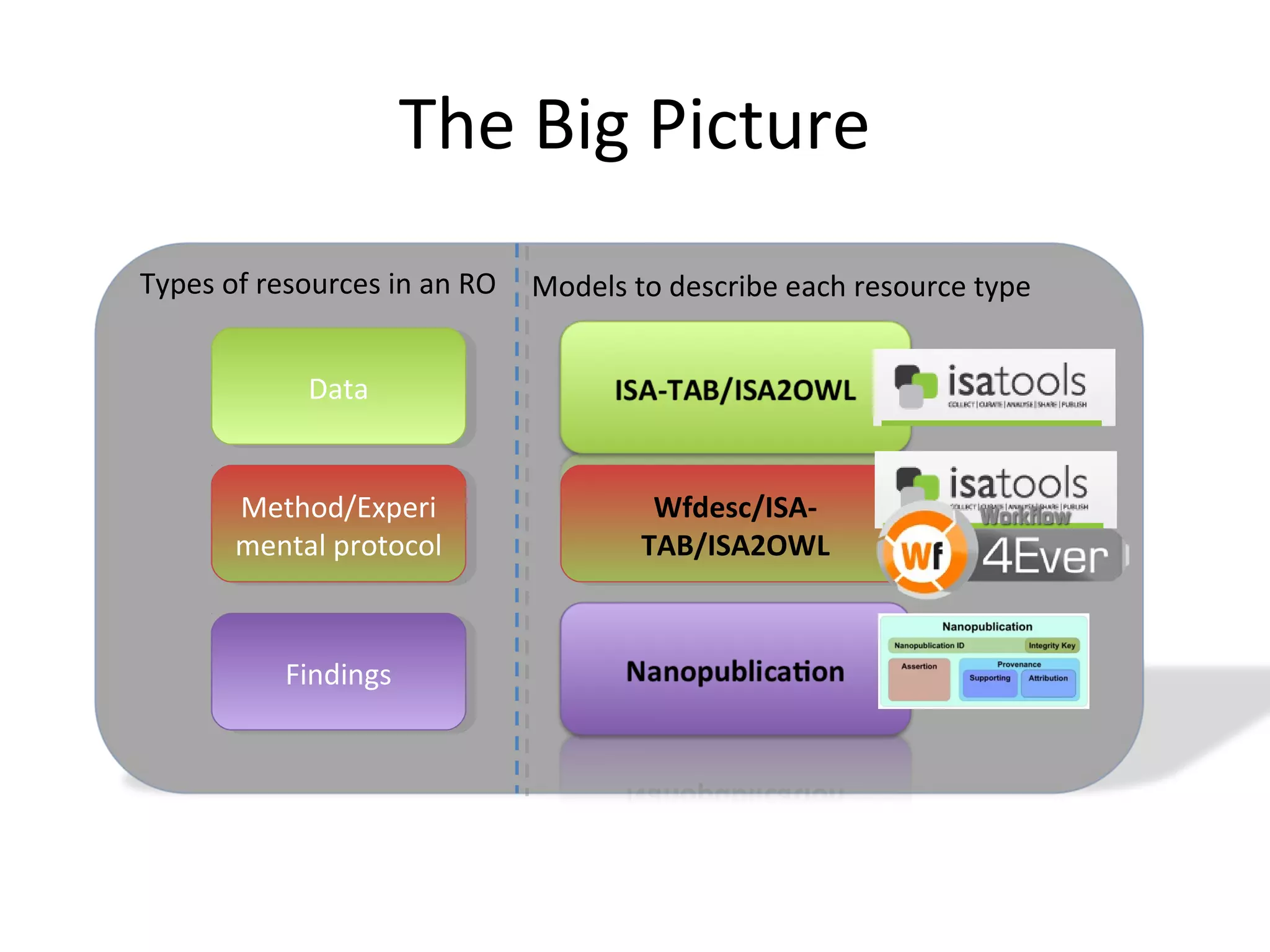
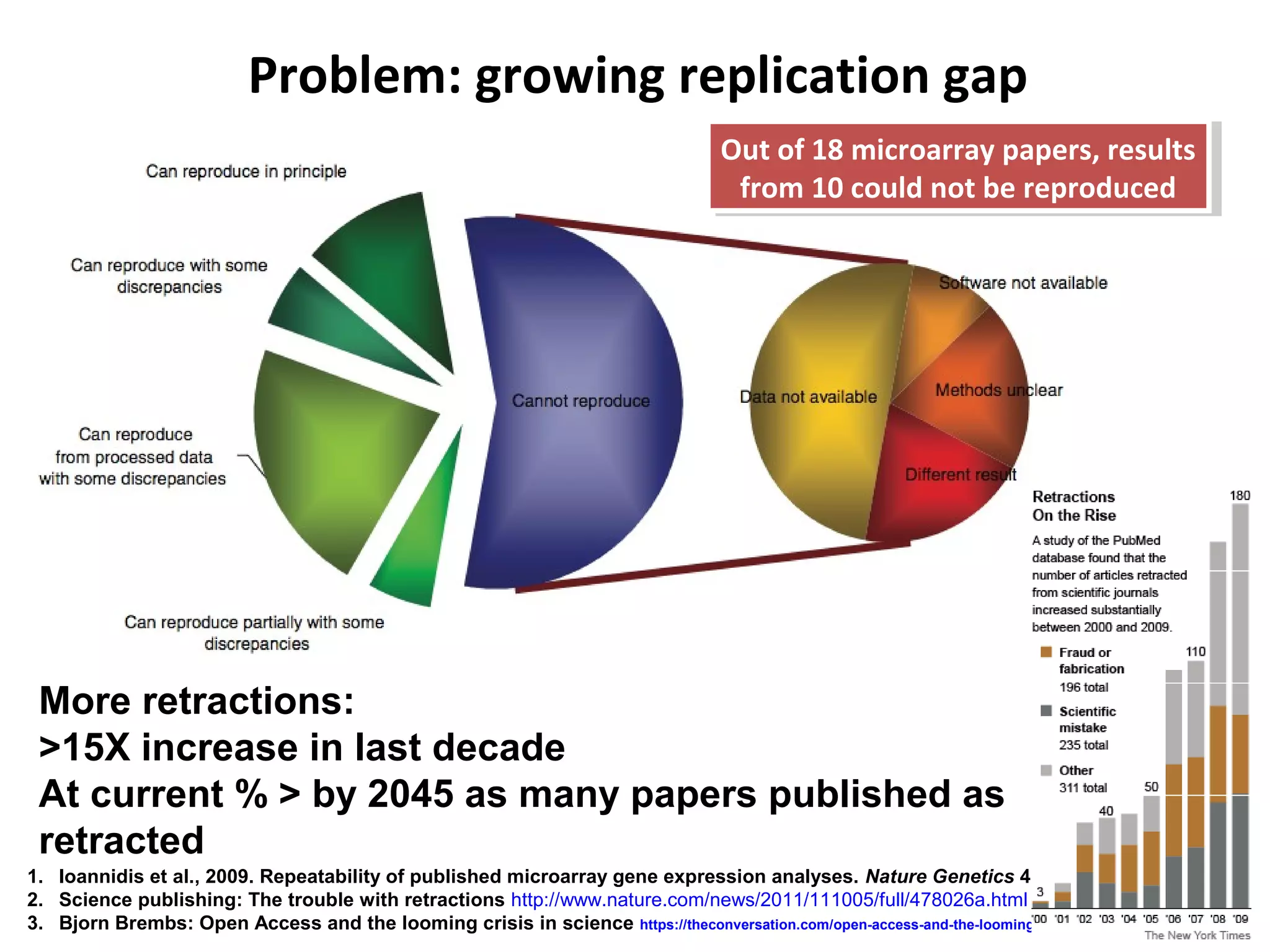
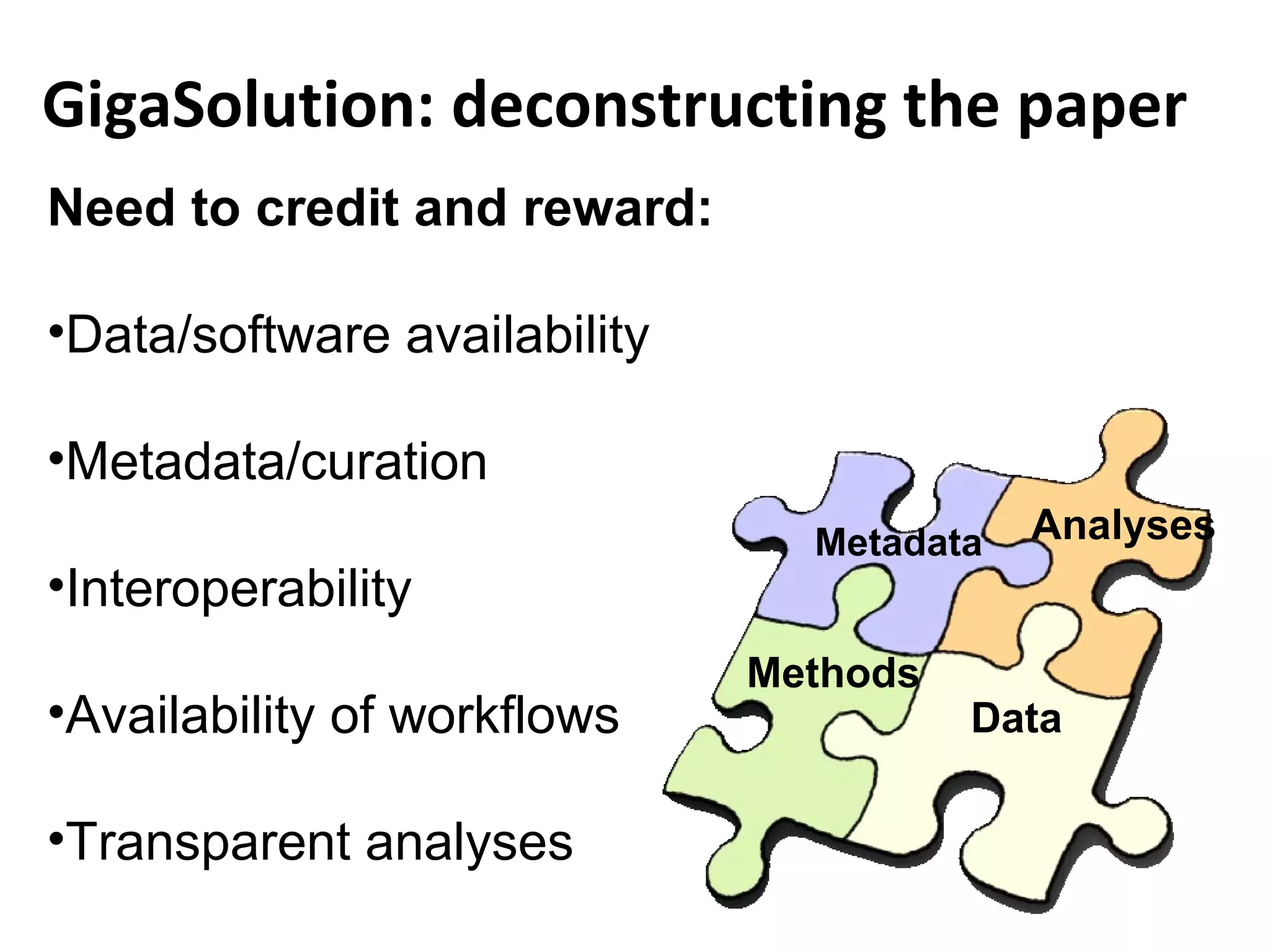
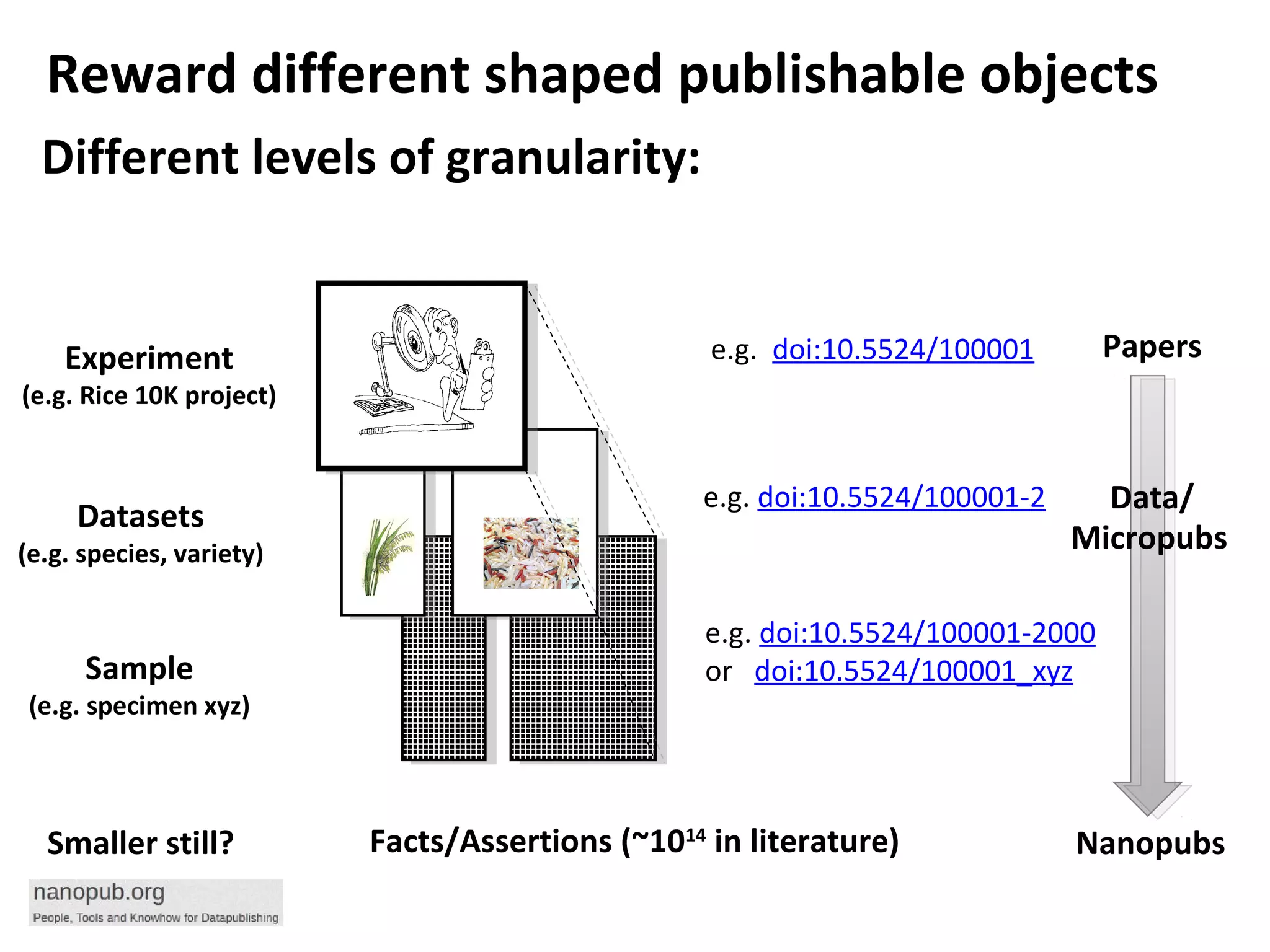
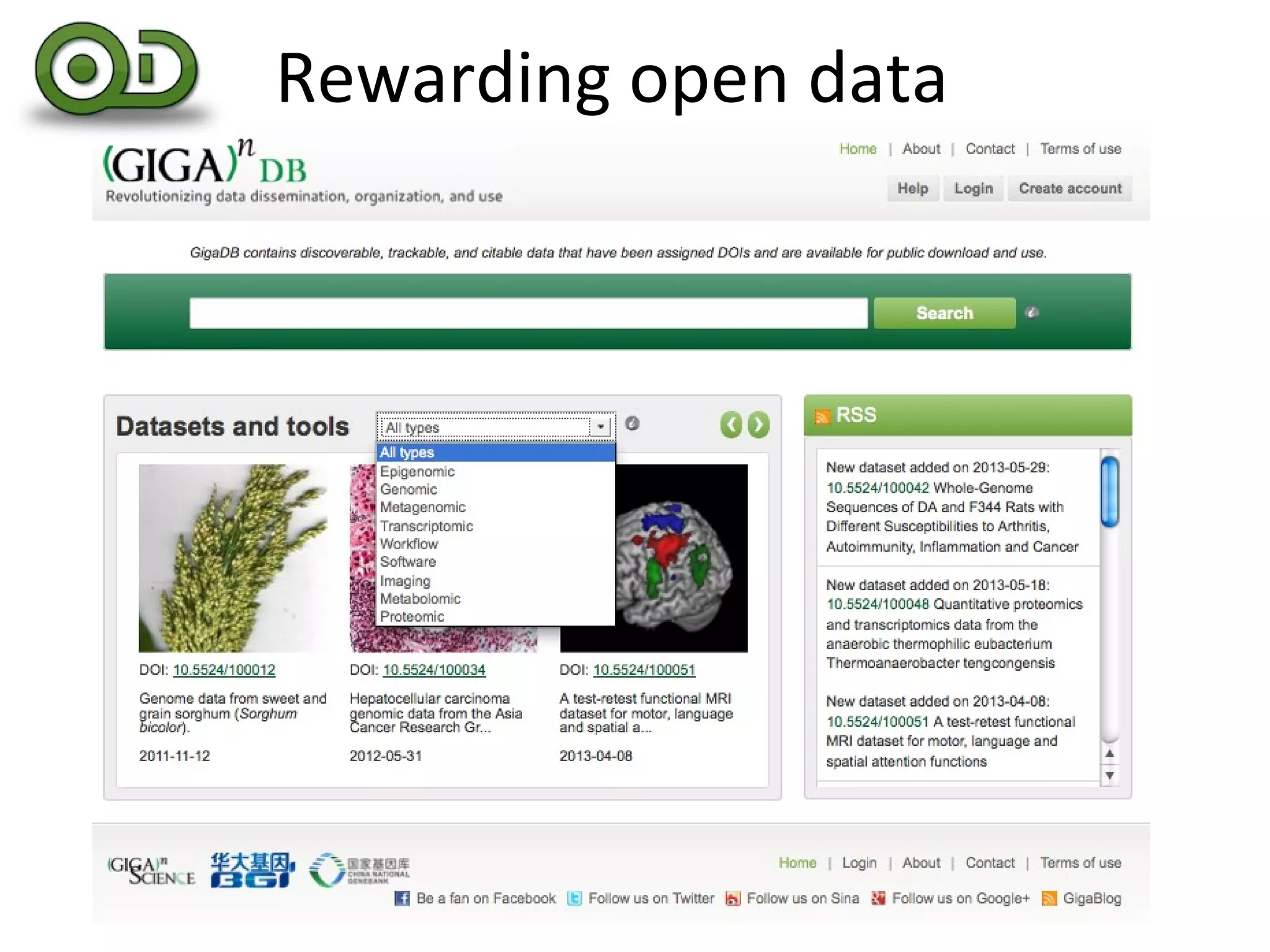
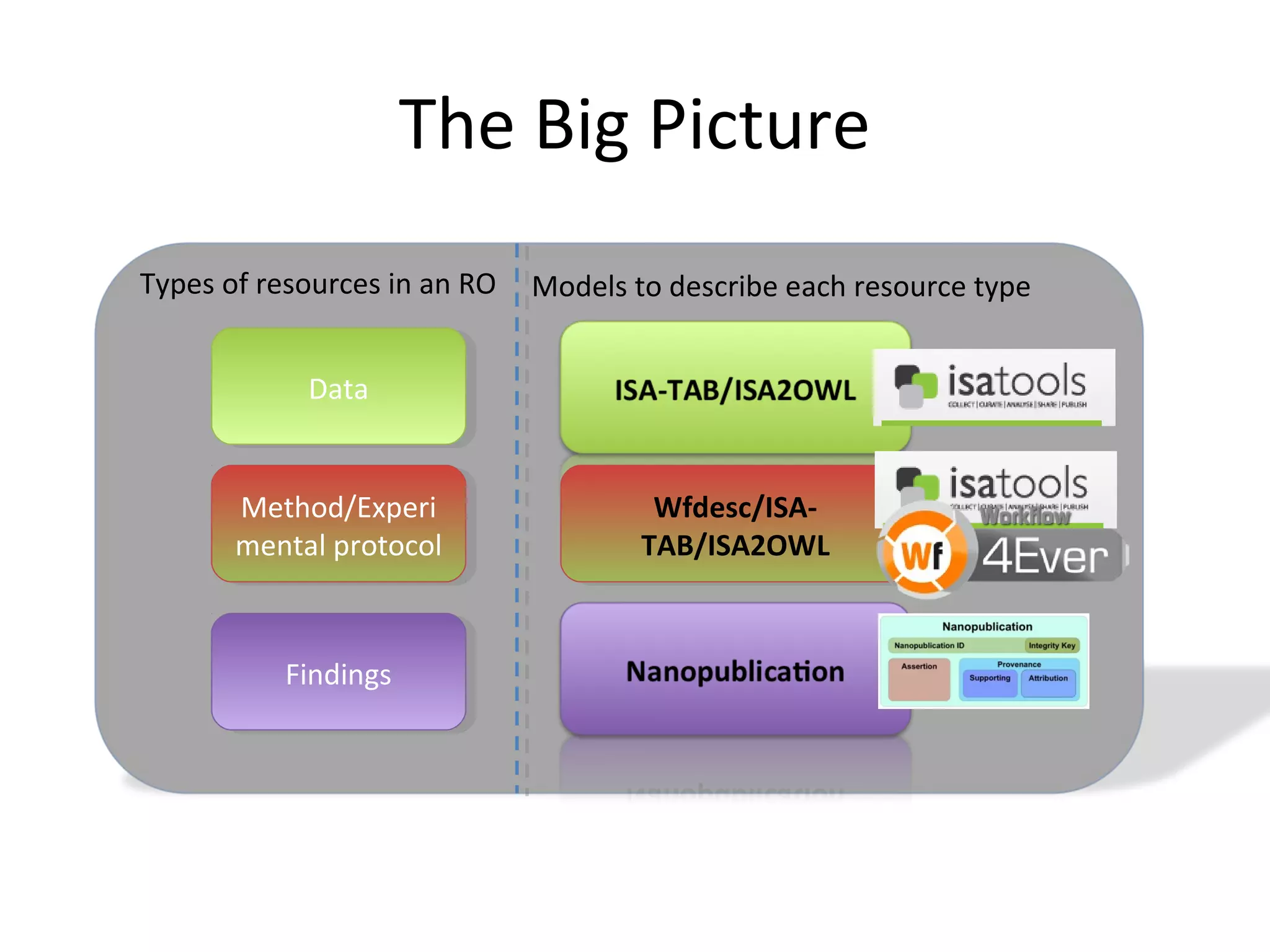
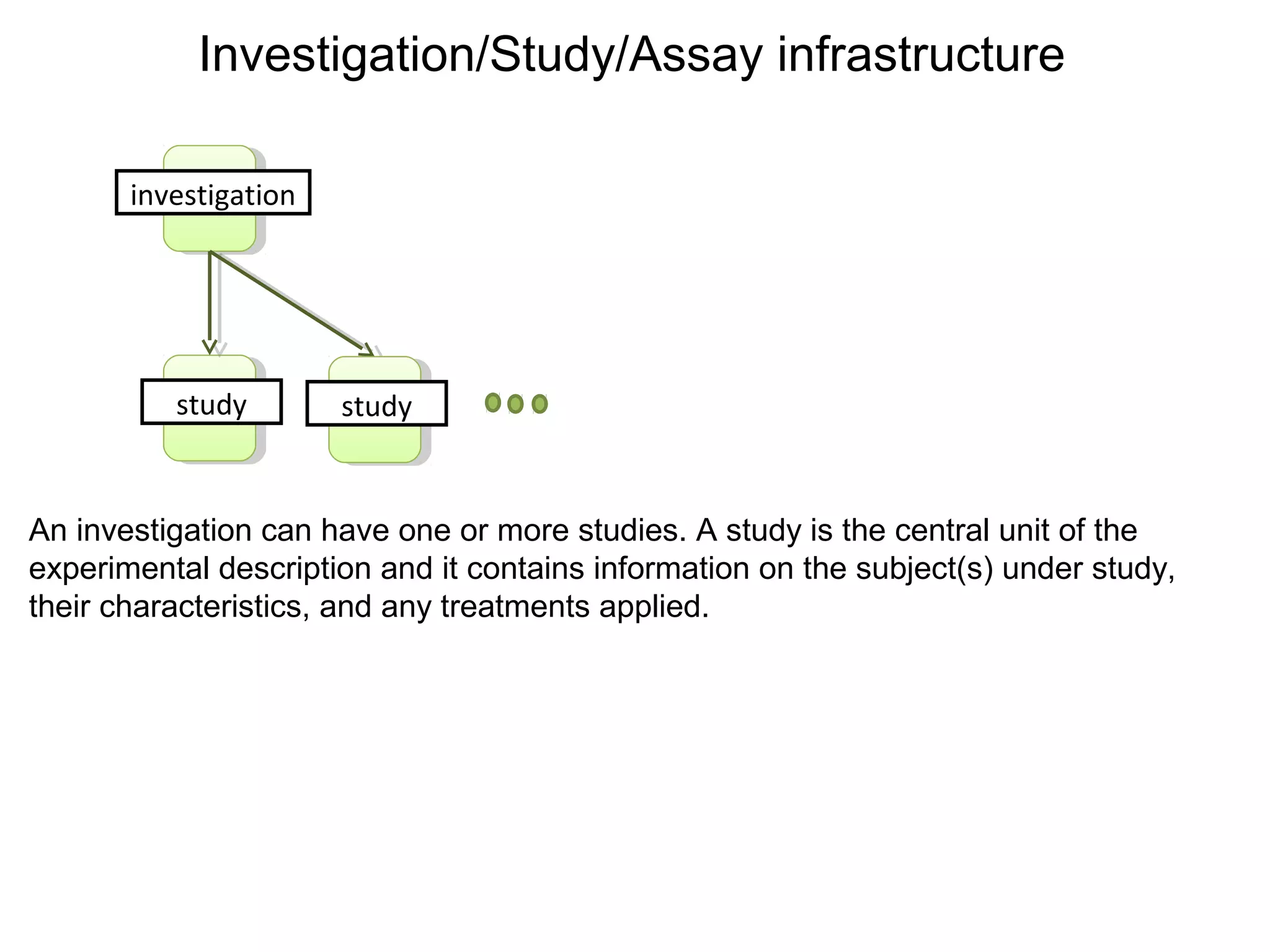
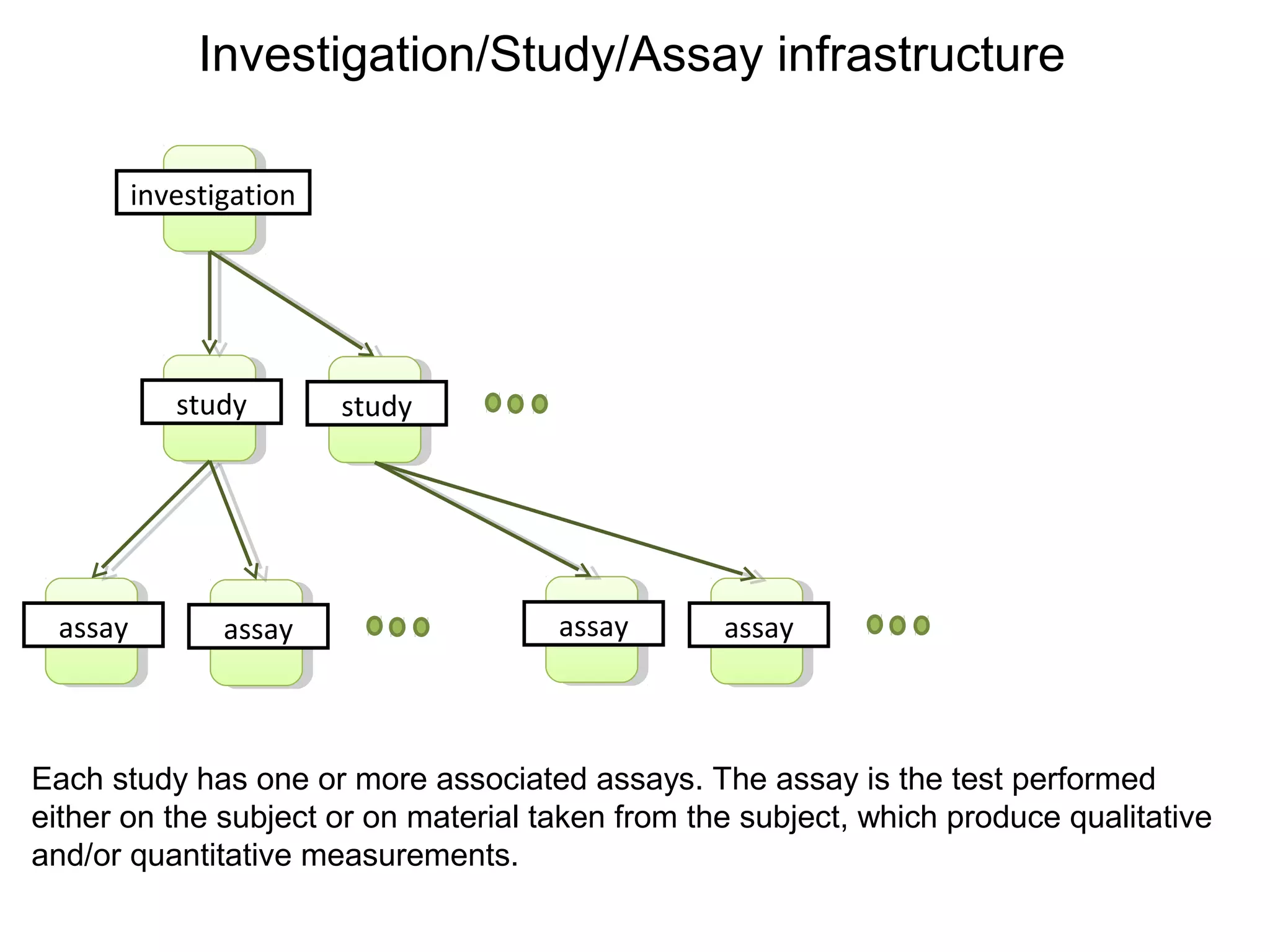
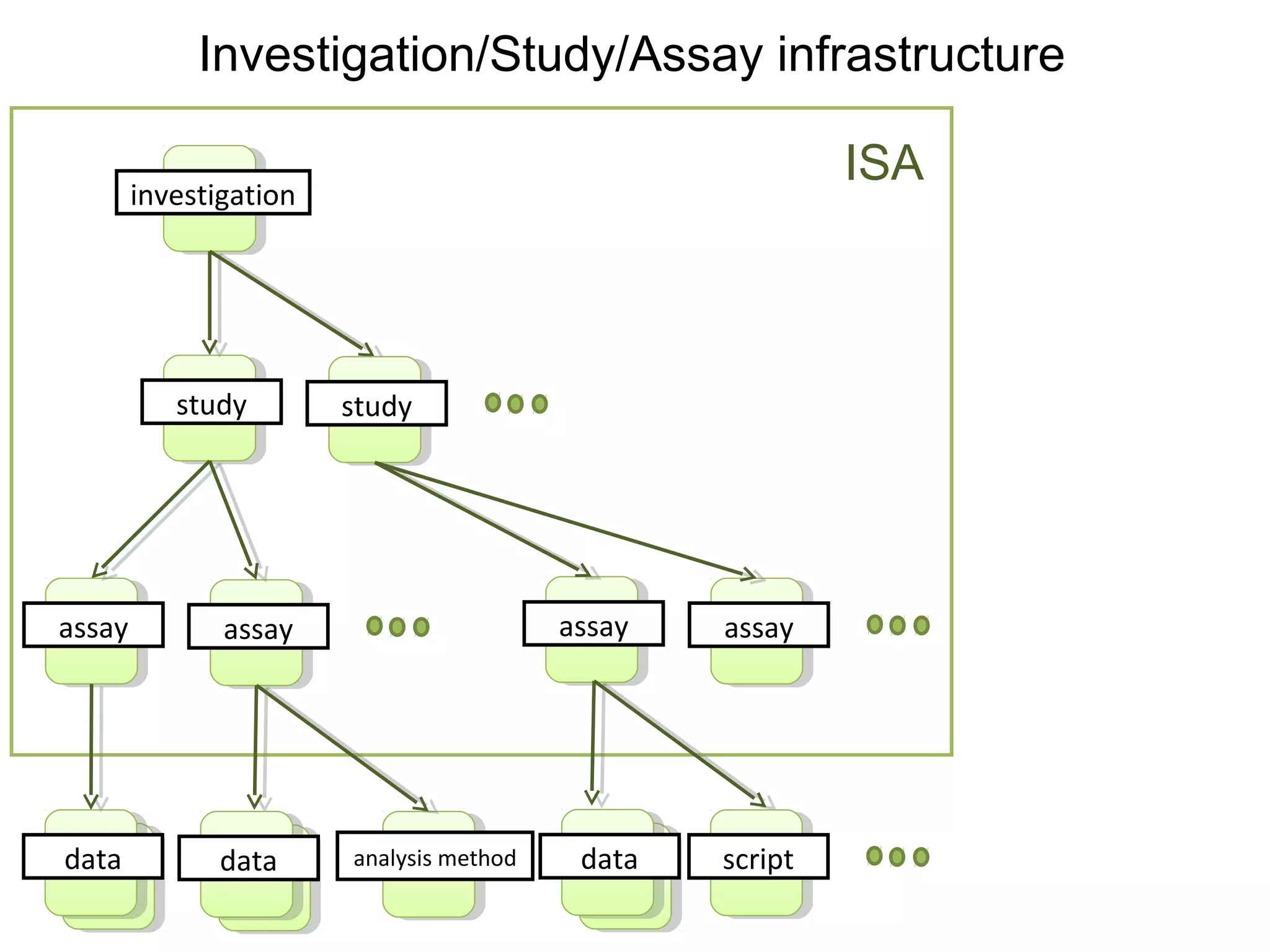
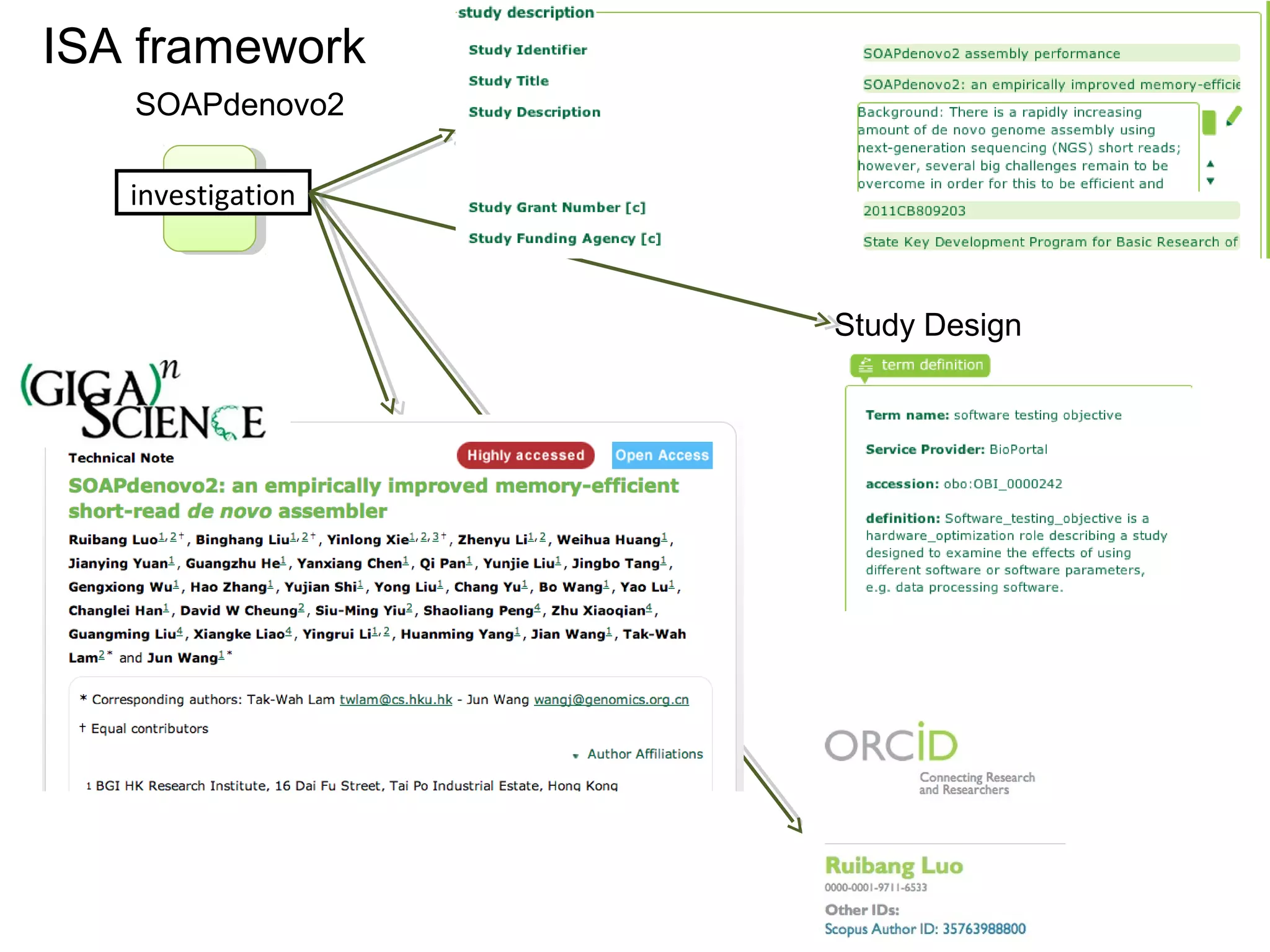
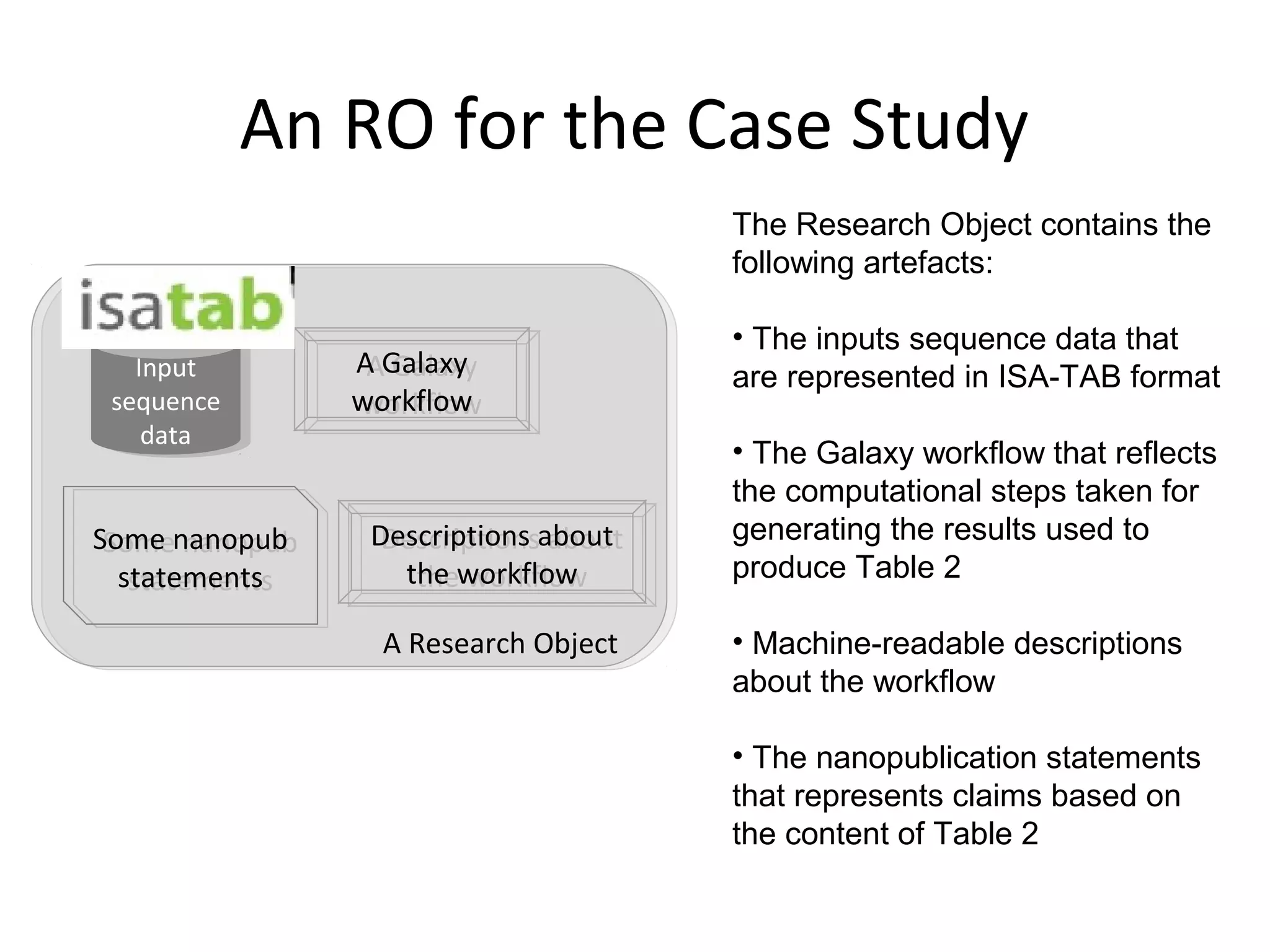
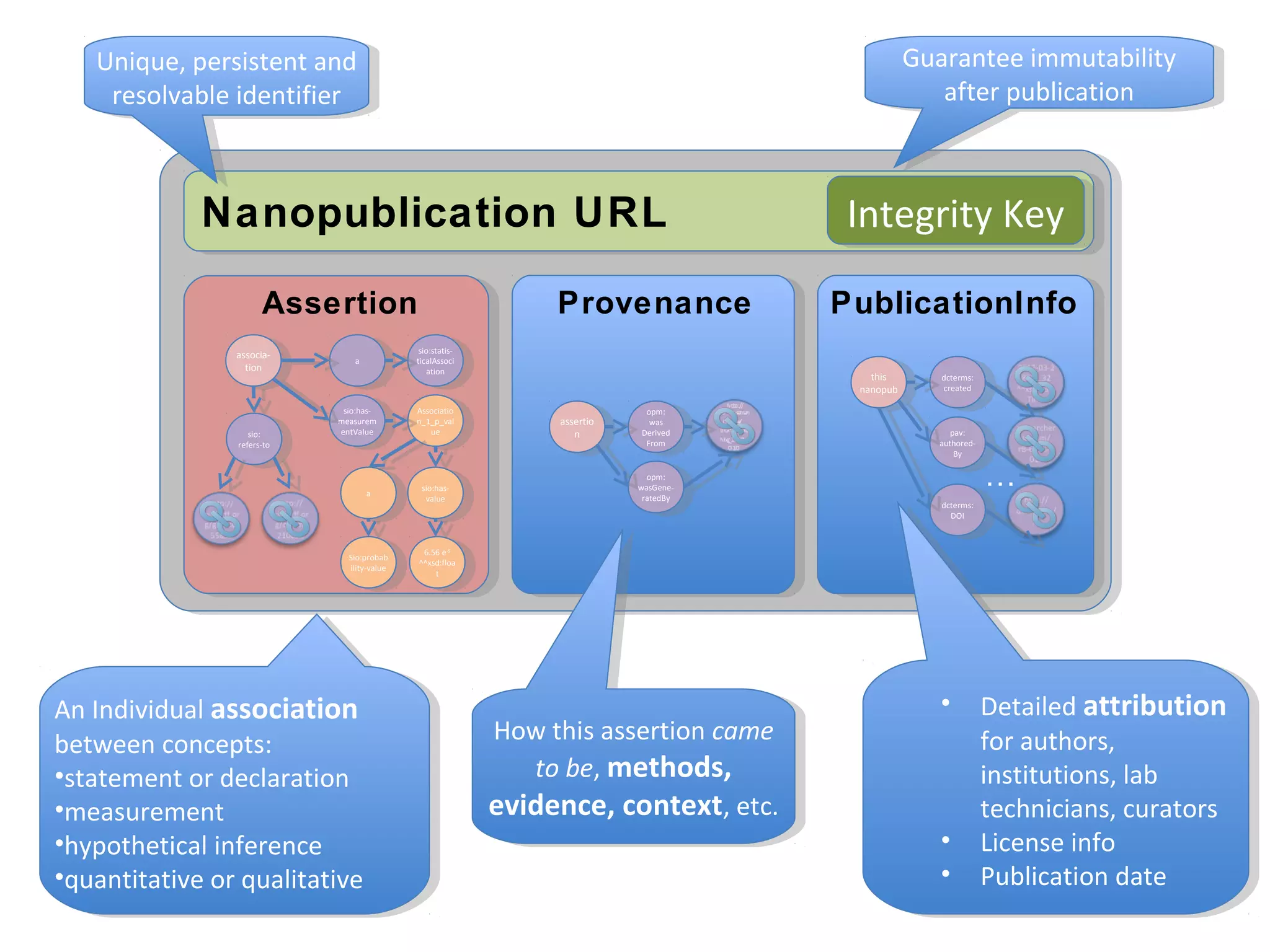
This document discusses improving reproducibility and transparency in scholarly publishing by rewarding open sharing of data, methods, and results. It presents a case study using the ISA framework to describe experimental data and workflows, nanopublications to make assertions from results, and research objects to aggregate these scholarly artifacts. Combining these approaches could help address issues like increasing retractions by incentivizing open review, replication of analyses, and credit for sharing diverse scholarly contributions beyond publications. The document advocates extending this case study to provide community guidelines for integrated use of these models.





















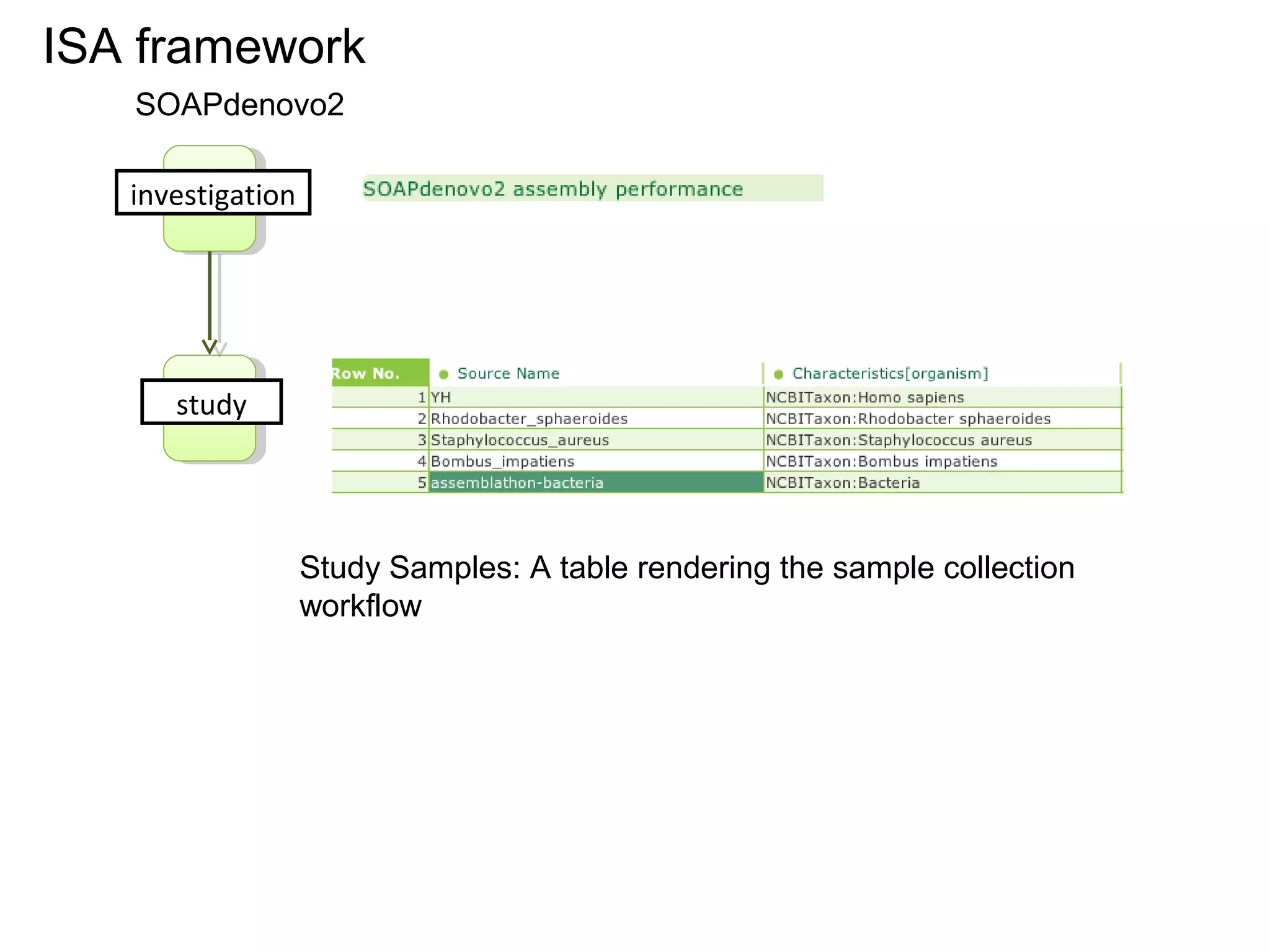
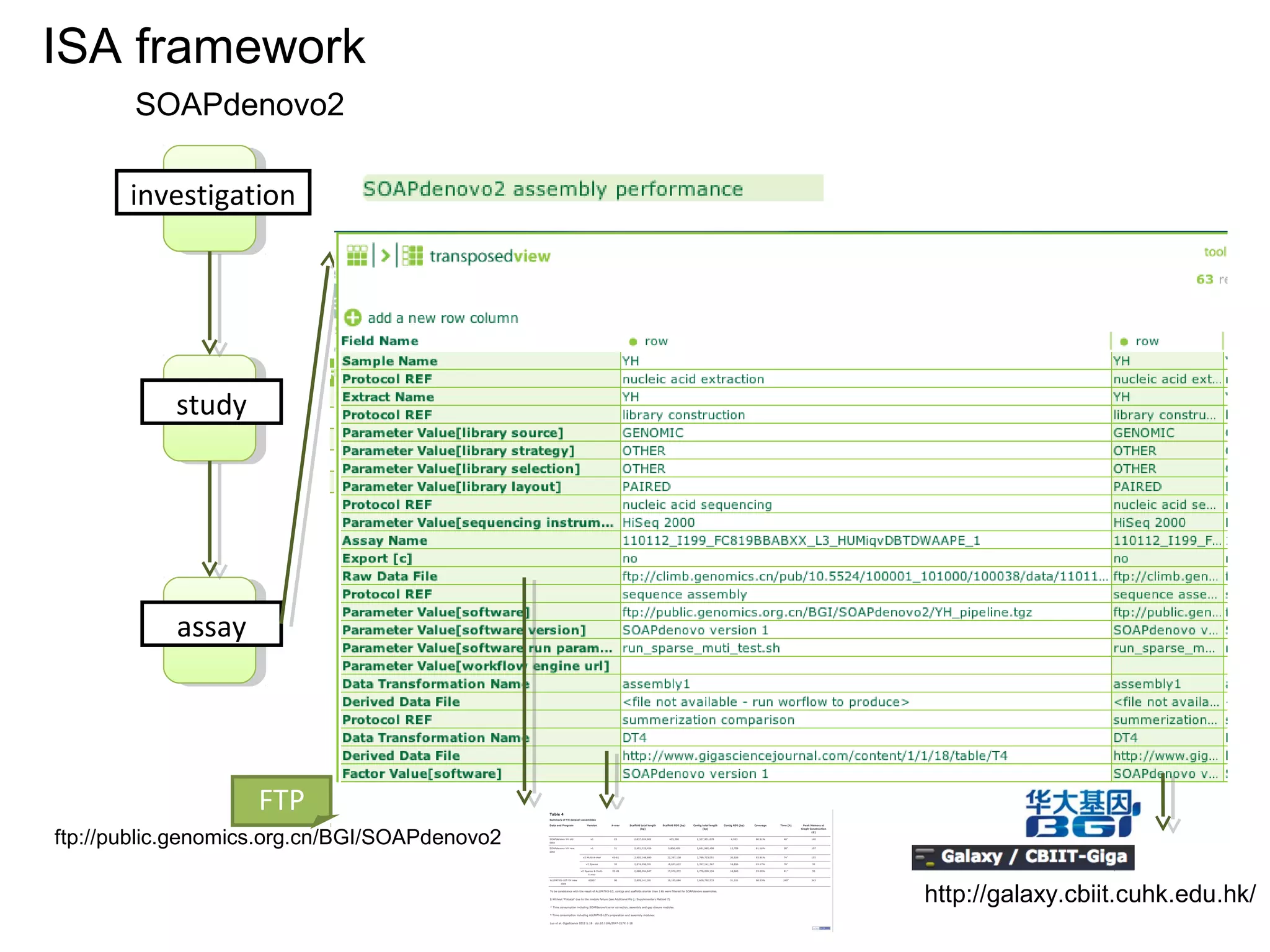
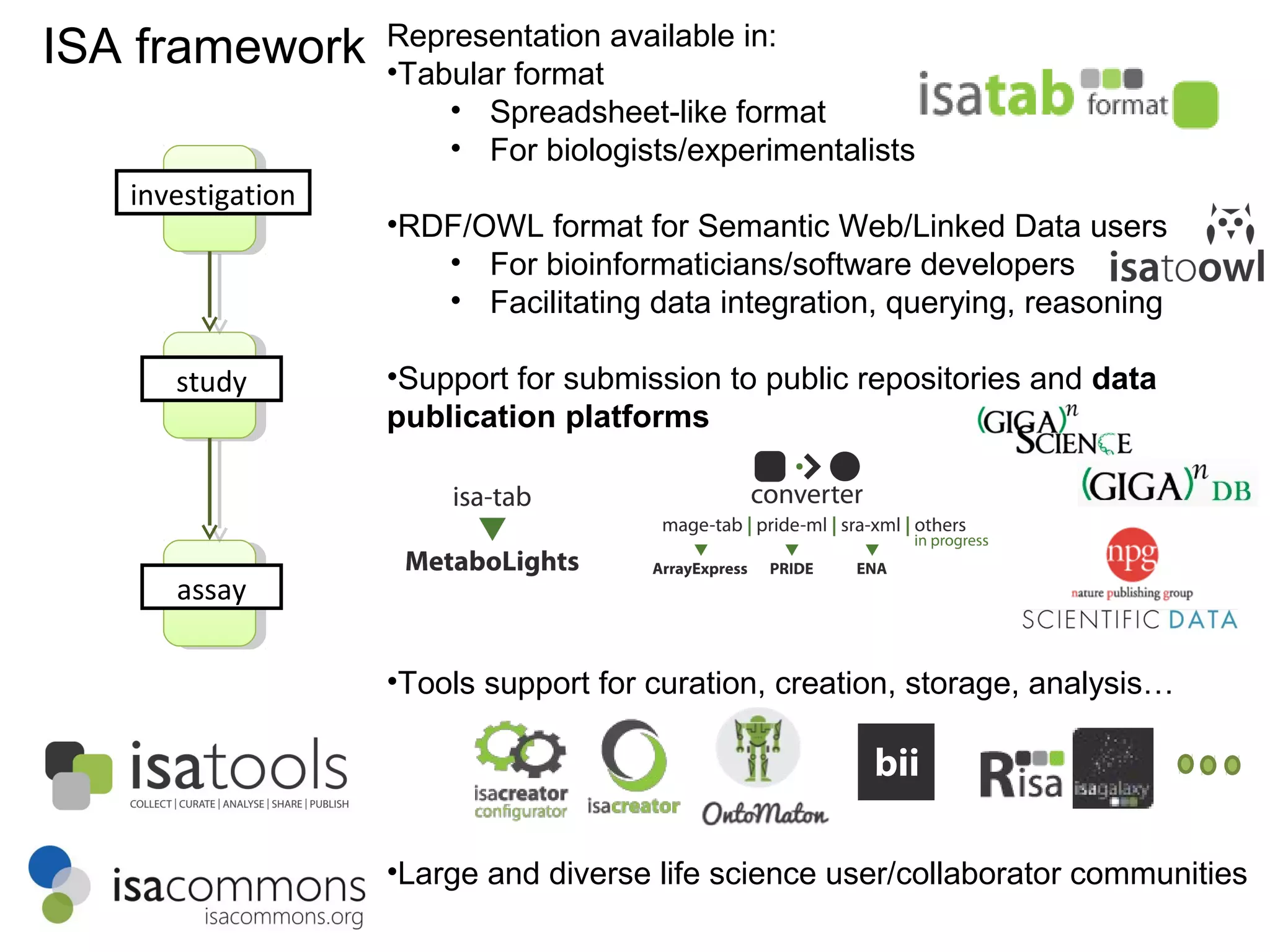
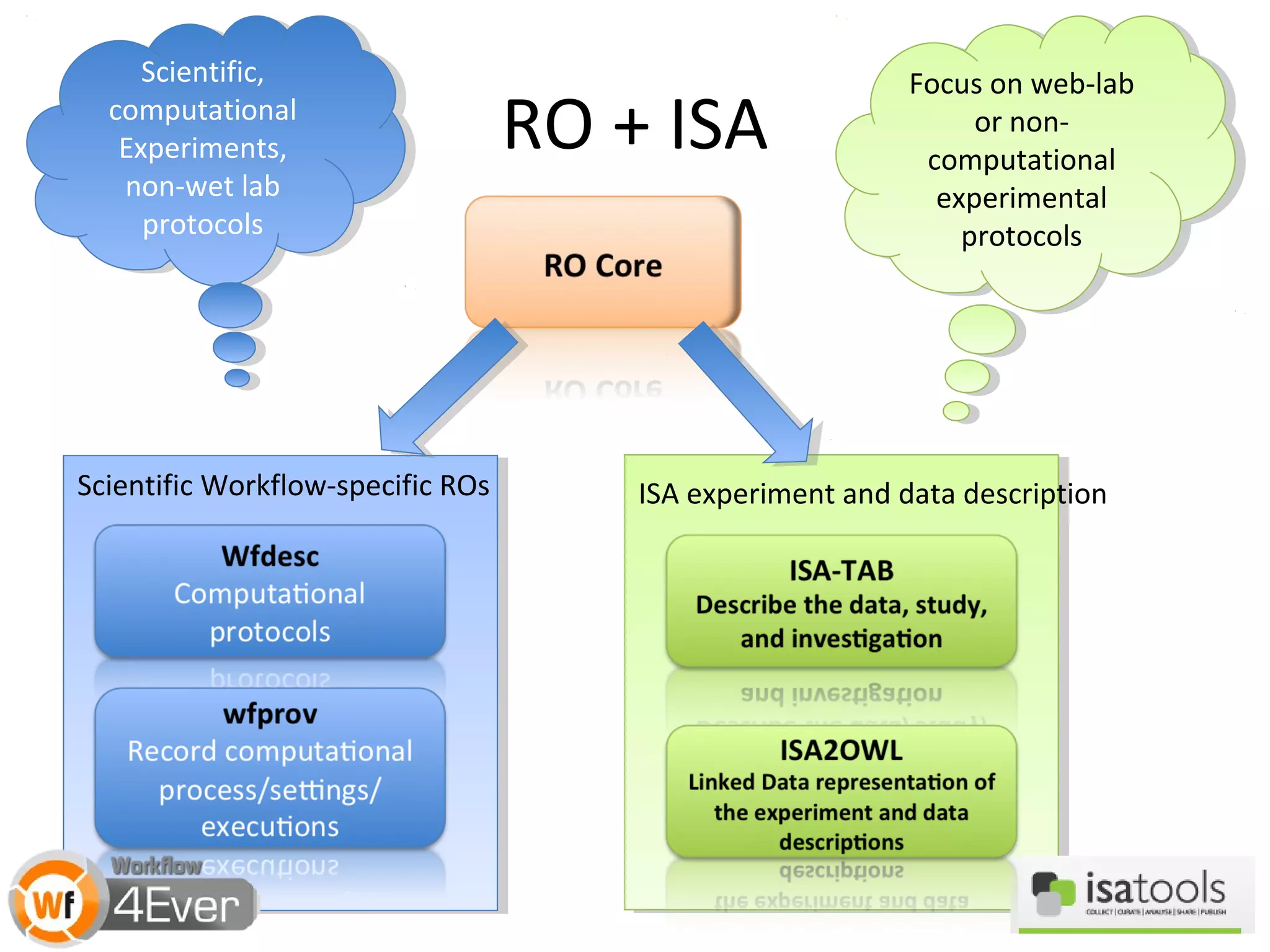
![A Nanopublication-Centric View
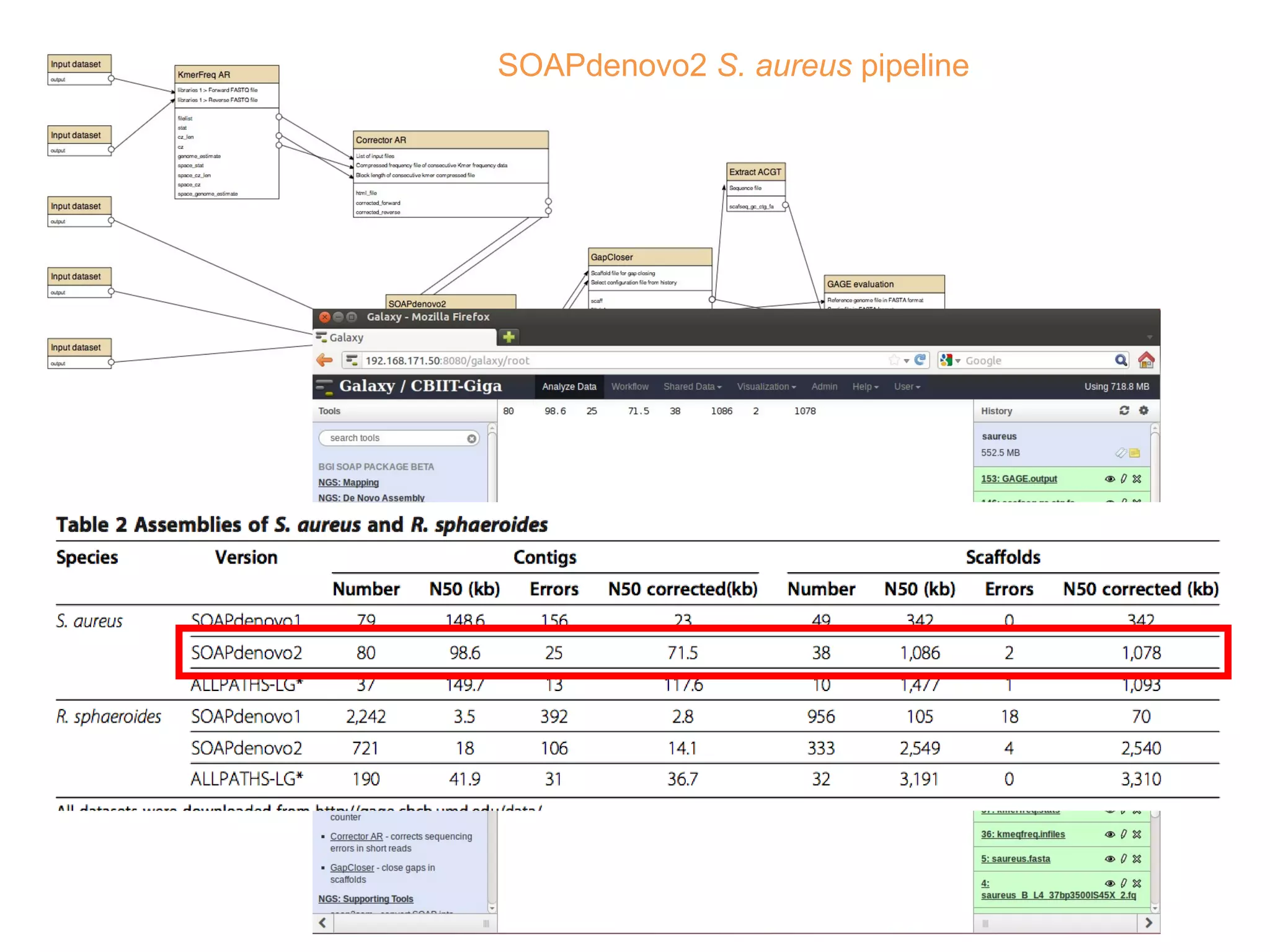
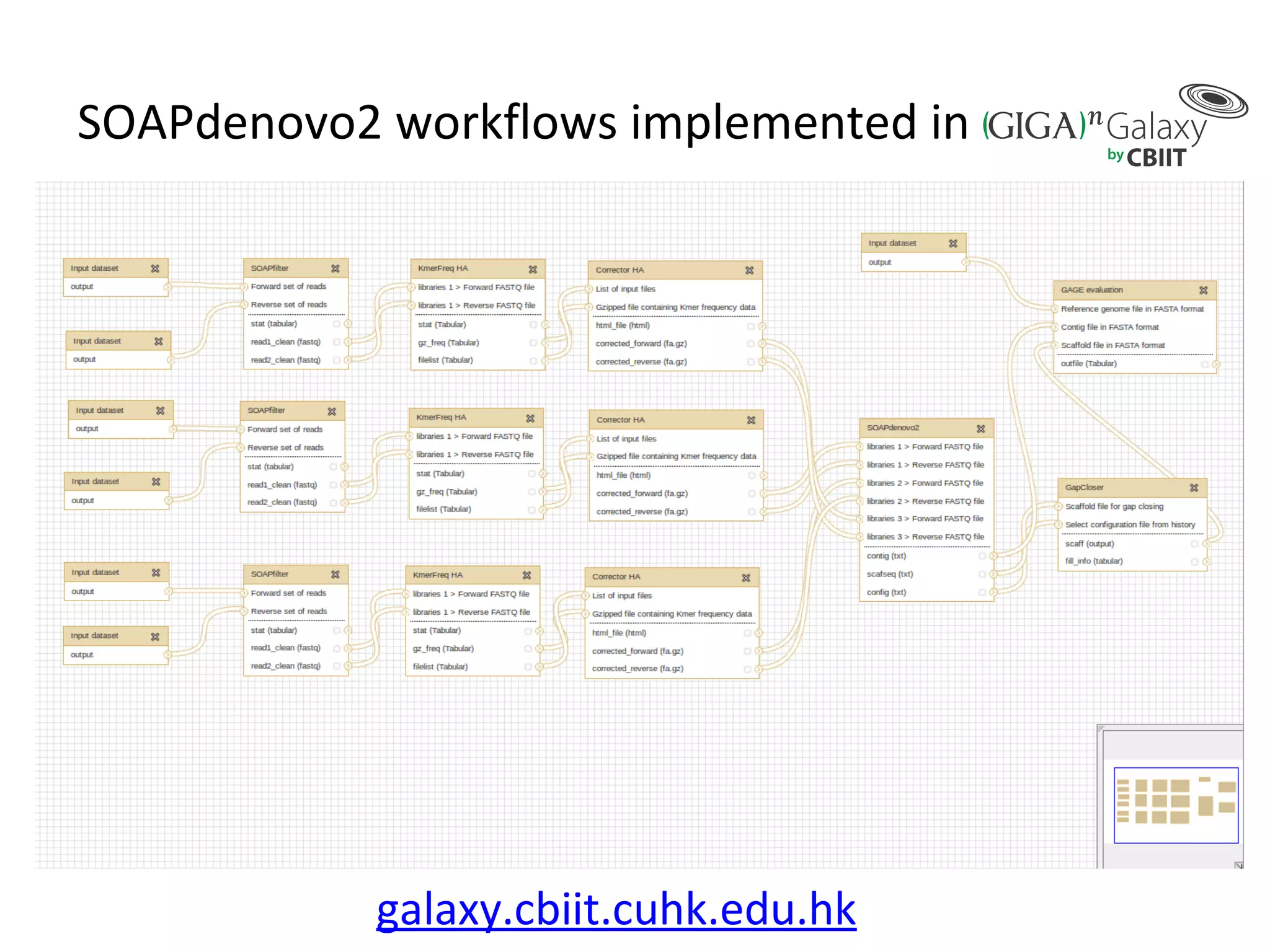
• Improvements of SOAPdenovo2 have also been observed in assembling GAGE [8]
dataset (see Additional file 1: Supplementary Method 6 and Tables 2 and 3). As
shown in Tables 2 and 3, the correct assembly length of SOAPdenovo2 increased
by approximately 3 to 80-fold comparing with that of SOAPdenovo1.](https://image.slidesharecdn.com/sceismbtalk-130722025756-phpapp01/75/Scott-Edmunds-ISMB-talk-on-Big-Data-Publishing-45-2048.jpg)