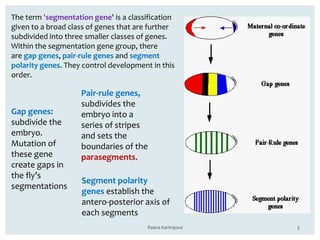
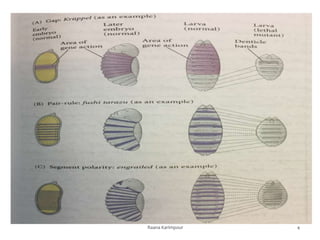
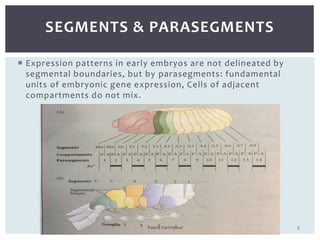
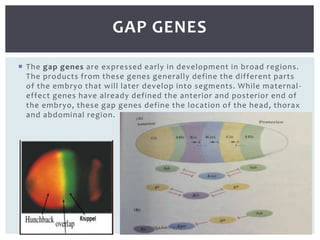
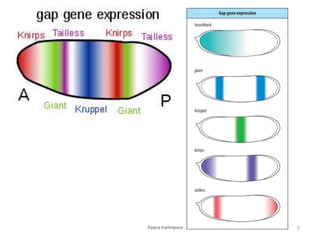
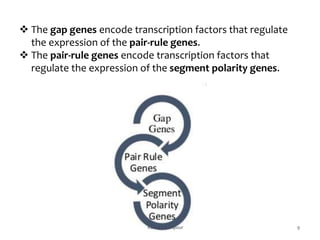
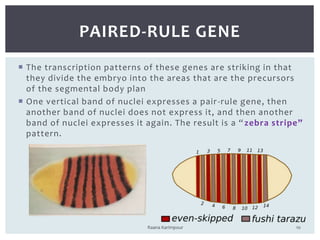
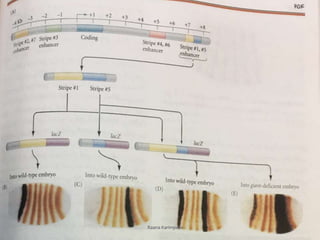
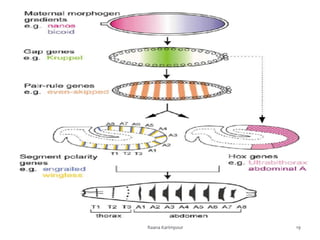
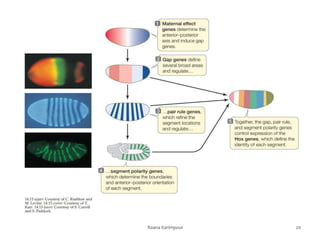
The document discusses the role of segmentation genes in Drosophila development, explaining their classification into gap genes, pair-rule genes, and segment polarity genes. It outlines how these genes control body plan segmentation through regulation of transcription factors, forming distinct patterns in early embryonic stages. The interactions between cells, mediated by segment polarity genes, determine cell fates within parasegments.