GWAS QTL Identification and Strategies to Increase Yield

Beyond GWAS QTL Identification and Strategies to Increase Yield
Rare Genetic Mutations Contribute to Alopecia Areata Etiology

Rare Genetic Mutations Contribute to Alopecia Areata Etiology
Heritability estimates of, genetic and phenotypic correlations among some sel...

Heritability estimates of, genetic and phenotypic correlations among some sel...
ADAR2 editing activity in newly diagnosed versus relapsed pediatric high-grad...

ADAR2 editing activity in newly diagnosed versus relapsed pediatric high-grad...
Candidate causative mutation on BTA18 associated with calving and conformatio...

Candidate causative mutation on BTA18 associated with calving and conformatio...
Variant G6PD levels promote tumor cell proliferation or apoptosis via the STA...

Variant G6PD levels promote tumor cell proliferation or apoptosis via the STA...
OSU-03012 sensitizes breast cancers to lapatinib-induced cell killing: a role...

OSU-03012 sensitizes breast cancers to lapatinib-induced cell killing: a role...
Induction of HSP16.2 in C. elgans Using Caffeine as Stressor 

Induction of HSP16.2 in C. elgans Using Caffeine as Stressor
Data as research output, data as part of the scholarly record

Data as research output, data as part of the scholarly record
Viewers also liked
Poultry Breeding Poultry Breeding and Production manual by Dr.Shamoil Tariq

Poultry Breeding and Production manual by Dr.Shamoil TariqUniversity of Veterinary & Animal Sciences,Lahore
Viewers also liked (7)
Genetic variability, heritability, genetic advance, genetic advance as percen...

Genetic variability, heritability, genetic advance, genetic advance as percen...
Poultry Breeding and Production manual by Dr.Shamoil Tariq

Poultry Breeding and Production manual by Dr.Shamoil Tariq
Lecture 3 quantitative traits and heritability full

Lecture 3 quantitative traits and heritability full
Similar to Genomic evaluation of low-heritability traits: dairy cattle health as a model
Glioblastoma (GBM) is the most malignant brain tumor. Many abnormal secretion and
expression of cytokines have been found in GBM, initially speculated that the occurrence of
GBM may be involved in these abnormal secretion of cytokines. This study aims to detect the
association of cytokine genes with GBM.Genetic association between selected cytokine genes and glioblastoma in the H...

Genetic association between selected cytokine genes and glioblastoma in the H...Enrique Moreno Gonzalez
The use of gene expression data from Micro arrays coupled with WT QTL's linked to Tryp resistance phenotypes in Cattle to elucidate pertinent genetic changes underpinning phenotype in putative candidate genes A systematic, data driven approach to the combined analysis of microarray and...

A systematic, data driven approach to the combined analysis of microarray and...Laurence Dawkins-Hall
Elucidating changes in gene expression by Micro array genomic sweeps of genetic QTLs linked to Tryp resistance in WT cattle to identify putative candidates underpinning pathophysiology Systemic analysis of data combined from genetic qtl's and gene expression dat...

Systemic analysis of data combined from genetic qtl's and gene expression dat...Laurence Dawkins-Hall
Similar to Genomic evaluation of low-heritability traits: dairy cattle health as a model (20)
indrasen-chauhan-central-sheep-wool-research-institute-india.pptx

indrasen-chauhan-central-sheep-wool-research-institute-india.pptx
Genetic association between selected cytokine genes and glioblastoma in the H...

Genetic association between selected cytokine genes and glioblastoma in the H...
A systematic, data driven approach to the combined analysis of microarray and...

A systematic, data driven approach to the combined analysis of microarray and...
Genomic selection, prediction models, GEBV values, genomic selection in plant...

Genomic selection, prediction models, GEBV values, genomic selection in plant...
Systemic analysis of data combined from genetic qtl's and gene expression dat...

Systemic analysis of data combined from genetic qtl's and gene expression dat...
Genetic predisposition to papillary thyroid cancer by Albert de la Chapelle, ...

Genetic predisposition to papillary thyroid cancer by Albert de la Chapelle, ...
Statistical analysis of biological data (comparison of proportions)

Statistical analysis of biological data (comparison of proportions)
Establishment and analysis of a disease risk prediction model for chronic kid...

Establishment and analysis of a disease risk prediction model for chronic kid...
More from John B. Cole, Ph.D.
More from John B. Cole, Ph.D. (20)
Using genotypes to construct phenotypes for dairy cattle breeding programs an...

Using genotypes to construct phenotypes for dairy cattle breeding programs an...
If we would see further than others: research & technology today and tomorrow

If we would see further than others: research & technology today and tomorrow
Using genotyping and whole-genome sequencing to identify causal variants asso...

Using genotyping and whole-genome sequencing to identify causal variants asso...
The hunt for a functional mutation affecting conformation and calving traits ...

The hunt for a functional mutation affecting conformation and calving traits ...
An updated version of lifetime net merit incorporating additional fertility t...

An updated version of lifetime net merit incorporating additional fertility t...
An updated version of lifetime net merit incorporating additional fertility t...

An updated version of lifetime net merit incorporating additional fertility t...
Genetic Evaluation of Stillbirth in US Holsteins Using a Sire-maternal Grands...

Genetic Evaluation of Stillbirth in US Holsteins Using a Sire-maternal Grands...
Opportunities for genetic improvement of health and fitness traits

Opportunities for genetic improvement of health and fitness traits
Genomic selection and systems biology – lessons from dairy cattle breeding

Genomic selection and systems biology – lessons from dairy cattle breeding
Use of NGS to identify the causal variant associated with a complex phenotype

Use of NGS to identify the causal variant associated with a complex phenotype
Uso e valore economico dei test genomici in azienda

Uso e valore economico dei test genomici in azienda
The use and economic value of genomic testing for calves on dairy farms

The use and economic value of genomic testing for calves on dairy farms
New applications of genomic technology in the US dairy industry

New applications of genomic technology in the US dairy industry
Recently uploaded
Recently uploaded (20)
❤Jammu Kashmir Call Girls 8617697112 Personal Whatsapp Number 💦✅.

❤Jammu Kashmir Call Girls 8617697112 Personal Whatsapp Number 💦✅.
High Profile 🔝 8250077686 📞 Call Girls Service in GTB Nagar🍑

High Profile 🔝 8250077686 📞 Call Girls Service in GTB Nagar🍑
Vip profile Call Girls In Lonavala 9748763073 For Genuine Sex Service At Just...

Vip profile Call Girls In Lonavala 9748763073 For Genuine Sex Service At Just...
dkNET Webinar "Texera: A Scalable Cloud Computing Platform for Sharing Data a...

dkNET Webinar "Texera: A Scalable Cloud Computing Platform for Sharing Data a...
Locating and isolating a gene, FISH, GISH, Chromosome walking and jumping, te...

Locating and isolating a gene, FISH, GISH, Chromosome walking and jumping, te...
Seismic Method Estimate velocity from seismic data.pptx

Seismic Method Estimate velocity from seismic data.pptx
Biogenic Sulfur Gases as Biosignatures on Temperate Sub-Neptune Waterworlds

Biogenic Sulfur Gases as Biosignatures on Temperate Sub-Neptune Waterworlds
Module for Grade 9 for Asynchronous/Distance learning

Module for Grade 9 for Asynchronous/Distance learning
Asymmetry in the atmosphere of the ultra-hot Jupiter WASP-76 b

Asymmetry in the atmosphere of the ultra-hot Jupiter WASP-76 b
9654467111 Call Girls In Raj Nagar Delhi Short 1500 Night 6000

9654467111 Call Girls In Raj Nagar Delhi Short 1500 Night 6000
Genomic evaluation of low-heritability traits: dairy cattle health as a model
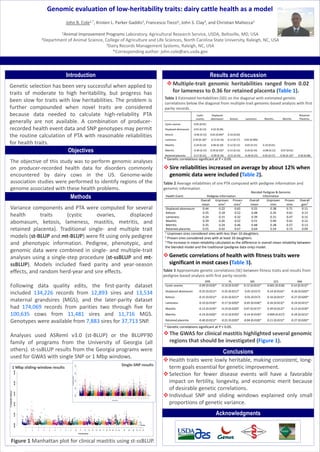
- 1. TEMPLATE DESIGN © 2008 www.PosterPresentations.com John B. Cole1,*, Kristen L. Parker Gaddis2, Francesco Tiezzi2, John S. Clay3, and Christian Maltecca2 1Animal Improvement Programs Laboratory, Agricultural Research Service, USDA, Beltsville, MD, USA 2Department of Animal Science, College of Agriculture and Life Sciences, North Carolina State University, Raleigh, NC, USA 3Dairy Records Management Systems, Raleigh, NC, USA *Corresponding author: john.cole@ars.usda.gov Introduction Genetic selection has been very successful when applied to traits of moderate to high heritability, but progress has been slow for traits with low heritabilities. The problem is further compounded when novel traits are considered because data needed to calculate high-reliability PTA generally are not available. A combination of producer- recorded health event data and SNP genotypes may permit the routine calculation of PTA with reasonable reliabilities for health traits. Results and discussion Multiple-trait genomic heritabilities ranged from 0.02 for lameness to 0.36 for retained placenta (Table 1). Methods Genomic evaluation of low-heritability traits: dairy cattle health as a model Objectives The objective of this study was to perform genomic analyses on producer-recorded health data for disorders commonly encountered by dairy cows in the US. Genome-wide association studies were performed to identify regions of the genome associated with these health problems. Variance components and PTA were computed for several health traits (cystic ovaries, displaced abomasum, ketosis, lameness, mastitis, metritis, and retained placenta). Traditional single- and multiple trait models (st-BLUP and mt-BLUP) were fit using only pedigree and phenotypic information. Pedigree, phenotypic, and genomic data were combined in single- and multiple-trait analyses using a single-step procedure (st-ssBLUP and mt- ssBLUP). Models included fixed parity and year-season effects, and random herd-year and sire effects. Following data quality edits, the first-parity dataset included 134,226 records from 12,893 sires and 13,534 maternal grandsires (MGS), and the later-parity dataset had 174,069 records from parities two through five for 100,635 cows from 11,481 sires and 11,716 MGS. Genotypes were available from 7,883 sires for 37,713 SNP. Analyses used ASReml v3.0 (st-BLUP) or the BLUPF90 family of programs from the University of Georgia (all others). st-ssBLUP results from the Georgia programs were used for GWAS with single SNP or 1 Mbp windows. Acknowledgments Conclusions Health traits were lowly heritable, making consistent, long- term goals essential for genetic improvement. Selection for fewer disease events will have a favorable impact on fertility, longevity, and economic merit because of desirable genetic correlations. Individual SNP and sliding windows explained only small proportions of genetic variance. Figure 1 Manhattan plot for clinical mastitis using st-ssBLUP. Single-SNP results 1 Mbp sliding-window results Health Event Pedigree Information Blended Pedigree & Genomic Information Overall mean Unproven sires 1 Proven sires 2 Overall mean Unproven sires Proven sires Overall gain 3 Displaced abomasum 0.44 0.22 0.65 0.55 0.38 0.71 0.11 Ketosis 0.35 0.18 0.52 0.48 0.35 0.61 0.13 Lameness 0.24 0.15 0.32 0.39 0.31 0.47 0.15 Mastitis 0.39 0.26 0.52 0.51 0.40 0.61 0.12 Metritis 0.35 0.24 0.46 0.48 0.38 0.57 0.13 Retained placenta 0.55 0.42 0.67 0.64 0.54 0.73 0.09 1 1 Unproven sires considered sires with less than 10 daughters. 2 Proven sires considered sires with at least 10 daughters. 3 The increase in mean reliability calculated as the difference in overall mean reliability between the blended model and the traditional (pedigree data only) model. Table 2 Average reliabilities of sire PTA computed with pedigree information and genomic information. Table 1 Estimated heritabilities (SD) on the diagonal with estimated genetic correlations below the diagonal from multiple-trait genomic-based analysis with first parity records. * Genetic correlations significant at P < 0.05. Sire reliabilities increased on average by about 12% when genomic data were included (Table 2). The GWAS for clinical mastitis highlighted several genomic regions that should be investigated (Figure 1). Genetic correlations of health with fitness traits were significant in most cases (Table 3). * Genetic correlations significant at P < 0.05. Table 3 Approximate genetic correlations (SE) between fitness traits and results from pedigree-based analysis with first parity records .
