Plasmid Conjugation Systems Explained
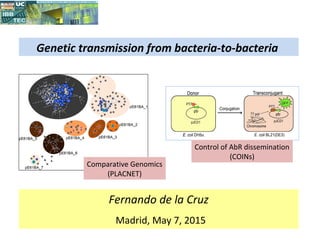
- 1. Fernando de la Cruz Madrid, May 7, 2015 Genetic transmission from bacteria-to-bacteria Comparative Genomics (PLACNET) Control of AbR dissemination (COINs)
- 2. THE DIVERSITY OF PLASMID CONJUGATION SYSTEMS Introductory slides
- 3. Elements of bacterial conjugation DONOR RECIPIENT Two corollaries: 1. The only element in common among all conjugatively transmissible plasmids is the relaxase 2. Conjugative plasmids have relaxases and VirB4s (the most conserved protein of T4SSs).
- 4. Distribution of conjugative, mobilizable and non-transmissible plasmids (1,730 plasmid genomes) according to plasmid size Smillie C, MP Garcillan-Barcia, MV Francia, EP Rocha and F de la Cruz. 2010. Mobility of plasmids. Microbiol Mol Biol Rev 74:434-52.
- 5. Main relaxase families in γ-proteobacteria 260 Tra+ plasmids in NCBI γ-proteobacteria (271 relaxases) DPMT: 19 primer pairs targeting 17 main subfamilies theoretically detect 193/271 (71%) γ- proteobacterial relaxases Garcillan-Barcia et al. 2009. The diversity of conjugative relaxases and its application in plasmid classification. FEMS Microbiol Rev 33:657-87. Alvarado et al. 2012. A Degenerate Primer MOB Typing (DPMT) Method to Classify Gamma-Proteobacterial Plasmids in Clinical and Environmental Settings. PloS One 7, e40438.
- 6. 1 2 1 Permissive host Non-permissive host 3 Relationship between transmissible plasmids and ICEs A. There are twice as many ICEs than plasmids in sequenced genomes B. ICEs and plasmids are indistinguishable by phylogeny or protein signatures C. Plasmids and ICEs frequently interconvert (at an evolutionary scale) D. A substantial fraction of integrated elements were domesticated as protein secretion specialists August 2011 | Vol 7 | e1002222
- 7. PLASMID ANALYSIS FROM WGS DATASETS Plasmid Genomics
- 8. PLACNET = Plasmid Constellation Networks PLACNET components 2 types of nodes Contigs Reference sequences 2 types of edges Scaffold links Homology to references In PLACNET, a plasmid is a connected component (a constellation of contigs) The network contains one or more disjoint connected components: chromosomes and plasmids Staphylococcus aureus 118 ST772 Chr: 2.9 Mb Plasmid: 12,819 kb Lanza et al. (2014). Plasmid flux in Escherichia coli ST131 sublineages, analyzed by plasmid constellation network (PLACNET), a new method for plasmid reconstruction from whole genome sequences. PLoS Genet 10: e1004766.
- 9. Example I. ESBL+ ST131 E. coli 3.629.034 bp core genome
- 10. IncF plasmids (ESBL genes) Hierarchical clustering dendrogram of reconstructed ST131 plasmids 43 plasmids 15 MOB families
- 12. Example II. ESBL+ E. coli of same ST type isolated from humans and animals (32 strains =148 plasmids)
- 13. Phylogeny of reconstructed MOBP12 plasmids demonstrates plasmid transmission between animals and humans
- 14. Hierarchical clustering dendrogram of reconstructed MOBP12/IncI plasmids
- 15. THE WORLD OF PLASMIDS Plasmidomics
- 16. 148 plasmids 70 plasmids 37 plasmids 59 strains 255 plasmids (4.3 plasmids / genome) Distribution of plasmids in three Escherichia coli collections 32 ESBL-positive strains Different locations in The Netherlands 2006-2011 Source: human, chicken, chicken meat, pig, de Been et al. Plos Genet 10: e1004776 19 Carbapenem-R strains Bioproject (PRJNA202876) Submitted by Broad Institute (May 2013) Source: human clinical isolates (three hospitals in Boston-USA) de Toro et al. Mic Spectrum (in press) 8 ST131 strains 4 strains virotypes A, B, C and D (Blanco et al., 2013) 4 strains (Bioproject PRJNA186413) Source: human, Lanza et al. Plos Genet 10: e1004766
- 17. 3 3.1 3.2 3.3 3.4 3.5 3.6 3.7 3.8 3.9 4 4.1 4.2 4.3 4.4 4.5 4.6 4.7 4.8 4.9 5 5.1 5.2 5.3 5.4 5.5 0 5 10 15 20 25 30 35 40 MOBH11 MOBC12 MOBC1 MOBF12 MOBF11 MOBQ12 MOBQu MOBP12 MOBP11 MOBP6 MOBP5 MOBP3 MOBV2 No-MOB Size of plasmids (Kb) [logarithmic scale] Totalnumber Distribution of MOB types (255 E.coli plasmids) 1 1.5 3 5 10 30 100 300 Smillie C, Garcillán-Barcia MP, Francia MV, Rocha EPC, de la Cruz F. Mobility of plasmids. Microbiol Mol Biol Rev 74, 434–52 (2010). Garcillán-Barcia MP, Alvarado A, de la Cruz F. Identification of bacterial plasmids based on mobility and plasmid population biology. FEMS Microbiol Rev 33, 936-56 (2011). De Toro M, Garcillán-Barcia MP, de la Cruz F. (2014) Microb Spectrum (in press)
- 18. 3 3.1 3.23.3 3.43.5 3.6 3.7 3.83.9 4 4.1 4.24.3 4.4 4.5 4.6 4.7 4.84.9 5 5.1 5.25.35.4 5.5 0 5 10 15 20 25 30 35 40 Rep-3 [pfam01051] Rep-HTH-Crp2 [pfam13545] Rep-1 [pfam01446] Rep-HTH-27 [pfam13463] Rep-HTH-36 [pfam13730] RepFIB (pILF82-like) RepQ12 (pSC101-like) RepQu (pMG828-2-like) RepQu (pIGWZ12-like) RepQu (ColE9-J-like) IncHI2 IncHI1 IncI2 IncI1, IncK, IncB/O IncN IncX IncP1 IncF ColE1-like N.D. Size of plasmids (Kb) [logarithmic scale] Totalnumber Distribution of Inc-types (255 E.coli plasmids) 1 1.5 3 5 10 30 100 300
- 19. Some pending questions (for plasmid biology) Why some plasmids appear frequently and others do not? What is unique about IncF or IncI plasmids and their adaptation to E. coli? What are the causes for synergistic plasmid combinations (e.g., IncF and ColE)? TRA/MOB? Why so many cryptic (selfish?) no-MOB plasmids?
- 21. Disarming MDR/Vir bacteria Of the 3 potential target processes, only conjugation seems to occur by a single major mechanism Lesson from bacteria: Fight AbR dissemination now, or it might be too late!
- 22. Finding COINs: R388-based high throughput conjugation assay Fernandez-Lopez et al. (2005) Unsaturated fatty acids are inhibitors of bacterial conjugation. Microbiol 151:3517-26.
- 24. COINs act on donor cells and are effective in all tested hosts
- 25. Halting plasmid dissemination with COINs Effect of 2-HDA on the kinetics of plasmid R1drd19 conjugation in liquid medium COINs effectively inhibit plasmid infection of a recipient population Experiments should be reproduced in natural environments We are testing a freshwater microcosms for inhibition of MOBF12/IncF plasmid infection by uFAs
- 26. MGEs and timescales in bacterial evolution 1. Short scale (few years): Antibiotic resistance (the first evolution experiment at a planetary scale) • Diversity • Novelty • Globality ... Appearance of MGEs (plasmids and Tns) 2. Intermediate scale (hundreds of years): Degradation of xenobiotic compounds (more local, more complex) same mechanisms but: • Sometimes phages • Sometimes inserted in chromosomes 3. Large scale (millions of years): Pathogenicity (slower, even more complex) • Mostly genomic (pathogenicity) islands ““Horizontal gene transfer and the origin of species: lessons from bacteria”Horizontal gene transfer and the origin of species: lessons from bacteria” de la Cruz and Davies (2000)de la Cruz and Davies (2000) Trends in Microbiol.Trends in Microbiol. 88: 128–133.: 128–133.
- 27. Acknowledgements The UC team Other collaborators Julien Guglielmini, Eduardo Rocha (Institut Pasteur, Paris) Marta Martinez, Antonio Fernandez (Instituto Biomar, León) Teresa Coque, Val F. Lanza, Fernando Baquero (Hosp RyC, Madrid) Jorge Blanco (Univ. Santiago de Compostela) Mark de Been, Willem van Schaik, Rob Willems (Univ. Utrech)
Editor's Notes
- (25 min + 5 min discussion) Outline: Gene transfer overview - PLACNET look at plasmid diversity - COINs
- Two corollaries: 1. The only element in common among all conjugatively transmissible plasmids is the relaxase 2. Conjugative plasmids have relaxases and VirB4s (the most conserved protein of T4SSs).
- Figure 6– Distribution of conjugative, mobilizable and non-conjugative plasmids according to plasmid size (curves were created from polynomial interpolation of the histograms of each class).
- PLACNET is a network of contig relationships (graph representation) PLACNET reconstruction for Staphylococcus aureus genome (S. aureus 118 ST772, ID: PRJNA82607). Assembly data: Number of libraries: 1; read length: 75 bp; number of contigs: 73; total bp: 2,798,022 bp; N50: 224673 and Kmer: 73. One plasmid (12,819 bp) reconstructed. No REL or RIP proteins detected.
- Figure 2. Phylogenetic tree of ST131 E. coli, based on 3.629.034 bp core genome (3,734 orthologous genes: 90% identity and 90% coverage) and 100 bootstrapping replicates.
- Figure 4. UPGMA Dendrogram based on protein cluster analysis (60% id, 80% coverage). 38 plasmids + 5 black = 43 plasmids It is important to complement this dendrogram with a second one including reference sequences. From this second one we know if a plasmid is related to other plasmids but lacks marker genes. Example: delta-MOB IncF plasmids.
- DPMT column: only groups included in Alvarado (2012) are included (ergo, some gamma-proteobacterial groups are missing (e.g. MOBP6). MPF column: type of T4SS for the given DPMT subfamily (left triangles). - : no T4SS present -> mobilizable plasmids. MPMT column: plasmid groups included are shown in darker color and indicated in black letters. Plasmids not included are shown in red letters.
- The figure shows the results of MPMT in 83 isolates. 6 multiplex reactions to identify 23 groups of closely related relaxases, which belong to 11 relaxase MOB subfamilies. FI (40 + out of 83 isolates), FII (37 + out of 83 isolates) -> 5 isolates were positive for both (FI and FII-like), 11 isolates were negative for both, 35 isolates were + only for FI and 32 isolates were only positive for FII -> 5+11+35+32=83.
- Not a single protein is conserved in all IncF genomes (no core)
- Two collections: (1) strain pairs of human and poultry associated strains that had previously been considered to be identical based on Multi-Locus Sequence Typing, plasmid typing and antibiotic resistance gene sequencing. (2) isolates from farmers and their pigs.
- Here, plasmids found in study represent separate subsets (lineages) of IncK or IncI1 plasmids epidemic plasmids
- In the small-size plasmids (1-2 Kb) no REP protein was detected; ColE1, RepQu and RepQ12 replicons were common in 2.5-10 Kb size plasmids. In large plasmids, IncF and IncI groups the most abundant.
- These questions belong to the realm of functional plasmidomics, a field in which we want to work for the next few years
- 2-HDA affected growth of Vibrio strain, but not of the others. CF = conjugation frequency
- In our view, plasmids are the spearheads of bacterial evolution
