Embed presentation
Download as PDF, PPTX





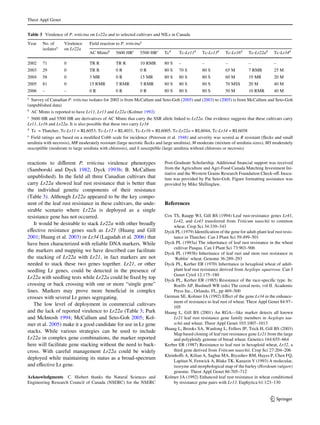

This study mapped the adult-plant leaf rust resistance gene Lr22a in wheat using microsatellite markers (SSRs). Lr22a was previously introgressed from Aegilops tauschii into wheat. Comparing SSR alleles between the donor line and resistant backcross lines identified 2-5 markers co-introgressed with Lr22a spanning 9-20 cM. Testing these markers in an F2 population confirmed linkage of the closest marker GWM296 to Lr22a, located 2.9 cM away. GWM296 accurately predicted lines known to carry Lr22a and could be useful for marker-assisted selection of this resistance gene.







