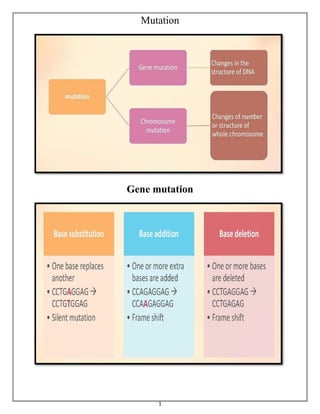
Gene mutation & Chromosomal Mutations
- 2. 2 ◘ Point Mutations: Point mutations are changes in one base pair of a cell's DNA sequence. For example, if an A in the DNA code is changed to a C, that is a point mutation. Point mutations in the coding region of a gene can have different effects depending on the resulting changes to the codons in the messenger RNA. There are a few major kinds of point mutations: missense mutations, nonsense mutations, silent mutations and read through mutations. Here, we're going to focus on nonsense mutations, which are mutations that introduce a premature stop codon into the coding sequence of a gene. ◘ Nonsense Mutation Definition When you think about a mutant, you might think about sci-fi movies where mutated creatures become powerful and evil and then attempt to destroy the world. But what are mutations, really? Mutations are changes to a cell's DNA sequence, and there are several different types. A nonsense mutation is a point mutation that introduces a premature stop codon into the part of the gene that encodes a protein.
- 3. 3 A stop codon is like a period at the end of a sentence. It instructs the ribosome to stop making the protein. So, if a mutation leads to an early stop codon, only part of the protein will be made. Half-baked proteins that result from nonsense mutations are often nonfunctional or defective. Now let's learn more about how nonsense mutations work and their consequences. Gene vs. Chromosomal Mutations Gene Mutations Chromosomal Mutations • Mistakes that affect individual genes on a chromosome. • One base substitutes for another on a DNA strand and leads to the wrong protein being made; this affects one or more functions within the organism. • Mistakes that affect the whole chromosome. • There are four types of chromosomal mutations: duplication, deletion, inversion, and translocation. • ALL MUTATIONS ARE CAUSED BY MUTAGENS. ♣ A chromosome abnormality, disorder, anomaly, aberration, or mutation is a missing, extra, or irregular portion of chromosomal DNA.
- 4. 4
- 5. 5
- 6. 6 ◘ Aberration: Whether chromosome aberrations (after the mitosis) are induced by single-strand breaks or double-strand breaks in the structure determines the fate of the cell. In single-strand breaks, the chromosome tends to repair by joining the two fragments in a process called restitution, provided sufficient time is allowed. The cell becomes functionally normal and replicates normally. However, if the fragments are replicated during DNA synthesis prior to restitution, two strands with centromeres and two strands without centromeres will be produced. Random combination of these fragments will then produce acentric and dicentric chromatids. Such chromosomes suffer severe consequences due to the mismatch of genetic information.
- 7. 7 In another scenario, radiation can cause two breaks in one arm of a chromosome, resulting in three fragments: Only two of which combine with the loss of the third. Such a process is called deletion. Combination of all three fragments into a chromosome with changes along the broken line. This process is called inversion, ◘ If radiation produces single-strand break in two separate chromosomes, then there are four ways of recombining the broken ends: A. The dicentric and acentric combinations are similar to those formed after replication of single strands in the same chromosome. B. Translocation is a process in which two fragments • One with a centromere from one chromosome and one without a centromere from another chromosome combine to form a new chromosome.
- 8. 8 ◘ Philadelphia chromosome or Philadelphia translocation • Is a specific chromosomal abnormality that is associated with chronic myelogenous leukemia (CML). • It is the result of a reciprocal translocation between chromosome 9 and 22, and is specifically designated t(9;22)(q34;q11). • The presence of this translocation is a highly sensitive test for CML, since 95% of people with CML have this abnormality.
- 9. 9 • However, the presence of the Philadelphia (Ph) chromosome is not sufficiently specific to diagnose CML, since it is also found in acute lymphoblastic leukemia and occasionally in acute myelogenous leukemia (AML). • The result of the translocation is the oncogenic BCR-ABL gene fusion, located on the shorter derivative 22 chromosome. • This translocation is designated t (9;22). • It results in one chromosome 9 longer than normal and one chromosome 22 shorter than normal. • The latter is called the Philadelphia chromosome and designated Ph1 . • The DNA removed from chromosome 9 contains most of the proto- oncogene designated c-ABL. • The break in chromosome 22 occurs in the middle of a gene designated BCR. • The resulting Philadelphia chromosome has the 5' section of BCR fused with most of c-ABL. Genetic make up
- 10. 10 ◘ Genes playing role in cancer development: • Oncogenes. • Tumor suppressor genes. • DNA repair genes. ◘ Viruses and Cancer Those are directly oncogenic, infecting and persisting in the cancer cells, e.g. most DNA tumor viruses. Those that act indirectly, promoting conditions that allow tumor cells to progress to cancer, • e.g. hepatitis C virus and Human Immunodeficiency Virus (HIV).
- 11. 11 ♣ Retroviruses: • Comprise a group of related viruses in which the viral genetic information is converted from RNA to a DNA form during infection of animal cells. • In its DNA form the viral information inserted into the host chromosomes. • From a chromosomal position, the viral DNA directs the synthesis of viral RNA and proteins, which then spontaneously assemble to form progeny virus. • Viruses of this type have been detected in many animal species, and some have been popularly called RNA tumor viruses because of their association with tumors and leukemias. ♠ Retroviruses are thought to cause tumors in two general ways: ♣ First, viral integration can occur near or within an important control gene in the host chromosome, raising the level of expression of that gene in a way that leads to uncontrolled cellular growth. • In general, the probability of this happening is low because viral integration is not specifically targeted for these sites.
- 12. 12 ♣ Second, the virus itself may carry a gene whose product disrupts normal cellular control pathways. • These virally borne ‘control genes’ are called oncogene, and they tend to produce tumor cells at a high frequency. • Viral oncogenes are often defective copies of cellular regulatory genes. • Oncogenic viruses cause cancer by inducing changes that affect cell growth and division • Oncogenes frequently interfere with the normal functioning of the viral genes responsible for producing new virus. • Thus tumor viruses are often defective and sometimes require ‘helper’ viruses to reproduce. • One important class of oncogene is called ras genes are a small family of related genes found normally in a variety of eukaryotic organisms ranging from yeast to humans. • They are associated with a variety of human and rodent tumors, and genes ras have been found as oncogenes in mouse and rat retroviruses. • Nucleotide sequence comparison between normal ras genes and oncogenic ones in retroviruses indicate that oncogenic ras genes contain mutations. • Thus it may be that some carcinogens cause cancer by chemically modifying the ras region of the DNA. • Normal ras genes can also produce malignancy if the ras proteins are produced in very large amounts.
- 13. 13 • This can occur if a very strong promoter is placed immediately upstream from the genes. • Cell transformation: the ability of viruses to alter cell in culture has been critical to identifying oncogenes. Control Of Gene expression ◘ Operons • Operons are groups of genes that function to produce proteins needed by the cell. ♦ There are two different kinds of genes in operons: 1- Structural genes code for proteins needed for the normal operation of the cell. - For example, they may be proteins needed for the breakdown of sugars. - The structural genes are grouped together and a single mRNA molecule is produced during their transcription. 2- Regulator genes code for proteins that regulate other genes.
- 14. 14 • An operon is a functioning unit of key nucleotide sequences including an operator, a common promoter, and one or more structural genes, which is controlled as a unit to produce messenger RNA (mRNA), in the process of protein transcription. • Operons have not been found in eukaryotes ◘ The Operon as a unit of transcription • An operon contains one or more structural genes which are transcribed into one polycistronic mRNA: a single mRNA molecule that codes for more than one protein. • Upstream of the structural genes lies a promoter sequence which provides a site for RNA polymerase to bind and initiate transcription. • Close to the promoter lies a section of DNA called an operator. • The operon may also contain regulatory genes such as a repressor gene which codes for a regulatory protein that binds to the operator and inhibits transcription. • Regulatory genes need not be part of the operon itself, but may be located elsewhere in the genome. • The repressor molecule will reach the operator to block the transcription of the structural genes.
- 15. 15 • In bacteria, control of the rate of transcriptional initiation is the predominant site for control of gene expression. • As with the majority of prokaryotic genes, initiation is controlled by two DNA sequence elements that are approximately 35 bases and 10 bases, respectively, upstream of the site of transcriptional initiation and as such are identified as the -35 and -10 positions.
- 16. 16 • These 2 sequence elements are termed promoter sequences, because they promote recognition of transcriptional start sites by RNA polymerase. • The consensus sequence for the -35 position is TTGACA, and for the -10 position, TATAAT. • (The -10 position is also known as the Pribnow-box.) • These promoter sequences are recognized and contacted by RNA polymerase. Promoter
- 17. 17 Prokaryotic Promoter ♦ The lac operon Glucose Absent, Lactose Present: Transcription of the lac operon. • The lac operon of the model bacterium Escherichia coli was the first operon to be discovered and provides a typical example of operon function. • It consists of three adjacent structural genes, a promoter, a terminator, and an operator.
- 18. 18 • The lac operon is regulated by several factors including the availability of glucose and lactose. • For E. coli, glucose is the preferred sugar. • When glucose is present, there is no need to use lactose: these genes are not transcribed. • When there is no glucose, E. coli has to use other sugars. • IF lactose is present, the genes for lactose utilization will be made. • Conversely, if there is no lactose, they genes remain locked down. Glucose Present: No transcription Glucose Absent: Minimal transcription
- 19. 19 ♦There are two modes of regulation of the initiation of transcription in operons: Positive control mode - Where the interaction between the regulatory protein and regulatory region on the DNA turn the transcription on. - The genes are off by default and are turned on by the activators. - Transcription factors interact with the RNA polymerase and assist the enzyme in initiating transcription at the promoter. - This positive fashion of controlling gene expression is more common in eukaryotes than in prokaryotes. Negative control mode: - Where the interaction turns the genes off. - In this case, a repressor protein binds the operator, a DNA sequence of approximately 20 to 25 nucleotides, which is next to the promoter or juxtaposed, and prevents the RNA polymerase from initiating transcription. - To switch on the system, small molecules called inducers trigger the production of proteins by binding to the repressor protein and changing its conformation. - This change alters the operator-repressor interaction, so that the repressor can no longer remain attached to the operator. - Negative control is widely used among prokaryotes, which need to respond swiftly to changes in the environment.
- 20. 20 Regulation in Prokaryotes - Bacteria have a simple general mechanism for coordinating the regulation of genes that encode products involved in a set of related processes. - The gene cluster and promoter, plus additional sequences that function together in regulation are called an operon. The Lactose Operon (lac operon) - The lactose operon of E. coli encodes the enzyme b- galactosidase which hydrolyzes lactose into galactose and glucose.
- 21. 21 Lac operon • The Z gene encodes for b-galactosidase. • The Y gene encodes a permease that facilitates the transport of lactose into the bacterium. • The A gene encodes a thiogalactoside transacetylase whose function is not known. • All three of these genes are transcribed as a single, polycistronic mRNA. • Polycistronic RNA contains multiple genetic messages each with its own translational initiation and termination signals. ♦ Negative Control of the lac Operon • The protein that inhibits transcription of the lac operon is a tetramer with four identical subunits called lac repressor. • The lac repressor is encoded by the lacI gene, located upstream of the lac operon and has its own promoter. • Expression of the lacI gene is not regulated and very low levels of the lac repressor are continuously synthesized. • Genes whose expression is not regulated are called constitutive genes.
- 22. 22 • In the absence of lactose the lac repressor blocks the expression of the lac operon by binding to the DNA at a site, called the operator that is downstream of the promoter and upstream of the transcriptional initiation site. • The operator consists of a specific nucleotide sequence that is recognized by the repressor which binds very tightly, physically blocking (strangling) the initiation of transcription. • The lac repressor has a high affinity for lactose. • When a small amount of lactose is present the lac repressor will bind it causing dissociation from the DNA operator thus freeing the operon for gene expression. • Substrates that cause repressors to dissociate from their operators are called inducers and the genes that are regulated by such repressors are called inducible genes.
- 23. 23 Lac Operon
- 24. 24 Negative regulation by repressor •الdiagramsللتوضيح كدا غير علينا اللي هي اسود بإطار اللي♥
